Abstract
The RAG endonuclease initiates Igh locus V(D)J recombination in progenitor (pro)-B cells1. Upon binding a recombination centre-based JH, RAG scans upstream chromatin via loop extrusion, potentially mediated by cohesin, to locate Ds and assemble a DJH-based recombination centre2. CTCF looping factor-bound elements (CBEs) within IGCR1 upstream of Ds impede RAG scanning3,4,5; however, their inactivation allows scanning to proximal VHs, where additional CBEs activate rearrangement and impede scanning any further upstream5. Distal VH utilization is thought to involve diffusional access to the recombination centre following large-scale Igh locus contraction6,7,8. Here we test the potential of linear RAG scanning to mediate distal VH usage in G1-arrested v-Abl pro-B cell lines9, which undergo robust D-to-JH but little VH-to-DJH rearrangements, presumably owing to lack of locus contraction2,5. Through an auxin-inducible approach10, we degraded the cohesin component RAD2110,11,12 or CTCF12,13 in these G1-arrested lines. Degradation of RAD21 eliminated all V(D)J recombination and interactions associated with RAG scanning, except for reecombination centre-located DQ52-to-JH joining, in which synapsis occurs by diffusion2. Remarkably, while degradation of CTCF suppressed most CBE-based chromatin interactions, it promoted robust recombination centre interactions with, and robust VH-to-DJH joining of, distal VHs, with patterns similar to those of ‘locus-contracted’ primary pro-B cells. Thus, downmodulation of CTCF-bound scanning-impediment activity promotes cohesin-driven RAG scanning across the 2.7-Mb Igh locus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
HTGTS-V(D)J-seq, 3C-HTGTS, ChIP–seq and GRO-seq data reported in this study have been deposited in the Gene Expression Omnibus (GEO) database under the accession number GSE142781. Source data are provided with this paper.
Code availability
HTGTS-V(D)J-seq and 3C-HTGTS data were processed via the published pipeline (http://robinmeyers.github.io/transloc_pipeline/). D usage extraction from VHDJH joins was processed via a custom VDJ_annotation pipeline (https://github.com/Yyx2626/VDJ_annotation/).
References
Teng, G. & Schatz, D. G. Regulation and evolution of the RAG recombinase. Adv. Immunol. 128, 1–39 (2015).
Zhang, Y. et al. The fundamental role of chromatin loop extrusion in physiological V(D)J recombination. Nature 573, 600–604 (2019).
Guo, C. et al. CTCF-binding elements mediate control of V(D)J recombination. Nature 477, 424–430 (2011).
Hu, J. et al. Chromosomal loop domains direct the recombination of antigen receptor genes. Cell 163, 947–959 (2015).
Jain, S., Ba, Z., Zhang, Y., Dai, H.-Q. & Alt, F. W. CTCF-binding elements mediate accessibility of RAG substrates during chromatin scanning. Cell 174, 102–116.e14 (2018).
Bossen, C., Mansson, R. & Murre, C. Chromatin topology and the regulation of antigen receptor assembly. Annu. Rev. Immunol. 30, 337–356 (2012).
Ebert, A., Hill, L. & Busslinger, M. Spatial regulation of V-(D)J recombination at antigen receptor loci. Adv. Immunol. 128, 93–121 (2015).
Proudhon, C., Hao, B., Raviram, R., Chaumeil, J. & Skok, J. A. Long-range regulation of V(D)J recombination. Adv. Immunol. 128, 123–182 (2015).
Bredemeyer, A. L. et al. ATM stabilizes DNA double-strand-break complexes during V(D)J recombination. Nature 442, 466–470 (2006).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944.e22 (2017).
Yatskevich, S., Rhodes, J. & Nasmyth, K. Organization of chromosomal DNA by SMC complexes. Annu. Rev. Genet. 53, 445–482 (2019).
Peters, J.-M., Tedeschi, A. & Schmitz, J. The cohesin complex and its roles in chromosome biology. Genes Dev. 22, 3089–3114 (2008).
Haarhuis, J. H. I., Elbatsh, A. M. O. & Rowland, B. D. Cohesin and its regulation: on the logic of X-shaped chromosomes. Dev. Cell 31, 7–18 (2014).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Zhang, X. et al. Fundamental roles of chromatin loop extrusion in antibody class switching. Nature 575, 385–389 (2019).
Lin, S. G. et al. Highly sensitive and unbiased approach for elucidating antibody repertoires. Proc. Natl Acad. Sci. USA 113, 7846–7851 (2016).
Choi, N. M. et al. Deep sequencing of the murine IgH repertoire reveals complex regulation of nonrandom V gene rearrangement frequencies. J. Immunol. 191, 2393–2402 (2013).
Bolland, D. J. et al. Two mutually exclusive local chromatin states drive efficient V(D)J recombination. Cell Rep. 15, 2475–2487 (2016).
Degner, S. C. et al. CCCTC-binding factor (CTCF) and cohesin influence the genomic architecture of the Igh locus and antisense transcription in pro-B cells. Proc. Natl Acad. Sci. USA 108, 9566–9571 (2011).
Lucas, J. S., Zhang, Y., Dudko, O. K. & Murre, C. 3D trajectories adopted by coding and regulatory DNA elements: first-passage times for genomic interactions. Cell 158, 339–352 (2014).
Khoury, A. et al. Constitutively bound CTCF sites maintain 3D chromatin architecture and long-range epigenetically regulated domains. Nat. Commun. 11, 54 (2020).
Nakahashi, H. et al. A genome-wide map of CTCF multivalency redefines the CTCF code. Cell Rep. 3, 1678–1689 (2013).
Canzio, D. et al. Antisense lncRNA transcription mediates DNA demethylation to drive stochastic protocadherin α promoter choice. Cell 177, 639–653.e15 (2019).
Hansen, A. S., Cattoglio, C., Darzacq, X. & Tjian, R. Recent evidence that TADs and chromatin loops are dynamic structures. Nucleus 9, 20–32 (2018).
Li, Y. et al. The structural basis for cohesin–CTCF-anchored loops. Nature 578, 472–476 (2020).
Pugacheva, E. M. et al. CTCF mediates chromatin looping via N-terminal domain-dependent cohesin retention. Proc. Natl Acad. Sci. USA 117, 2020–2031 (2020).
Hansen, A. S. et al. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell 76, 395–411.e13 (2019).
Saldaña-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422.e5 (2019).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707.e14 (2017).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Wutz, G. et al. ESCO1 and CTCF enable formation of long chromatin loops by protecting cohesinSTAG1 from WAPL. eLife 9, e52091 (2020).
Hesslein, D. G. T. et al. Pax5 is required for recombination of transcribed, acetylated, 5' IgH V gene segments. Genes Dev. 17, 37–42 (2003).
Fuxa, M. et al. Pax5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes Dev. 18, 411–422 (2004).
Liu, H. et al. Yin Yang 1 is a critical regulator of B-cell development. Genes Dev. 21, 1179–1189 (2007).
Medvedovic, J. et al. Flexible long-range loops in the VH gene region of the Igh locus facilitate the generation of a diverse antibody repertoire. Immunity 39, 229–244 (2013).
Donohoe, M. E., Zhang, L.-F., Xu, N., Shi, Y. & Lee, J. T. Identification of a Ctcf cofactor, Yy1, for the X chromosome binary switch. Mol. Cell 25, 43–56 (2007).
Ebert, A. et al. The distal V(H) gene cluster of the Igh locus contains distinct regulatory elements with Pax5 transcription factor-dependent activity in pro-B cells. Immunity 34, 175–187 (2011).
Seitan, V. C. et al. A role for cohesin in T-cell-receptor rearrangement and thymocyte differentiation. Nature 476, 467–471 (2011).
Zhao, L. et al. Orientation-specific RAG activity in chromosomal loop domains contributes to Tcrd V(D)J recombination during T cell development. J. Exp. Med. 213, 1921–1936 (2016).
Ribeiro de Almeida, C. et al. The DNA-binding protein CTCF limits proximal Vκ recombination and restricts κ enhancer interactions to the immunoglobulin κ light chain locus. Immunity 35, 501–513 (2011).
Lin, S. G., Ba, Z., Alt, F. W. & Zhang, Y. RAG chromatin scanning during V(D)J recombination and chromatin loop extrusion are related processes. Adv. Immunol. 139, 93–135 (2018).
Hsieh, T.-H. S. et al. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell 78, 539–553.e8 (2020).
Alt, F. W., Zhang, Y., Meng, F.-L., Guo, C. & Schwer, B. Mechanisms of programmed DNA lesions and genomic instability in the immune system. Cell 152, 417–429 (2013).
Shinkai, Y. et al. RAG-2-deficient mice lack mature lymphocytes owing to inability to initiate V(D)J rearrangement. Cell 68, 855–867 (1992).
Strasser, A. et al. Enforced BCL2 expression in B-lymphoid cells prolongs antibody responses and elicits autoimmune disease. Proc. Natl Acad. Sci. USA 88, 8661–8665 (1991).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132, 487–498 (2008).
Ji, Y. et al. The in vivo pattern of binding of RAG1 and RAG2 to antigen receptor loci. Cell 141, 419–431 (2010).
Hu, J. et al. Detecting DNA double-stranded breaks in mammalian genomes by linear amplification-mediated high-throughput genome-wide translocation sequencing. Nat. Protoc. 11, 853–871 (2016).
Frock, R. L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33, 179–186 (2015).
Orlando, D. A. et al. Quantitative ChIP-seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Acknowledgements
We thank lab members for stimulating discussions. This work was supported by NIH R01 AI020047 (to F.W.A.). R.C. is partially funded by the NIH Regulome Project. F.W.A. is an investigator of the Howard Hughes Medical Institute. Z.B. was and H.-Q.D. is a Cancer Research Institute Irvington fellow.
Author information
Authors and Affiliations
Contributions
Z.B., J.L., R.C. and F.W.A. designed the study. Z.B. and J.L. performed most of the experiments with assistance from E.W.D., S.G.L. and K.-R.K.-K. on certain experiments. A.Y.Y. and N.K. designed some of the bioinformatics pipelines for data analysis. H.-Q.D. and S.J. provided insights. Z.B., J.L., and F.W.A. analysed and interpreted data, designed figures and wrote the paper. R.C., A.Y.Y., S.G.L., H.-Q.D. and S.J. helped edit the paper. F.W.A. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
F.W.A. is a co-founder of Otoro Biopharmaceuticals.
Additional information
Peer review information Nature thanks David Schatz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation and characterization of Rad21-degron lines.
a, Diagram of the Rad21-degron. b, c, Left: schematic of the targeting strategy for introducing in-frame mAID-Clover sequences into both mouse endogenous Rad21 alleles (b), and OsTir1-V5 expression cassette into both Rosa26 alleles (c). Positions of Cas9/sgRNAs and southern blotting probes are indicated. B: BglII; S: SspI. Right: Southern blotting confirmation of correctly targeted alleles as indicated (n=3 biologically independent repeats with similar results for each targeting). d, Representative Clover signal-based flow-cytometry plots for Rad21-degron cells that are non-treated (NT) or treated with IAA (+IAA) for 6h (top) followed by treatment with STI-571 (without or with IAA) for 4 days to induce G1 arrest (bottom). The parental cells were used as Clover negative controls. Three biological repeats with similar results. e, Time-course cell viability assay for Rad21-degron cells without (NT) or with (+IAA) IAA treatment following STI-571 treatment for G1-arrest (+STI). Average percentage ± s.d. of viable cells for each timepoint and under each condition was shown (n=4 biologically independent samples). p values were calculated using unpaired two-tailed t-test. f, Fucci cell cycle assay of Rad21-degron cells without (NT) or with (+IAA) IAA treatment followed by 4-day STI-571 treatment. Representative flow-cytometry plots and average percentage ± s.d. of cells arrested in G1-stage at indicated condition (n=3 biologically independent samples) are shown. The parental cells without Fucci were used as negative controls. g, Rad21 ChIP-seq profiles at indicated Igh locus in NT and IAA-treated G1-arrested Rad21-degron cells (n=2 biological repeats with similar results). h, Rad21 ChIP-seq signal within ±1kb region across all peaks genome-wide called in G1-arrested NT Rad21-degron cells (n=2 biological repeats with similar results). Top: average enrichment. IAA treatment leads to a depletion of chromatin-bound Rad21 genome-wide. See Methods for details.
Extended Data Fig. 2 Effects of cohesin loss on Igh loop domains and transcription across Igh and genome-wide in G1-arrested Rad21-degron cells.
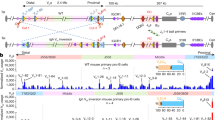
a, b, 3C-HTGTS chromatin interaction profiles of RC/iEμ bait (a, red asterisk) and VH81X-CBE bait (b, blue asterisk) at indicated Igh locus in NT and IAA-treated G1-arrested Rad21-degron cells. Three biologically independent repeats for each bait with similar results. c, Western blotting for Rad21 and RAG2 using histone H3 as a loading control on nuclear extracts of NT and IAA-treated G1-arrested Rad21-degron cells. Two biologically independent repeats with similar results. d, Scatter plots of transcriptome-wide GRO-seq counts in NT (x axis) and IAA-treated (y axis) G1-arrested Rad21-degron cells. Three biologically independent repeats with similar results. Representative known requisite genes for V(D)J recombination and chromatin interaction are highlighted by red circles and blue arrows. Spearman’s correlation coefficient (r) and p values determined by two-sided Spearman’s correlation test are presented. e, GRO-seq analyses of NT and IAA-treated G1-arrested Rad21-degron cells across the entire Igh locus as indicated. Three biologically independent repeats with similar results. S/AS: sense/anti-sense transcription. f, 3C-HTGTS chromatin interaction profiles of RC/iEμ bait (red asterisk) at indicated Igh locus in NT and IAA-treated G1-arrested IGCR1-deleted Rad21-degron cells. Two biologically independent repeats with similar results. See Methods for details.
Extended Data Fig. 3 Generation and characterization of CTCF-tagged, OsTir1-expressing and CTCF-degron lines.
a, Diagram of the CTCF-degron. b, Left: targeting strategy for introducing in-frame mAID-GFP sequences into both Ctcf alleles. Cas9/sgRNAs and southern blotting probe are indicated. B: BglII. Right: Southern blotting confirmation of correctly targeted alleles (n=3 biological repeats with similar results). c, d, Southern blotting confirmation of correctly targeted Rosa26 alleles in CTCF-degron (c, n=3 biological repeats with similar results) and OsTir1-expressing (d, left, n=2 biological repeats with similar results) lines following OsTir1-V5 targeting strategy shown in Extended Data Fig. 1c. Western blotting on nuclear extracts of G1-arresed OsTir1-expressing cells with correctly targeted Rosa26 alleles confirms the constitutive expression of OsTir1-V5 protein (d, right, n=2 biological repeats with similar results). Histone H3 was used as a loading control. e, Representative GFP signal-based flow-cytometry plots for CTCF-degron cells that are non-treated (NT) or treated with IAA (+IAA) for 6h (top) followed by 4-day STI-571 treatment (without or with IAA) (bottom). The parental cells were used as GFP negative controls. Three biological repeats with similar results. f, Fucci cell cycle assay of CTCF-degron cells without (NT) or with IAA treatment followed by 4-day STI-571 treatment. Representative flow-cytometry plots and average percentage ± s.d. of cells arrested in G1-stage at indicated condition (n=3 biologically independent samples) are shown. The parental cells without Fucci were used as negative controls. g, Western blotting on nuclear extracts of G1-arrested cells as indicated using indicated antibodies (n=3 biological repeats with similar results). h, Time-course cell viability assay for parental and IAA-treated CTCF-degron cells, following STI-571 treatment for G1-arrest (+STI). Average percentage ± s.d. of viable cells for each timepoint and under each condition was shown (n=3 biologically independent samples). p values were calculated using unpaired two-tailed t-test. See Methods.
Extended Data Fig. 4 Characterization of chromatin CTCF/Rad21 binding across the Igh downstream part and genome-wide in G1-arrested parental and IAA-treated CTCF-degron cells.
a, b, Average signal counts ± s.e.m. of CTCF (a) and Rad21 (b) ChIP-seq from three biologically independent experiments across 3’ part of Igh locus as diagrammed at the top in G1-arrested parental and IAA-treated CTCF-degron cells are plotted. < and > show orientation of indicated CBEs. c, d, CTCF (c) and Rad21 (d) ChIP-seq signal across all peaks genome-wide called in G1-arrested parental cells. See Extended Data Fig. 1h legend for other details. Three biologically independent repeats with similar results. IAA-treated CTCF-degron cells had greatly diminished CTCF and Rad21 chromatin occupancy, but, also had residual binding at some CBEs genome-wide. See Methods for details. Notably, CTCF binding increased at the RC (Extended Data Fig. 4a) and at a subset transcribed VHs (Fig. 3e, Supplementary Data) upon CTCF depletion. While increases in apparent CTCF-binding to the non-CBE-containing RC and also transcribed VHs upon CTCF depletion could be considered surprising, it may likely occur indirectly due to cohesin-mediated loop extrusion-driven dynamic associations of the RC dynamic anchor with residual-CTCF-bound CBEs across Igh.
Extended Data Fig. 5 Characterization of transcription and RC chromatin interactions across the Igh downstream part and transcription genome-wide in G1-arrested parental and IAA-treated CTCF-degron cells.
a, Average signal counts ± s.e.m. of GRO-seq from three biologically independent experiments across 3’ part of Igh locus as diagramed at the top in G1-arrested parental and IAA-treated CTCF-degron cells are plotted. S/AS: sense/anti-sense transcription. b, Average 3C-HTGTS signal counts ± s.e.m. from three independent experiments with RC-based baits (red asterisk) across 3’ part of Igh locus as diagramed at the top in G1-arrested parental and IAA-treated CTCF-degron cells are plotted. c, Scatter plots of transcriptome-wide GRO-seq counts in G1-arrested parental (x axis) and IAA-treated CTCF-degron (y axis) cells. Three biologically independent repeats with similar results. Representative known requisite genes for V(D)J recombination and chromatin interaction are highlighted by red circles and blue arrows. Spearman’s correlation coefficient (r) and p values determined by two-sided Spearman’s correlation test are presented. d, Scatter plots of normalized GRO-seq read counts (x axis) and 3C-HTGTS junction counts (y axis) calculated from regions within ±5kb of highly-utilized VHs across four indicated VH domains in IAA-treated CTCF-degron cells. Data were normalized and quantified among three biologically independent experiments for each assay. Spearman’s correlation coefficient (r) and p values determined by two-sided Spearman’s correlation test are presented. See Methods for details.
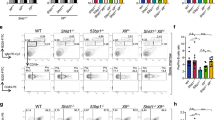
Extended Data Fig. 6 A positive impact of reduced CTCF levels on RAG scanning across the VH locus.
a–h, HTGTS-V(D)J-seq analysis showing average utilization frequencies ± s.d. of VH segments across four highlighted VH domains in RAG2-complemented and G1-arrested parental (a, b), OsTir1-expressing (c, d), CTCF-tagged (e, f), and CTCF-degron (g, h) cells without or with IAA treatment. n=4 (a–f) or n=6 (g, h) biologically independent repeats with similar results. Utilization of VHs presented in b–h are compared to those in NT parental cells shown in a, with red, blue, and grey bars indicating VHs with significantly increased, decreased, and unchanged utilization, respectively. To facilitate direct comparisons, the HTGTS-V(D)J-seq data for the IAA-treated parental and CTCF-degron cells shown in b and h are the same as those presented in Fig. 3b, c, respectively, with different y-axis scales. p values were calculated using unpaired two-tailed t-test (*: p ≤ 0.05, **: p ≤ 0.01, and ***: p ≤ 0.001). See Supplementary Table 3 for exact p values. See Methods for details.
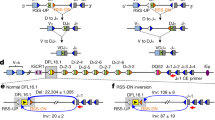
Extended Data Fig. 7 Substantial CTCF depletion promotes robust RAG utilization of cryptic RSSs across the VH locus in G1-arrested IAA-treated CTCF-degron cells.
a, Diagram of joining between RAG-mediated breaks at initiating bona fide JH4-RSS (blue triangle) and cryptic RSSs, mostly represented by CAC motifs (green triangles), across Igh locus, upon down-modulation of CTCF/CBEs (indicated by light purple arrowheads). Deletional and inversional joining is denoted by red (+) and blue (−) curves, respectively. b, Pooled HTGTS junction profiles across highlighted VH domains for deletional and inversional joining in RAG2-complemented, IAA-treated and G1-arrested parental (n=4 biological repeats with similar results) and CTCF-degron (n=6 biological repeats with similar results) cells. For presentation, all libraries from parental or CTCF-degron cells were pooled and normalized to 3,819 total off-target junctions. Note that the high-level deletional cryptic RSS joining signals (red bars) within IGCR1-JH region are off-scale on the y-axis for viewing of relatively lower level deletional cryptic RSS joining signals across the VH locus. The low-level, but reproducible, inversional joining events (blue bars) within IGCR1-JH region involve cryptic RSSs predominantly at IGCR1 and DH3-2 locales, as described2. The vast majority of deletional joining events across the VH locus are reproducible among replicates and involve, at a minimum the CAC of an RSS. Almost all of the handful of the very low-level inversional joining events interspersed across VH locus are not reproducible among replicates consistent with being background events. c, Average frequencies ± s.d. of deletional (left) and inversional (right) joining events within indicated regions in RAG2-complemented, IAA-treated and G1-arrested parental (n=4 biological repeats with similar results) and CTCF-degron (n=6 biological repeats with similar results) cells. Each of the biological library replicates was normalized to 415 off-target junctions for statistical analysis. p values were calculated using unpaired two-tailed t-test. NA: not applicable. See Methods.
Extended Data Fig. 8 A working model for CTCF orchestrating long-range cohesin-driven V(D)J recombinational scanning.
a, Diagram shows mouse VH locus with 100-plus VHs in four domains23,24 as described in Fig. 2a, b legend and the main text. b, With normal CTCF/CBE impediment activity (red arrowheads), RAG scanning is strongly impeded by CBE-based IGCR1 and non-CBE-based DJHRC impediments2,5, allowing scanning-based D-to-JH rearrangement to precede VH-to-DJH rearrangement2. Low-level scanning beyond IGCR1 allows proximal VH-to-DJH rearrangement in a minor subset of pro-B cells. Proximal VH-CBEs impedes further upstream scanning5,47. c, Down-modulation of CTCF reduces CTCF-occupancy of CBEs (grey arrowheads) across Igh, but residual binding is retained at certain CBEs (light blue arrowheads). Dampened CTCF/CBE activity allows RC-bound RAG to scan various linear distances across Igh in some cases the full-length. Upon reaching remaining scanning impediments, which could reflect CBEs with residual CTCF binding and/or transcription (dark arrows)2,21,48 and perhaps others not yet defined (see main text, Fig. 3e and Supplementary Data), impeded scanning focuses RC-based RAG activity on sequences in the impeded region2 for VH-to-DJH joining49. d, Down-modulation of CTCF binding activity is one manner for extending extrusion but this could also be achieved by circumventing CBE impediments, for example, through modulation of cohesin activity (see text). Finally, extrusion of chromatin impediments past the RC may be more efficient in the absence of RAG binding2. With respect to the VH locus, a RAG-bound RC may be more resistant passing impediments and also lead to V(D)J rearrangements that could progressively decrease the level of scanning that proceeds further upstream.
Supplementary information
Supplementary Figure 1
Uncropped Southern and Western blot scans.
Supplementary Data
This file contains VH usage, GRO-seq, 3C-HTGTS, ChIP-seq and predicted CBE motifs for all VHs.
Supplementary Tables 1-4
Supplementary Table 1: Relative utilization of Igh V and D segments on IGCR1-intact and IGCR1-deleted alleles in RAG2-complemented G1-arrested Rad21-degron cells without (NT) or with (+IAA) IAA treatment from > 3 biological repeats. Supplementary Table 2: Summary of observed and predicted utilization of DQ52 segment obtained from DJH and VHDJH joins on IGCR1-intact and IGCR1-deleted alleles in RAG2-complemented G1-arrested Rad21-degron cells without (NT) or with (IAA) IAA treatment from >3 biological repeats. Supplementary Table 3: Relative utilization of Igh V and D segments in RAG2-complemented G1-arrested parental, OsTir1-expressing, CTCF-tagged, and CTCF-degron v-Abl cells without (NT) or with (+IAA) IAA treatment, and bone marrow (BM) pro-B cells from >4 biological repeats. Supplementary Table 4: Oligos used in this study.
Rights and permissions
About this article
Cite this article
Ba, Z., Lou, J., Ye, A.Y. et al. CTCF orchestrates long-range cohesin-driven V(D)J recombinational scanning. Nature 586, 305–310 (2020). https://doi.org/10.1038/s41586-020-2578-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2578-0
This article is cited by
-
Boundary stacking interactions enable cross-TAD enhancer–promoter communication during limb development
Nature Genetics (2024)
-
Genetic variation in the immunoglobulin heavy chain locus shapes the human antibody repertoire
Nature Communications (2023)
-
CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion
Nature (2023)
-
New insights into genome folding by loop extrusion from inducible degron technologies
Nature Reviews Genetics (2023)
-
Genome control by SMC complexes
Nature Reviews Molecular Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.