Abstract
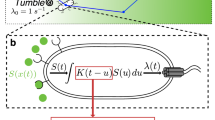
Organisms acquire and use information from their environment to guide their behaviour. However, it is unclear whether this information quantitatively limits their performance at behavioural tasks. Here we relate information to the ability of Escherichia coli to navigate up chemical gradients, the behaviour known as chemotaxis. First, we derive a theoretical limit on the speed with which cells climb gradients, given the rate at which they acquire information. Next, we measure cells’ gradient-climbing speeds and the rate of information acquisition by their chemotaxis signalling pathway. We find that E. coli cells make behavioural decisions with much less than the one bit required to determine whether they are swimming up the gradient. Some of this information is irrelevant to gradient climbing, and some is lost in communication to behaviour. Despite these limitations, E. coli cells climb gradients at speeds within a factor of two of the theoretical bound. Thus, information can limit the performance of an organism, and sensory–motor pathways may have evolved to efficiently use information acquired from the environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Source data are provided with this paper. Source data for the Supplementary figures are contained in a Supplementary Data file. Source data for figures in the main text are provided online with the manuscript.
Code availability
Code to reproduce the main text figures are available with the source data. All the algorithms used are described in detail in the Supplementary Information.
References
Cheong, R., Rhee, A., Wang, C. J., Nemenman, I. & Levchenko, A. Information transduction capacity of noisy biochemical signaling networks. Science 334, 354–358 (2011).
Uda, S. et al. Robustness and compensation of information transmission of signaling pathways. Science 341, 558–561 (2013).
Selimkhanov, J. et al. Accurate information transmission through dynamic biochemical signaling networks. Science 346, 1370–1373 (2014).
Granados, A. A. et al. Distributed and dynamic intracellular organization of extracellular information. Proc. Natl Acad. Sci. USA 115, 6088–6093 (2018).
Bowsher, C. G. & Swain, P. S. Environmental sensing, information transfer, and cellular decision-making. Curr. Opin. Biotechnol. 28, 149–155 (2014).
Tkačik, G. & Bialek, W. Information processing in living systems. Annu. Rev. Condens. Matter Phys. 7, 89–117 (2016).
Laughlin, S. A simple coding procedure enhances a neuron’s information capacity. Z. Naturforsch. C J. Biosci. 36, 910–912 (1981).
Rieke, F., Warland, D. & Bialek, W. Coding efficiency and information rates in sensory neurons. Europhys. Lett. 22, 151–156 (1993).
Petkova, M. D., Tkačik, G., Bialek, W., Wieschaus, E. F. & Gregor, T. Optimal decoding of cellular identities in a genetic network. Cell 176, 844–855.e15 (2019).
Bialek, W., De Ruyter Van Steveninck, R. R. & Tishby, N. Efficient representation as a design principle for neural coding and computation. In 2006 IEEE International Symposium on Information Theory 659–663 (IEEE, 2006).
Berg, H. C. E. coli in Motion (Springer, 2004).
Lazova, M. D., Ahmed, T., Bellomo, D., Stocker, R. & Shimizu, T. S. Response rescaling in bacterial chemotaxis. Proc. Natl Acad. Sci. USA 108, 13870–13875 (2011).
Tu, Y. Quantitative modeling of bacterial chemotaxis: signal amplification and accurate adaptation. Annu. Rev. Biophys. 42, 337–359 (2013).
Parkinson, J. S., Hazelbauer, G. L. & Falke, J. J. Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol. 23, 257–266 (2015).
Berg, H. C. & Purcell, E. M. Physics of chemoreception. Biophys. J. 20, 193–219 (1977).
Korobkova, E., Emonet, T., Vilar, J. M. G., Shimizu, T. S. & Cluzel, P. From molecular noise to behavioural variability in a single bacterium. Nature 428, 574–578 (2004).
Colin, R., Rosazza, C., Vaknin, A. & Sourjik, V. Multiple sources of slow activity fluctuations in a bacterial chemosensory network. eLife 6, e26796 (2017).
Keegstra, J. M. et al. Phenotypic diversity and temporal variability in a bacterial signaling network revealed by single-cell FRET. eLife 6, e27455 (2017).
Lan, G., Sartori, P., Neumann, S., Sourjik, V. & Tu, Y. The energy–speed–accuracy trade-off in sensory adaptation. Nat. Phys. 8, 422–428 (2012).
Clausznitzer, D., Micali, G., Neumann, S., Sourjik, V. & Endres, R. G. Predicting chemical environments of bacteria from receptor signaling. PLoS Comput. Biol. 10, e1003870 (2014).
Ito, S. & Sagawa, T. Maxwell’s demon in biochemical signal transduction with feedback loop. Nat. Commun. 6, 7498 (2015).
Micali, G. & Endres, R. G. Maximal information transmission is compatible with ultrasensitive biological pathways. Sci. Rep. 9, 16898 (2019).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Tostevin, F. & ten Wolde, P. R. Mutual information between input and output trajectories of biochemical networks. Phys. Rev. Lett. 102, 218101 (2009).
Long, J., Zucker, S. W. & Emonet, T. Feedback between motion and sensation provides nonlinear boost in run-and-tumble navigation. PLoS Comput. Biol. 13, e1005429 (2017).
Hartich, D., Barato, A. C. & Seifert, U. Stochastic thermodynamics of bipartite systems: transfer entropy inequalities and a Maxwell’s demon interpretation. J. Stat. Mech. 2014, P02016 (2014).
Meijers, M., Ito, S. & ten Wolde, P. R. Behavior of information flow near criticality. Phys. Rev. E 103, L010102 (2021).
Schreiber, T. Measuring information transfer. Phys. Rev. Lett. 85, 461–464 (2000).
Sigtermans, D. Towards a framework for observational causality from time series: when Shannon meets Turing. Entropy 22, 426 (2020).
de Gennes, P.-G. Chemotaxis: the role of internal delays. Eur. Biophys. J. 33, 691–693 (2004).
Clark, D. A. & Grant, L. C. The bacterial chemotactic response reflects a compromise between transient and steady-state behavior. Proc. Natl Acad. Sci. USA 102, 9150–9155 (2005).
Locsei, J. T. Persistence of direction increases the drift velocity of run and tumble chemotaxis. J. Math. Biol. 55, 41–60 (2007).
Celani, A. & Vergassola, M. Bacterial strategies for chemotaxis response. Proc. Natl Acad. Sci. USA 107, 1391–1396 (2010).
Becker, N. B., Mugler, A. & ten Wolde, P. R. Optimal prediction by cellular signaling networks. Phys. Rev. Lett. 115, 258103 (2015).
Nakamura, K. & Kobayashi, T. J. Connection between the bacterial chemotactic network and optimal filtering. Phys. Rev. Lett. 126, 128102 (2021).
Spudich, J. L. & Koshland, D. E. Non-genetic individuality: chance in the single cell. Nature 262, 467–471 (1976).
Masson, J.-B., Voisinne, G., Wong-Ng, J., Celani, A. & Vergassola, M. Noninvasive inference of the molecular chemotactic response using bacterial trajectories. Proc. Natl Acad. Sci. USA 109, 1802–1807 (2012).
Waite, A. J., Frankel, N. W. & Emonet, T. Behavioral variability and phenotypic diversity in bacterial chemotaxis. Annu. Rev. Biophys. 47, 595–616 (2018).
Moore, J. P., Kamino, K. & Emonet, T. Non-genetic diversity in chemosensing and chemotactic behavior. Int. J. Mol. Sci. 22, 6960 (2021).
Shannon, C. E. Communication in the presence of noise. Proc. IRE 37, 10–21 (1949).
Rieke, F., Bodnar, D. A. & Bialek, W. Naturalistic stimuli increase the rate and efficiency of information transmission by primary auditory afferents. Proc. R. Soc. Lond. B 262, 259–265 (1995).
Kamino, K., Keegstra, J. M., Long, J., Emonet, T. & Shimizu, T. S. Adaptive tuning of cell sensory diversity without changes in gene expression. Sci. Adv. 6, eabc1087 (2020).
Kalinin, Y. V., Jiang, L., Tu, Y. & Wu, M. Logarithmic sensing in Escherichia coli bacterial chemotaxis. Biophys. J. 96, 2439–2448 (2009).
Sourjik, V. & Berg, H. C. Receptor sensitivity in bacterial chemotaxis. Proc. Natl Acad. Sci. USA 99, 123–127 (2002).
Shimizu, T. S., Tu, Y. & Berg, H. C. A modular gradient-sensing network for chemotaxis in Escherichia coli revealed by responses to time-varying stimuli. Mol. Syst. Biol. 6, 382 (2010).
Francis, N. R. et al. Subunit organization in a soluble complex of Tar, CheW, and CheA by electron microscopy. J. Biol. Chem. 277, 36755–36759 (2002).
Levit, M. N., Grebe, T. W. & Stock, J. B. Organization of the receptor-kinase signaling array that regulates Escherichia coli chemotaxis. J. Biol. Chem. 277, 36748–36754 (2002).
Colin, R., Zhang, R. & Wilson, L. G. Fast, high-throughput measurement of collective behaviour in a bacterial population. J. R. Soc. Interface 11, 20140486 (2014).
Potter, G. D., Byrd, T. A., Mugler, A. & Sun, B. Dynamic sampling and information encoding in biochemical networks. Biophys. J. 112, 795–804 (2017).
Anders, A., Ghosh, B., Glatter, T. & Sourjik, V. Design of a MAPK signalling cascade balances energetic cost versus accuracy of information transmission. Nat. Commun. 11, 3494 (2020).
Razo-Mejia, M. et al. First-principles prediction of the information processing capacity of a simple genetic circuit. Phys. Rev. E 102, 022404 (2020).
Shannon, C. E. Coding theorems for a discrete source with a fidelity criterion. IRE Nat. Conv. Rec. 7, 142–163 (1959).
Andrews, B. W. & Iglesias, P. A. An information-theoretic characterization of the optimal gradient sensing response of cells. PLoS Comput. Biol. 3, e153 (2007).
Mora, T. & Wingreen, N. S. Limits of sensing temporal concentration changes by single cells. Phys. Rev. Lett. 104, 248101 (2010).
Waite, A. J. et al. Non-genetic diversity modulates population performance. Mol. Syst. Biol. 12, 895 (2016).
Ni, B., Colin, R., Link, H., Endres, R. G. & Sourjik, V. Growth-rate dependent resource investment in bacterial motile behavior quantitatively follows potential benefit of chemotaxis. Proc. Natl Acad. Sci. USA 117, 595–601 (2020).
Berg, H. C. & Brown, D. A. Chemotaxis in Escherichia coli analysed by three-dimensional tracking. Nature 239, 500–504 (1972).
Dufour, Y. S., Fu, X., Hernandez-Nunez, L. & Emonet, T. Limits of feedback control in bacterial chemotaxis. PLoS Comput. Biol. 10, e1003694 (2014).
Armstrong, J. B., Adler, J. & Dahl, M. M. Nonchemotactic mutants of Escherichia coli. J. Bacteriol. 93, 390–398 (1967).
Bachmann, B. J. Pedigrees of some mutant strains of Escherichia coli K-12. Bacteriol. Rev. 36, 525–557 (1972).
Barker, C. S., Prüß, B. M. & Matsumura, P. Increased motility of Escherichia coli by insertion sequence element integration into the regulatory region of the flhD operon. J. Bacteriol. 186, 7529–7537 (2004).
Wolfe, A. J. & Berg, H. C. Migration of bacteria in semisolid agar. Proc. Natl Acad. Sci. USA 86, 6973–6977 (1989).
Licata, N. A., Mohari, B., Fuqua, C. & Setayeshgar, S. Diffusion of bacterial cells in porous media. Biophys. J. 110, 247–257 (2016).
Kurzthaler, C. et al. A geometric criterion for the optimal spreading of active polymers in porous media. Preprint at https://arxiv.org/abs/2106.05366 (2021).
Alon, U., Surette, M. G., Barkai, N. & Leibler, S. Robustness in bacterial chemotaxis. Nature 397, 168–171 (1999).
Neumann, S., Vladimirov, N., Krembel, A. K., Wingreen, N. S. & Sourjik, V. Imprecision of adaptation in Escherichia coli chemotaxis. PLoS ONE 9, e84904 (2014).
Wong-Ng, J., Melbinger, A., Celani, A. & Vergassola, M. The role of adaptation in bacterial speed races. PLoS Comput. Biol. 12, e1004974 (2016).
Wong-Ng, J., Celani, A. & Vergassola, M. Exploring the function of bacterial chemotaxis. Curr. Opin. Microbiol. 45, 16–21 (2018).
Tu, Y. & Grinstein, G. How white noise generates power-law switching in bacterial flagellar motors. Phys. Rev. Lett. 94, 208101 (2005).
Park, H., Oikonomou, P., Guet, C. C. & Cluzel, P. Noise underlies switching behavior of the bacterial flagellum. Biophys. J. 101, 2336–2340 (2011).
Huo, H., He, R., Zhang, R. & Yuan, J. Swimming Escherichia coli cells explore the environment by Lévy walk. Appl. Environ. Microbiol. 87, e02429-20 (2021).
Flores, M., Shimizu, T. S., ten Wolde, P. R. & Tostevin, F. Signaling noise enhances chemotactic drift of E. coli. Phys. Rev. Lett. 109, 148101 (2012).
Sneddon, M. W., Pontius, W. & Emonet, T. Stochastic coordination of multiple actuators reduces latency and improves chemotactic response in bacteria. Proc. Natl Acad. Sci. USA 109, 805–810 (2012).
Qin, D., Xia, Y. & Whitesides, G. M. Soft lithography for micro- and nanoscale patterning. Nat. Protoc. 5, 491–502 (2010).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using µManager. Curr. Protoc. Mol. Biol. 92, 14.20.1–14.20.17 (2010).
Sourjik, V. & Berg, H. C. Functional interactions between receptors in bacterial chemotaxis. Nature 428, 437–441 (2004).
Frymier, P. D., Ford, R. M., Berg, H. C. & Cummings, P. T. Three-dimensional tracking of motile bacteria near a solid planar surface. Proc. Natl Acad. Sci. USA 92, 6195–6199 (1995).
Grognot, M. & Taute, K. M. A multiscale 3D chemotaxis assay reveals bacterial navigation mechanisms. Commun. Biol. 4, 1–8 (2021).
Acknowledgements
We thank J. Moore and X. Zhang for their help in setting up the experimental assays. We also thank K. Taute and M. Grognot for helpful discussions about the gradient experiments. We thank P. R. ten Wolde for providing detailed feedback on an earlier version of this manuscript; I. Nemenman and S. Ito for helpful discussions; and A. Wachtel, I. Graf, and D. Clark for commenting on the text. We acknowledge T. Shimizu and V. Sourjik for bacteria strains and R. Gomez-Sjoberg, Microfluidics Lab, for providing information and software to control the solenoid valves in the microfluidic setup. This work was supported by NIH awards R01GM106189 (H.H.M., K.K. and T.E.), R01GM138533 (H.H.M., K.K. and T.E.), F32GM131583 (H.H.M.) and R35GM138341 (B.B.M.); by a Yale PEB Seed Grant (T.E. and B.B.M.); and by Simons Investigator Award 624156 (B.B.M.).
Author information
Authors and Affiliations
Contributions
H.H.M. and K.K. contributed equally to this work. H.H.M., K.K., B.B.M. and T.E. designed the research. H.H.M. and B.B.M. derived the theoretical bound with inputs from T.E. and K.K. H.H.M. performed the bacteria-tracking experiments. K.K. performed the single-cell FRET experiments. H.H.M., K.K. and T.E. validated the data. H.H.M., K.K., B.B.M. and T.E. discussed the data analysis. H.H.M. and K.K. performed the data analysis. H.H.M., K.K., B.B.M. and T.E. wrote the initial draft and all the revisions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Physics thanks Lev Tsimring and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, Table 1 and text.
Supplementary Data
Source data plotted in the Supplementary figures, as well as a README file.
Source data
Source Data Fig. 1
Source data and code for generating the figure, as well as a README file.
Source Data Fig. 2
Source data and code for generating the figure, as well as a README file.
Source Data Fig. 3
Source data and code for generating the figure, as well as a README file.
Rights and permissions
About this article
Cite this article
Mattingly, H.H., Kamino, K., Machta, B.B. et al. Escherichia coli chemotaxis is information limited. Nat. Phys. 17, 1426–1431 (2021). https://doi.org/10.1038/s41567-021-01380-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-021-01380-3
This article is cited by
-
Generative learning for nonlinear dynamics
Nature Reviews Physics (2024)
-
Microbes in porous environments: from active interactions to emergent feedback
Biophysical Reviews (2024)
-
Smart micro- and nanorobots for water purification
Nature Reviews Bioengineering (2023)
-
The ecological roles of bacterial chemotaxis
Nature Reviews Microbiology (2022)