Abstract
The field of single-cell genomics and spatial technologies is rapidly evolving and has already provided unprecedented insights into complex tissues. Major advances have been made in dissecting the cellular composition and spatiotemporal interactions that mediate developmental processes in the fetal kidney. Single-cell technologies have also provided detailed insights into the heterogeneity of cell types within the healthy adult and shed light on the complex cellular mechanisms that contribute to kidney disease. The in-depth characterization of specific cell types associated with acute kidney injury and glomerular diseases has potential for the development of prognostic biomarkers and new therapeutics. Analyses of pathway activity in clear-cell renal cell carcinoma can predict the sensitivity of tumour cells to specific inhibitors. The identification of the cell of origin of renal cell carcinoma and of new cell types within the tumour microenvironment also has implications for the development of targeted therapeutics. Similarly, single-cell sequencing has provided new insights into the mechanisms underlying kidney fibrosis, specifically our understanding of myofibroblast origins and the contribution of cell crosstalk within the fibrotic niche to disease progression. These and future studies will enable the creation of a map to aid our understanding of the cellular processes and interactions in the developing, healthy and diseased kidney.
Key points
-
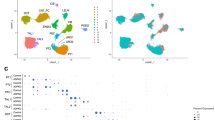
Single-cell RNA sequencing has enabled dissection of the cellular heterogeneity of complex tissues, as well as the characterization of rare cell populations.
-
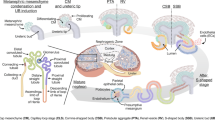
Differential gene expression analyses and pseudotemporal predictions have yielded insights into the mechanisms of mesenchymal-to-epithelial transition, nephron progenitor self-renewal and podocyte development.
-
Inference of pathway activity for druggable pathways in clear-cell renal cell carcinoma can predict the sensitivity of tumour cells to pathway inhibitors, which could facilitate the identification of optimal combinational therapy.
-
Investigation of fibrosis-related gene expression in single-cell RNA sequencing data has enabled precise understanding of fibroblast and pericyte-to-myofibroblast differentiation trajectories.
-
Single-cell RNA sequencing has facilitated characterization of a dedifferentiated VCAM1+ population of proximal tubule cells and revealed its broad relevance in renal cell carcinoma, kidney injury and kidney fibrosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl Acad. Sci. USA 89, 1827–1831 (1992).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 39, 313–319 (2021).
Takasato, M. & Little, M. H. The origin of the mammalian kidney: implications for recreating the kidney in vitro. Development 142, 1937–1947 (2015).
Rosenblum, N. D. Developmental biology of the human kidney. Semin. Fetal Neonatal Med. 13, 125–132 (2008).
Lindström, N. O. et al. Progressive recruitment of mesenchymal progenitors reveals a time-dependent process of cell fate acquisition in mouse and human nephrogenesis. Dev. Cell 45, 651–660.e4 (2018).
Matsui, I. et al. Single cell RNA sequencing uncovers cellular developmental sequences and novel potential intercellular communications in embryonic kidney. Sci. Rep. 11, 73 (2021).
Hochane, M. et al. Single-cell transcriptomics reveals gene expression dynamics of human fetal kidney development. PLoS Biol. 17, e3000152 (2019).
Lawlor, K. T. et al. Nephron progenitor commitment is a stochastic process influenced by cell migration. Elife 8, e41156 (2019).
Brunskill, E. W. et al. Single cell dissection of early kidney development: multilineage priming. Development 141, 3093–3101 (2014).
Magella, B. et al. Cross-platform single cell analysis of kidney development shows stromal cells express Gdnf. Dev. Biol. 434, 36–47 (2018).
Lindström, N. O. et al. Spatial transcriptional mapping of the human nephrogenic program. Dev. Cell 56, 2381–2398.e6 (2021).
Wineberg, Y. et al. Single-cell RNA sequencing reveals mRNA splice isoform switching during kidney development. J. Am. Soc. Nephrol. 31, 2278–2291 (2020).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Van Itallie, C. M. et al. Two splice variants of claudin-10 in the kidney create paracellular pores with different ion selectivities. Am. J. Physiol. Renal Physiol. 291, F1288–F1299 (2006).
Tran, T. et al. In vivo developmental trajectories of human podocyte inform in vitro differentiation of pluripotent stem cell-derived podocytes. Dev. Cell 50, 102–116.e6 (2019).
Menon, R. et al. Single-cell analysis of progenitor cell dynamics and lineage specification in the human fetal kidney. Development 145, dev164038 (2018).
Yamada, H. et al. MAGI-2 orchestrates the localization of backbone proteins in the slit diaphragm of podocytes. Kidney Int. 99, 382–395 (2021).
Tsujimoto, H. et al. A modular differentiation system maps multiple human kidney lineages from pluripotent stem cells. Cell Rep. 31, 107476 (2020).
Subramanian, A. et al. Single cell census of human kidney organoids shows reproducibility and diminished off-target cells after transplantation. Nat. Commun. 10, 5462 (2019).
Low, J. H. et al. Generation of human PSC-derived kidney organoids with patterned nephron segments and a de novo vascular network. Cell Stem Cell 25, 373–387.e9 (2019).
Combes, A. N., Zappia, L., Er, P. X., Oshlack, A. & Little, M. H. Single-cell analysis reveals congruence between kidney organoids and human fetal kidney. Genome Med. 11, 3 (2019).
Phipson, B. et al. Evaluation of variability in human kidney organoids. Nat. Methods 16, 79–87 (2019).
Shankar, A. S. et al. Human kidney organoids produce functional renin. Kidney Int. 99, 134–147 (2021).
Wu, H. & Humphreys, B. D. Single cell sequencing and kidney organoids generated from pluripotent stem cells. Clin. J. Am. Soc. Nephrol. 15, 550–556 (2020).
Lähnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Dumas, S. J. et al. Phenotypic diversity and metabolic specialization of renal endothelial cells. Nat. Rev. Nephrol. 17, 441–464 (2021).
Stewart, B. J., Ferdinand, J. R. & Clatworthy, M. R. Using single-cell technologies to map the human immune system — implications for nephrology. Nat. Rev. Nephrol. 16, 112–128 (2020).
Wu, H., Kirita, Y., Donnelly, E. L. & Humphreys, B. D. Advantages of single-nucleus over single-cell RNA sequencing of adult kidney: rare cell types and novel cell states revealed in fibrosis. J. Am. Soc. Nephrol. 30, 23–32 (2019).
He, B. et al. Single-cell RNA sequencing reveals the mesangial identity and species diversity of glomerular cell transcriptomes. Nat. Commun. 12, 2141 (2021).
Lu, Y., Ye, Y., Yang, Q. & Shi, S. Single-cell RNA-sequence analysis of mouse glomerular mesangial cells uncovers mesangial cell essential genes. Kidney Int. 92, 504–513 (2017).
Karaiskos, N. et al. A single-cell transcriptome atlas of the mouse glomerulus. J. Am. Soc. Nephrol. 29, 2060–2068 (2018).
Chung, J.-J. et al. Single-cell transcriptome profiling of the kidney glomerulus identifies key cell types and reactions to injury. J. Am. Soc. Nephrol. 31, 2341–2354 (2020).
Kuppe, C. et al. Decoding myofibroblast origins in human kidney fibrosis. Nature 589, 281–286 (2021).
Menon, R. et al. Single cell transcriptomics identifies focal segmental glomerulosclerosis remission endothelial biomarker. JCI Insight 5, e133267 (2020).
Muto, Y. et al. Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney. Nat. Commun. 12, 2190 (2021).
Ransick, A. et al. Single-cell profiling reveals sex, lineage, and regional diversity in the mouse kidney. Dev. Cell 51, 399–413.e7 (2019).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Lake, B. B. et al. A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys. Nat. Commun. 10, 2832 (2019).
Dhillon, P. et al. The nuclear receptor ESRRA protects from kidney disease by coupling metabolism and differentiation. Cell Metab. 33, 379–394.e8 (2021).
Kirita, Y., Wu, H., Uchimura, K., Wilson, P. C. & Humphreys, B. D. Cell profiling of mouse acute kidney injury reveals conserved cellular responses to injury. Proc. Natl Acad. Sci. USA 117, 15874–15883 (2020).
Chen, L., Chou, C.-L. & Knepper, M. A. Targeted single-cell RNA-seq identifies minority cell types of kidney distal nephron. J. Am. Soc. Nephrol. https://doi.org/10.1681/ASN.2020101407 (2021).
Kompatscher, A. et al. Loss of transcriptional activation of the potassium channel Kir5.1 by HNF1β drives autosomal dominant tubulointerstitial kidney disease. Kidney Int. 92, 1145–1156 (2017).
Kompatscher, A. et al. Transcription factor HNF1β regulates expression of the calcium-sensing receptor in the thick ascending limb of the kidney. Am. J. Physiol. Renal Physiol. 315, F27–F35 (2018).
Cavodeassi, F., Modolell, J. & Gómez-Skarmeta, J. L. The Iroquois family of genes: from body building to neural patterning. Development 128, 2847–2855 (2001).
Reggiani, L., Raciti, D., Airik, R., Kispert, A. & Brändli, A. W. The prepattern transcription factor Irx3 directs nephron segment identity. Genes Dev. 21, 2358–2370 (2007).
Chen, L. et al. Renal-tubule epithelial cell nomenclature for single-cell RNA-sequencing studies. J. Am. Soc. Nephrol. 30, 1358–1364 (2019).
Madsen, K. M. & Tisher, C. C. Structural-functional relationship along the distal nephron. Am. J. Physiol. 250, F1–F15 (1986).
Chen, L. et al. Transcriptomes of major renal collecting duct cell types in mouse identified by single-cell RNA-seq. Proc. Natl Acad. Sci. USA 114, E9989–E9998 (2017).
Hinze, C. et al. Kidney single-cell transcriptomes predict spatial corticomedullary gene expression and tissue osmolality gradients. J. Am. Soc. Nephrol. 32, 291–306 (2021).
Saxena, V. et al. Kidney intercalated cells are phagocytic and acidify internalized uropathogenic Escherichia coli. Nat. Commun. 12, 2405 (2021).
Kim, J., Kim, Y. H., Cha, J. H., Tisher, C. C. & Madsen, K. M. Intercalated cell subtypes in connecting tubule and cortical collecting duct of rat and mouse. J. Am. Soc. Nephrol. 10, 1–12 (1999).
Werth, M. et al. Transcription factor patterns cells in the mouse kidney collecting ducts. Elife 6, e24265 (2017).
Trepiccione, F., Capasso, G., Nielsen, S. & Christensen, B. M. Evaluation of cellular plasticity in the collecting duct during recovery from lithium-induced nephrogenic diabetes insipidus. Am. J. Physiol. Renal Physiol. 305, F919–F929 (2013).
Jamous, M. et al. In young primary cultures of rabbit kidney cortical collecting ducts intercalated cells originate from principal or undifferentiated cells. Eur. J. Cell Biol. 66, 192–199 (1995).
Wu, H. et al. Aqp2-expressing cells give rise to renal intercalated cells. J. Am. Soc. Nephrol. 24, 243–252 (2013).
Fejes-Tóth, G. & Náray-Fejes-Tóth, A. Differentiation of renal beta-intercalated cells to alpha-intercalated and principal cells in culture. Proc. Natl Acad. Sci. USA 89, 5487–5491 (1992).
Gao, X. et al. Deletion of hensin/DMBT1 blocks conversion of beta- to alpha-intercalated cells and induces distal renal tubular acidosis. Proc. Natl Acad. Sci. USA 107, 21872–21877 (2010).
Padala, S. A. et al. Epidemiology of renal cell carcinoma. World J. Oncol. 11, 79–87 (2020).
Zhang, Y. et al. Single-cell analyses of renal cell cancers reveal insights into tumor microenvironment, cell of origin, and therapy response. Proc. Natl Acad. Sci. USA 118, e2103240118 (2021).
Peired, A. J. et al. Acute kidney injury promotes development of papillary renal cell adenoma and carcinoma from renal progenitor cells. Sci. Transl. Med. 12, eaaw6003 (2020).
Lombardi, D., Becherucci, F. & Romagnani, P. How much can the tubule regenerate and who does it? An open question. Nephrol. Dial. Transpl. 31, 1243–1250 (2016).
Zhang, S. et al. Immune infiltration in renal cell carcinoma. Cancer Sci. 110, 1564–1572 (2019).
Wu, T. D. et al. Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature 579, 274–278 (2020).
Borcherding, N. et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun. Biol. 4, 122 (2021).
Hu, J. et al. Single-cell transcriptome analysis reveals intratumoral heterogeneity in ccRCC, which results in different clinical outcomes. Mol. Ther. 28, 1658–1672 (2020).
Iwai, Y. et al. Involvement of PD-L1 on tumor cells in the escape from host immune system and tumor immunotherapy by PD-L1 blockade. Proc. Natl Acad. Sci. USA 99, 12293–12297 (2002).
Pruenster, M. & Rot, A. Throwing light on DARC. Biochem. Soc. Trans. 34, 1005–1008 (2006).
Courtney, K. D. & Choueiri, T. K. Updates on novel therapies for metastatic renal cell carcinoma. Ther. Adv. Med. Oncol. 2, 209–219 (2010).
Kim, K.-T. et al. Application of single-cell RNA sequencing in optimizing a combinatorial therapeutic strategy in metastatic renal cell carcinoma. Genome Biol. 17, 80 (2016).
Li, P. et al. Histopathologic correlates of kidney function: insights from nephrectomy specimens. Am. J. Kidney Dis. 77, 336–345 (2021).
Falke, L. L., Gholizadeh, S., Goldschmeding, R., Kok, R. J. & Nguyen, T. Q. Diverse origins of the myofibroblast — implications for kidney fibrosis. Nat. Rev. Nephrol. 11, 233–244 (2015).
Kramann, R. et al. Perivascular Gli1+ progenitors are key contributors to injury-induced organ fibrosis. Cell Stem Cell 16, 51–66 (2015).
Kramann, R. et al. Parabiosis and single-cell RNA sequencing reveal a limited contribution of monocytes to myofibroblasts in kidney fibrosis. JCI Insight 3, e99561 (2018).
Chevalier, R. L. The proximal tubule is the primary target of injury and progression of kidney disease: role of the glomerulotubular junction. Am. J. Physiol. Renal Physiol. 311, F145–F161 (2016).
Conway, B. R. et al. Kidney single-cell atlas reveals myeloid heterogeneity in progression and regression of kidney disease. J. Am. Soc. Nephrol. 31, 2833–2854 (2020).
Rudman-Melnick, V. et al. Single-cell profiling of AKI in a murine model reveals novel transcriptional signatures, profibrotic phenotype, and epithelial-to-stromal crosstalk. J. Am. Soc. Nephrol. 31, 2793–2814 (2020).
Wilson, P. C. et al. The single-cell transcriptomic landscape of early human diabetic nephropathy. Proc. Natl Acad. Sci. USA 116, 19619–19625 (2019).
Lake, B. B. et al. An atlas of healthy and injured cell states and niches in the human kidney. bioRxiv https://doi.org/10.1101/2021.07.28.454201 (2021).
Abedini, A. et al. Urinary single-cell profiling captures the cellular diversity of the kidney. J. Am. Soc. Nephrol. 32, 614–627 (2021).
Lusco, M. A., Najafian, B., Alpers, C. E. & Fogo, A. B. AJKD atlas of renal pathology: Pierson syndrome. Am. J. Kidney Dis. 71, e3–e4 (2018).
Zhang, L. et al. Genetic and preimplantation diagnosis of cystic kidney disease with ventriculomegaly. J. Hum. Genet. 65, 455–459 (2020).
Arazi, A. et al. Publisher correction: the immune cell landscape in kidneys of patients with lupus nephritis. Nat. Immunol. 20, 1404 (2019).
Zhang, T. et al. Association of urine sCD163 with proliferative lupus nephritis, fibrinoid necrosis, cellular crescents and intrarenal M2 macrophages. Front. Immunol. 11, 671 (2020).
Fava, A. et al. Integrated urine proteomics and renal single-cell genomics identify an IFN-γ response gradient in lupus nephritis. JCI Insight 5, e138345 (2020).
Der, E. et al. Tubular cell and keratinocyte single-cell transcriptomics applied to lupus nephritis reveal type I IFN and fibrosis relevant pathways. Nat. Immunol. 20, 915–927 (2019).
Der, E. et al. Single cell RNA sequencing to dissect the molecular heterogeneity in lupus nephritis. JCI Insight 2, e93009 (2017).
Zheng, Y. et al. Single-cell transcriptomics reveal immune mechanisms of the onset and progression of IgA nephropathy. Cell Rep. 33, 108525 (2020).
Tang, R. et al. A partial picture of the single-cell transcriptomics of human IgA nephropathy. Front. Immunol. 12, 645988 (2021).
Fu, J. et al. Single-cell RNA profiling of glomerular cells shows dynamic changes in experimental diabetic kidney disease. J. Am. Soc. Nephrol. 30, 533–545 (2019).
Sheng, X. et al. Mapping the genetic architecture of human traits to cell types in the kidney identifies mechanisms of disease and potential treatments. Nat. Genet. https://doi.org/10.1038/s41588-021-00909-9 (2021).
Doke, T. et al. Transcriptome-wide association analysis identifies DACH1 as a kidney disease risk gene that contributes to fibrosis. J. Clin. Invest 131, e141801 (2021).
Miao, Z. et al. Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets. Nat. Commun. 12, 2277 (2021).
Marshall, J. L. et al. High resolution Slide-seqV2 spatial transcriptomics enables discovery of disease-specific cell neighborhoods and pathways. bioRxiv https://doi.org/10.1101/2021.10.10.463829 (2021).
Author information
Authors and Affiliations
Contributions
Both authors researched data for the article, contributed substantially to discussion of the content and wrote the article. R.K. reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Nephrology thanks David Ferenbach, Benjamin Stewart and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- RNA-velocity
-
A computational method that predicts future states of cells by comparing the quantitative abundance of spliced and unspliced mRNA of genes.
- Pseudotime analysis
-
A computational method that orders single cells along a developmental trajectory by identifying similarities and continuous changes in the transcriptome of these cells.
- Alternative splicing
-
A process, in which different mRNA molecules are generated from the same gene by removing different parts of the gene during transcription. This process enables different proteins to be encoded by the same gene.
- Isoform switching
-
The process by which a cell switches from producing one isoform of a transcript to another isoform.
Rights and permissions
About this article
Cite this article
Schreibing, F., Kramann, R. Mapping the human kidney using single-cell genomics. Nat Rev Nephrol 18, 347–360 (2022). https://doi.org/10.1038/s41581-022-00553-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41581-022-00553-4
This article is cited by
-
TRPM channels in health and disease
Nature Reviews Nephrology (2024)
-
Mapping human tissues with highly multiplexed RNA in situ hybridization
Nature Communications (2024)
-
Pathological consequences of DNA damage in the kidney
Nature Reviews Nephrology (2023)
-
Single-cell lipidomics enabled by dual-polarity ionization and ion mobility-mass spectrometry imaging
Nature Communications (2023)
-
Single-cell transcriptomics in tissue engineering and regenerative medicine
Nature Reviews Bioengineering (2023)