Abstract
Influenza vaccines that confer broad and durable protection against diverse viral strains would have a major effect on global health, as they would lessen the need for annual vaccine reformulation and immunization1. Here we show that computationally designed, two-component nanoparticle immunogens2 induce potently neutralizing and broadly protective antibody responses against a wide variety of influenza viruses. The nanoparticle immunogens contain 20 haemagglutinin glycoprotein trimers in an ordered array, and their assembly in vitro enables the precisely controlled co-display of multiple distinct haemagglutinin proteins in defined ratios. Nanoparticle immunogens that co-display the four haemagglutinins of licensed quadrivalent influenza vaccines elicited antibody responses in several animal models against vaccine-matched strains that were equivalent to or better than commercial quadrivalent influenza vaccines, and simultaneously induced broadly protective antibody responses to heterologous viruses by targeting the subdominant yet conserved haemagglutinin stem. The combination of potent receptor-blocking and cross-reactive stem-directed antibodies induced by the nanoparticle immunogens makes them attractive candidates for a supraseasonal influenza vaccine candidate with the potential to replace conventional seasonal vaccines3.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All images and data were generated and analysed by the authors, and will be made available by the corresponding authors (B.S.G., N.P.K. and M.K.) upon reasonable request. Uncropped images of all gels are provided in Supplementary Fig. 1. Structural models and density maps have been deposited in the Protein Data Bank (PDB) and Electron Microscopy Data Bank (EMDB) under accession numbers EMD-22935 (H1-I53_dn5 nanoparticle), EMD-22937 and PDB 7KNA (localized reconstruction of H1 HA), EMD-22940 (H5 HA bound to 3 polyclonal Fabs), EMD-22939 (H5 HA bound to 2 polyclonal Fabs), and EMD-22938 (H5 HA bound to 1 polyclonal Fab). Influenza reverse genetics plasmids were provided by the St Jude Children’s Research Hospital under a material transfer agreement with the NIH. Requests for these reagents should be made to the St Jude Children’s Research Hospital. Source data are provided with this paper.
References
Wei, C.-J. et al. Next-generation influenza vaccines: opportunities and challenges. Nat. Rev. Drug Discov. 19, 239–252 (2020).
Ueda, G. et al. Tailored design of protein nanoparticle scaffolds for multivalent presentation of viral glycoprotein antigens. eLife 9, e57659 (2020).
Kanekiyo, M. & Graham, B. S. Next-generation influenza vaccines. Cold Spring Harb. Perspect. Med. a038448 (2020).
Iuliano, A. D. et al. Estimates of global seasonal influenza-associated respiratory mortality: a modelling study. Lancet 391, 1285–1300 (2018).
Flannery, B. et al. Interim estimates of 2017-18 seasonal influenza vaccine effectiveness - United States, February 2018. MMWR Morb. Mortal. Wkly. Rep. 67, 180–185 (2018).
Ellebedy, A. H. et al. Induction of broadly cross-reactive antibody responses to the influenza HA stem region following H5N1 vaccination in humans. Proc. Natl Acad. Sci. USA 111, 13133–13138 (2014).
Andrews, S. F. et al. Immune history profoundly affects broadly protective B cell responses to influenza. Sci. Transl. Med. 7, 316ra192 (2015).
Tan, H.-X. et al. Subdominance and poor intrinsic immunogenicity limit humoral immunity targeting influenza HA stem. J. Clin. Invest. 129, 850–862 (2019).
Yassine, H. M. et al. Hemagglutinin-stem nanoparticles generate heterosubtypic influenza protection. Nat. Med. 21, 1065–1070 (2015).
Impagliazzo, A. et al. A stable trimeric influenza hemagglutinin stem as a broadly protective immunogen. Science 349, 1301–1306 (2015).
Corbett, K. S. et al. Design of nanoparticulate group 2 influenza virus hemagglutinin sstem antigens that activate unmutated ancestor B cell receptors of broadly neutralizing antibody lineages. mBio 10, e02810-18 (2019).
Boyoglu-Barnum, S. et al. Glycan repositioning of influenza hemagglutinin stem facilitates the elicitation of protective cross-group antibody responses. Nat. Commun. 11, 791 (2020).
Steel, J. et al. Influenza virus vaccine based on the conserved hemagglutinin stalk domain. mBio 1, e00018-10 (2010).
Bommakanti, G. et al. Design of an HA2-based Escherichia coli expressed influenza immunogen that protects mice from pathogenic challenge. Proc. Natl Acad. Sci. USA 107, 13701–13706 (2010).
Krammer, F., Pica, N., Hai, R., Margine, I. & Palese, P. Chimeric hemagglutinin influenza virus vaccine constructs elicit broadly protective stalk-specific antibodies. J. Virol. 87, 6542–6550 (2013).
Marcandalli, J. et al. Induction of potent neutralizing antibody responses by a designed protein nanoparticle vaccine for respiratory syncytial virus. Cell 176, 1420–1431.e17 (2019).
Kanekiyo, M. et al. Self-assembling influenza nanoparticle vaccines elicit broadly neutralizing H1N1 antibodies. Nature 499, 102–106 (2013).
López-Sagaseta, J., Malito, E., Rappuoli, R. & Bottomley, M. J. Self-assembling protein nanoparticles in the design of vaccines. Comput. Struct. Biotechnol. J. 14, 58–68 (2015).
Tokatlian, T. et al. Innate immune recognition of glycans targets HIV nanoparticle immunogens to germinal centers. Science 363, 649–654 (2019).
Kanekiyo, M. et al. Mosaic nanoparticle display of diverse influenza virus hemagglutinins elicits broad B cell responses. Nat. Immunol. 20, 362–372 (2019).
Cohen, A. A. et al. Mosaic nanoparticles elicit cross-reactive immune responses to zoonotic coronaviruses in mice. Science 371, 735–741 (2021).
Georgiev, I. S. et al. Two-component ferritin nanoparticles for multimerization of diverse trimeric antigens. ACS Infect. Dis. 4, 788–796 (2018).
Cohen, A. A. et al. Construction, characterization, and immunization of nanoparticles that display a diverse array of influenza HA trimers. PLoS ONE 16, e0247963 (2021).
King, N. P. et al. Accurate design of co-assembling multi-component protein nanomaterials. Nature 510, 103–108 (2014).
Bale, J. B. et al. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science 353, 389–394 (2016).
Martín, J. et al. Studies of the binding properties of influenza hemagglutinin receptor-site mutants. Virology 241, 101–111 (1998).
Whittle, J. R. R. et al. Flow cytometry reveals that H5N1 vaccination elicits cross-reactive stem-directed antibodies from multiple Ig heavy-chain lineages. J. Virol. 88, 4047–4057 (2014).
Creanga, A. et al. A comprehensive influenza reporter virus panel for high-throughput deep profiling of neutralizing antibodies. Nat. Commun. https://doi.org/10.1038/s41467-021-21954-2 (2021).
Corti, D. et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 333, 850–856 (2011).
Bianchi, M. et al. Electron-microscopy-based epitope mapping defines specificities of polyclonal antibodies elicited during HIV-1 BG505 envelope trimer immunization. Immunity 49, 288–300.e8 (2018).
Kallewaard, N. L. et al. Structure and function analysis of an antibody recognizing all influenza A subtypes. Cell 166, 596–608 (2016).
Joyce, M. G. et al. Vaccine-induced antibodies that neutralize group 1 and group 2 influenza A viruses. Cell 166, 609–623 (2016).
Wei, C.-J. et al. Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060–1064 (2010).
Darricarrère, N. et al. Development of a Pan-H1 influenza vaccine. J. Virol. 92, e01349-18 (2018).
Giles, B. M. & Ross, T. M. A computationally optimized broadly reactive antigen (COBRA) based H5N1 VLP vaccine elicits broadly reactive antibodies in mice and ferrets. Vaccine 29, 3043–3054 (2011).
Broecker, F. et al. A mosaic hemagglutinin-based influenza virus vaccine candidate protects mice from challenge with divergent H3N2 strains. NPJ Vaccines 4, 31 (2019).
Sun, W. et al. Development of influenza B universal vaccine candidates using the “mosaic” hemagglutinin approach. J. Virol. 93, e00333-19 (2019).
Ng, S. et al. Novel correlates of protection against pandemic H1N1 influenza A virus infection. Nat. Med. 25, 962–967 (2019).
Fonville, J. M. et al. Antibody landscapes after influenza virus infection or vaccination. Science 346, 996–1000 (2014).
Gostic, K. M., Ambrose, M., Worobey, M. & Lloyd-Smith, J. O. Potent protection against H5N1 and H7N9 influenza via childhood hemagglutinin imprinting. Science 354, 722–726 (2016).
Throsby, M. et al. Heterosubtypic neutralizing monoclonal antibodies cross-protective against H5N1 and H1N1 recovered from human IgM+ memory B cells. PLoS ONE 3, e3942 (2008).
Hong, M. et al. Antibody recognition of the pandemic H1N1 influenza virus hemagglutinin receptor binding site. J. Virol. 87, 12471–12480 (2013).
Ekiert, D. C. et al. A highly conserved neutralizing epitope on group 2 influenza A viruses. Science 333, 843–850 (2011).
Iba, Y. et al. Conserved neutralizing epitope at globular head of hemagglutinin in H3N2 influenza viruses. J. Virol. 88, 7130–7144 (2014).
Lee, P. S. et al. Receptor mimicry by antibody F045-092 facilitates universal binding to the H3 subtype of influenza virus. Nat. Commun. 5, 3614 (2014).
Dreyfus, C. et al. Highly conserved protective epitopes on influenza B viruses. Science 337, 1343–1348 (2012).
Wu, Y. et al. A potent broad-spectrum protective human monoclonal antibody crosslinking two haemagglutinin monomers of influenza A virus. Nat. Commun. 6, 7708 (2015).
Kwakkenbos, M. J. et al. Generation of stable monoclonal antibody-producing B cell receptor-positive human memory B cells by genetic programming. Nat. Med. 16, 123–128 (2010).
Corti, D. et al. Cross-neutralization of four paramyxoviruses by a human monoclonal antibody. Nature 501, 439–443 (2013).
Studier, F. W. & William Studier, F. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Snijder, J. et al. Vitrification after multiple rounds of sample application and blotting improves particle density on cryo-electron microscopy grids. J. Struct. Biol. 198, 38–42 (2017).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Ilca, S. L. et al. Localized reconstruction of subunits from electron cryomicroscopy images of macromolecular complexes. Nat. Commun. 6, 8843 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Frenz, B. et al. Automatically fixing errors in glycoprotein structures with Rosetta. Structure 27, 134–139.e3 (2019).
Wang, R. Y.-R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Verkerke, H. P. et al. Epitope-independent purification of native-like envelope trimers from diverse HIV-1 isolates. J. Virol. 90, 9471–9482 (2016).
Guttman, M., Weis, D. D., Engen, J. R. & Lee, K. K. Analysis of overlapped and noisy hydrogen/deuterium exchange mass spectra. J. Am. Soc. Mass Spectrom. 24, 1906–1912 (2013).
Weis, D. D., Engen, J. R. & Kass, I. J. Semi-automated data processing of hydrogen exchange mass spectra using HX-Express. J. Am. Soc. Mass Spectrom. 17, 1700–1703 (2006).
Martínez-Sobrido, L. et al. Hemagglutinin-pseudotyped green fluorescent protein-expressing influenza viruses for the detection of influenza virus neutralizing antibodies. J. Virol. 84, 2157–2163 (2010).
Gao, Q. et al. The influenza A virus PB2, PA, NP, and M segments play a pivotal role during genome packaging. J. Virol. 86, 7043–7051 (2012).
Bloom, J. D., Gong, L. I. & Baltimore, D. Permissive secondary mutations enable the evolution of influenza oseltamivir resistance. Science 328, 1272–1275 (2010).
Kong, W.-P. et al. Protective immunity to lethal challenge of the 1918 pandemic influenza virus by vaccination. Proc. Natl Acad. Sci. USA 103, 15987–15991 (2006).
Yang, Z.-Y. et al. Immunization by avian H5 influenza hemagglutinin mutants with altered receptor binding specificity. Science 317, 825–828 (2007).
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: Adaptive regularization improves single particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2019).
Acknowledgements
We thank K. Foulds, A. Noe, S.-F. Kao, V. Ficca, N. Nji, D. Flebbe and E. McCarthy for help with non-human primate experiments; A. Taylor, H. Bao, C. Chiedi, M. Dillon, L. Gilman, G. Sarbador, E. McCarthy, J.-P. Todd and D. Scorpio for help with mouse, ferret and NHP experiments; H. Andersen, N. Jones and G. Patel for help with influenza challenge studies; R. Webby for providing influenza reverse genetics plasmids; Y. Tsybovsky and T. Stephens for initial electron microscopy screening; A. Reers and P. Myler for assistance with protein production; and members of the King laboratory and the Influenza Program at the VRC for comments on the manuscript. This study was supported by the intramural research program of the Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health (M.K. and B.S.G.); a gift from the Open Philanthropy Project (D.B. and N.P.K.); a gift from the Audacious Project (D.B. and N.P.K.); the Defense Threat Reduction Agency (HDTRA1-18-1-0001; D.B. and N.P.K.); the National Institute of General Medical Sciences (R01GM120553; D.V.); the National Institute of Allergy and Infectious Diseases (DP1AI158186 and HHSN272201700059C; D.V.); a Pew Biomedical Scholars Award (D.V.); an Investigators in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund (D.V.); and the National Institute of General Medical Sciences (R01GM099989; K.K.L.); and the University of Washington Arnold and Mabel Beckman Cryo-EM Center. Molecular graphics and analyses performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
Conceptualization: B.S.G., N.P.K., M.K. Modelling and design: D.E., G.U., N.P.K., M.K. Formal analysis: S.B.-B., D.E., R.A.G., Y.-J.P., O.J.A., M.J.W., S.K., K.K.L., M.G., D.V., N.P.K., M.K. Investigation: S.B.-B., D.E., R.A.G., G.B.H., A.C., Y.-J.P., O.J.A., S.M.M., R.R., M.M., D.P., N.M., L.C., M.J.W., S.K., S.A., J.R.V., K.K.L., M.G., D.V., N.P.K., M.K. Resources: A.C., G.U., L.S., D.B. Writing, original draft: S.B.-B., D.E., D.V., N.P.K., M.K. Writing, review and editing: all authors. Visualization: S.B.-B., D.E., Y.-J.P., D.V., N.P.K., M.K. Supervision: K.K.L., M.G., J.R.M., D.V., B.S.G., N.P.K., M.K. Project administration: M.C.C. Funding acquisition: L.S., D.V., J.R.M., B.S.G., D.B., N.P.K. R.A.G. and G.B.H. contributed equally.
Corresponding authors
Ethics declarations
Competing interests
S.B.-B., D.E., R.A.G., G.U., B.S.G., N.P.K. and M.K. are listed as inventors on a patent application based on the studies presented in this paper. D.V. is a consultant for Vir Biotechnology Inc. The Veesler laboratory has received an unrelated sponsored research agreement from Vir Biotechnology Inc. N.P.K. is a co-founder, shareholder, and chair of the scientific advisory board of Icosavax, Inc. L.S. is a shareholder of Icosavax, Inc. The King laboratory has received an unrelated sponsored research agreement from Pfizer. D.B. is a co-founder and shareholder of Icosavax, Inc. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks Steve Gamblin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Production and characterization of HA-I53_dn5 components and nanoparticle immunogens.
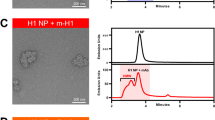
a, SEC purification of seasonal HAs fused to I53_dn5B trimeric components, using a Superdex 200 Increase 10/300 GL column. b, Reducing and non-reducing SDS–PAGE of SEC-purified trimeric HA-I53_dn5B fusions, pentameric I53_dn5A component, and I53_dn5B trimer lacking fused HA. c, SEC purification of nanoparticle immunogens after in vitro assembly, including I53_dn5 lacking displayed antigen, using a Superose 6 Increase 10/300 GL column. The nanoparticle immunogens elute at the void volume of the column, while I53_dn5 is resolved. Residual, unassembled trimeric and pentameric components elute around 15 ml and 18 ml, respectively. d, Reducing and non-reducing SDS–PAGE of SEC-purified nanoparticle immunogens and I53_dn5. e, Antigenic characterization of purified nanoparticle immunogens by ELISA. Symbols indicate the specificity of each monoclonal antibody. AUC, area under the curve. f, Analytical SEC of purified nanoparticle immunogens, compared to I53_dn5 nanoparticles lacking displayed antigen and trimeric H1-I53_dn5B, using a Sephacryl S-500 HR 16/60 column. g, Dynamic light scattering of SEC-purified nanoparticle immunogens, including I53_dn5. Dh, hydrodynamic diameter; Pd, polydispersity. h, Representative electron micrograph of H1-I53_dn5 embedded in vitreous ice. Scale bar, 100 nm. i, 2D class averages obtained using single-particle cryo-EM. Scale bar, 20 nm. j, Gold-standard Fourier shell correlation (FSC) curve for the H1-I53_dn5 density map presented in Fig. 1c. k, Gold-standard FSC curve for the localized reconstruction of H1 MI15 presented in Fig. 1c. All experiments except for electron microscopy data collection and processing were performed at least twice.
Extended Data Fig. 2 Hydrogen–deuterium exchange mass spectrometry (HDX-MS) of H1-foldon trimer and H1-I53_dn5 nanoparticle.
a, Amino acid sequence of H1 ectodomain expressed as a genetic fusion to both foldon and I53_dn5B. Underlined sequences correspond to peptides analysed by HDX-MS. b, Hydrogen–deuterium exchange percentages after 20 h for both samples mapped onto the structure of H1 HA (PDB 3LZG). c, Kinetics of hydrogen–deuterium exchange for both samples at multiple time points up to 20 h. Single asterisks denote peptides in which a negative percentage exchange was corrected to zero (<2% magnitude correction); double asterisks denote peptides that were missing a replicate at the 30 min time point.
Extended Data Fig. 3 Controllable co-display of multiple antigenic variants on two-component nanoparticle immunogens.
a, Sandwich BLI comparing qsCocktail-I53_dn5 and qsMosaic-I53_dn5. Biotinylated 5J8 immobilized on streptavidin probes was used to capture H1-containing nanoparticles from each sample. The captured particles were then exposed to antibodies specific to H3 (CR8020; left) or influenza B HA (CR8071; right). b, Numerical approximation of the H1 HA content of individual qsMosaic-I53_dn5 nanoparticles assuming an equimolar quadrivalent in vitro assembly reaction (that is, 25% of the input HA-I53_dn5B trimers bear H1 HA) and random incorporation of each HA-I53_dn5B trimer at each of the 20 trimeric positions into the nanoparticle. A distribution centred on 25% valency (5 H1 HA trimers per nanoparticle) is observed. c, Calculation of the fraction of individual mosaic nanoparticles displaying at least one H1 HA trimer as a function of the fractional concentration of H1-I53-dn5B in the in vitro assembly reaction ([H1]), expressed as: 1 − (1 − [H1])20. At the 25% fractional concentration used to assemble qsMosaic-I53_dn5, 99.7% of the individual nanoparticles are expected to display at least one H1 HA trimer. d, Quantification of HA antigen content by peptide mass spectrometry in three distinct qsMosaic-I53_dn5 nanoparticles with various antigen ratios before and after preparative SEC. Dashed lines represent the fractional concentration of each HA in the in vitro assembly reactions used to prepare the mosaic nanoparticle immunogens, main bars represent the mean values of four unique peptides from each HA, and error bars represent the standard deviation of measurements across the four unique peptides from each HA. The peptides used to quantify each HA are provided in Supplementary Table 3.
Extended Data Fig. 4 Vaccine-elicited antibody responses against vaccine-matched antigens.
a–c, HA-specific antibody titres in immunized mice (a), ferrets (b) and NHPs (c). Immunization schemes are shown at the top of each panel. All immunizations were given intramuscularly with AddaVax. Groups of BALB/cJ mice (n = 10), ferrets (n = 9), and rhesus macaques (n = 4) were used in each experiment. ELISA antibody titres are expressed as endpoint dilutions. Each symbol represents an individual animal and the horizontal bar indicates the geometric mean of the group. Individual NHPs are identified by unique symbols. d, Antibody responses against unmodified I53_dn5 nanoparticles lacking displayed HA. Immunization scheme is shown at the top of the panel. Groups of NHPs (n = 4) were immunized three times with either QIV, qsCocktail-I53-dn5 or qsMosaic-I53_dn5 with AddaVax at weeks 0, 8 and 16. Serum samples were collected 2 weeks after each immunization and tested for ELISA binding antibody against unmodified I53_dn5 particles. Antibody titres are expressed as endpoint dilutions. Individual NHPs are identified by unique symbols. The immunization study was performed once. e, Antibody responses against vaccine-matched antigens and viruses elicited by unadjuvanted vaccines in immunized mice. Immunization scheme is shown. All immunizations were given intramuscularly. Groups of BALB/cJ mice (n = 10) were used. HA-specific ELISA binding antibody (top), HAI (middle), and microneutralization titres (bottom) in immune sera are shown. Microneutralization titres are reported as IC50 values. Each symbol represents an individual animal, and the horizontal bar indicates the geometric mean of the group. P values were determined by nonparametric Kruskal–Wallis tests with Dunn’s multiple comparisons. All animal experiments except for NHP were performed at least twice and representative data are shown.
Extended Data Fig. 5 Antibody responses against vaccine-matched antigens and viruses elicited by 2018–2019 vaccines.
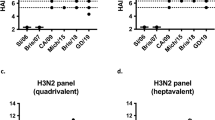
a, Immunization scheme. The commercial QIV, qsCocktail-I53_dn5 and qsMosaic-I53_dn5 vaccines used in this study comprised the 2018–2019 vaccine strains recommended by the WHO. Sequences for the HA-I53_dn5B fusion proteins—H1-I53_dn5, SG16-I53_dn5 (updated H3), B/Yam-I53_dn5, and CO17-I53_dn5 (updated B/Vic)—are provided in Supplementary Table 1. All immunizations were given intramuscularly with AddaVax. Groups of BALB/cJ mice (n = 10) were used. b–d, HA-specific antibody titres (b), HAI assay (c) and microneutralization titres (d) in immune sera. Microneutralization titres are reported as IC50 values. e, Heterosubtypic HA-specific antibody titres in immune sera. Each symbol represents an individual animal and the horizontal bar indicates the geometric mean of the group. P values were determined by nonparametric Kruskal–Wallis tests with Dunn’s multiple comparisons. The animal experiment was performed once.
Extended Data Fig. 6 Neutralization of historical H1N1 and H3N2 viruses.
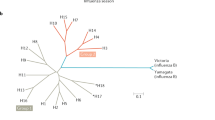
Immunization scheme for the ferret study. Groups of ferrets (n = 9) were used. Phylogenetic trees of HA sequences of human H1N1 (left) and H3N2 (right) viruses are shown (see Supplementary Table 4). Each symbol represents an individual animal and the horizontal bar indicates the geometric mean of the group. P values were determined by nonparametric Kruskal–Wallis tests with Dunn’s multiple comparisons. The ferret experiment was performed twice and representative data are shown.
Extended Data Fig. 7 Antibody responses elicited by a non-assembling immunogen.
a, Model of the I53_dn5B trimer, with the computationally designed interface that drives nanoparticle assembly indicated by the solid line (top), and the 1na0C3_int2 trimer, in which the interface mutations were reverted to their original identities (bottom). The dotted line indicates the inability of this molecule to drive nanoparticle assembly. b, Analytical SEC of the non-assembling immunogen (a mixture of four HA-1na0C3_int2 trimers with pentameric I53_dn5A) using a Superose 6 Increase 10/300 GL column. Only unassembled oligomeric components were observed. c, Reducing and non-reducing SDS–PAGE analysis of the non-assembling immunogen before and after analytical SEC. d, Negative-stain electron microscopy of the non-assembling immunogen, which confirmed the absence of higher-order structures indicated by analytical SEC. Scale bar, 100 nm. e, Immunization scheme in mice. All immunizations were given intramuscularly with AddaVax. Groups of BALB/cJ mice (n = 10) were used in the experiment. f, Microneutralization titres in immune sera against vaccine-matched or slightly mismatched viruses. Microneutralization titres are reported as IC50 values. g, Cross-reactive antibody titres in immune sera. Each symbol represents an individual animal, and the horizontal bar indicates the geometric mean of the group. P values were determined by nonparametric Kruskal–Wallis tests with Dunn’s multiple comparisons. All experiments were performed once.
Extended Data Fig. 8 Negative-stain electron microscopy analysis of H1 HA complexed with polyclonal antibody Fabs prepared from NHPs immunized with qsCocktail-I53_dn5 or qsMosaic-I53_dn5.
a, Negative-stain electron microscopy analysis of Fabs obtained from NHPs immunized with qsCocktail-I53_dn5 in complex with recombinant H1 MI15 HA trimers. Two-dimensional classifications were generated using 847,873 particles collected from 4,112 micrographs. The frequencies of complexes containing Fab fragments bound to RBD (81%), vestigial esterase (18%), or stem (1%) domains are presented as pie charts in Fig. 5c. b, Negative-stain electron microscopy analysis of Fabs obtained from NHPs immunized with qsMosaic-I53_dn5 in complex with recombinant H1 MI15 HA trimers. 2D classifications were generated using 997,557 particles collected from 3,237 micrographs. The frequencies of complexes containing Fab fragments bound to RBD (69%), vestigial esterase (24%), or stem (7%) domains are presented as pie charts in Fig. 5c. The top part of each panel shows representative reference-free 2D class averages. Scale bars, 20 nm. The bottom part of each panel shows seven representative 3D reconstructions of HA–Fab complexes. Single complexes containing Fabs of multiple specificities were counted once against each specificity. The coordinates of an H1 HA crystal structure (PDB 1RUZ) and a Fab fragment (PDB 3GBN) were fitted into the electron microscopy densities. Light blue ribbons, H1 HA; cyan or magenta ribbons, Fabs. All experiments were performed once.
Extended Data Fig. 9 Cryo-EM analysis of heterosubtypic H5 HA in complex with polyclonal antibody Fab fragments prepared from NHP immunized with qsMosaic-I53_dn5.
a, Representative cryo-electron micrograph. Scale bar, 100 nm. b, Reference-free 2D class averages. Scale bar, 20 nm. c, Gold-standard FSC curve for the asymmetric reconstruction shown in d. d, Asymmetric cryo-EM reconstruction of H5 HA–Fab complexes with Fab fragments bound to all three HA subunits at 3.6 Å resolution. The reconstruction is the same as that shown in the right panel in Fig. 5d, but here is coloured by local resolution. e, FSC curve for the asymmetric reconstruction shown in f. f, Two orthogonal orientations of an asymmetric cryo-EM reconstruction of H5 HA–Fab complexes with Fab fragments bound to two HA subunits at 4.1 Å resolution. g, FSC curve for the asymmetric reconstruction shown in h. h, Two orthogonal orientations of an asymmetric cryo-EM reconstruction of H5 HA–Fab complexes with Fab fragments bound to one HA subunit at 4.0 Å resolution. The reconstruction is the same as that shown in the left panel in Fig. 5d. All experiments were performed once.
Extended Data Fig. 10 Vaccine-elicited antibody responses against vaccine-matched viruses in NHPs with pre-existing influenza immunity.
Immunization scheme for the NHP study shown on the left. NHPs (n = 3) that had been immunized three times with either QIV 2017–2018 or qsMosaic-I53_dn5 2017–2018 were boosted 63 weeks later with a single dose (60 μg) of updated qsMosaic-I53_dn5 2018–2019. All immunizations were given intramuscularly with AddaVax. Microneutralization titres are reported IC50 values. Each symbol represents an individual animal and the horizontal bar indicates the geometric mean of the group. Individual NHPs are identified by unique symbols. P values were determined by paired t-tests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1 – 5 and Supplementary Tables 1 – 4.
Supplementary Data
Source Data for Supplementary Figs 2 – 5.
Rights and permissions
About this article
Cite this article
Boyoglu-Barnum, S., Ellis, D., Gillespie, R.A. et al. Quadrivalent influenza nanoparticle vaccines induce broad protection. Nature 592, 623–628 (2021). https://doi.org/10.1038/s41586-021-03365-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03365-x
This article is cited by
-
Enhancing antibody responses by multivalent antigen display on thymus-independent DNA origami scaffolds
Nature Communications (2024)
-
Mosaic quadrivalent influenza vaccine single nanoparticle characterization
Scientific Reports (2024)
-
Bringing immunofocusing into focus
npj Vaccines (2024)
-
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines
npj Vaccines (2024)
-
Diverse array of neutralizing antibodies elicited upon Spike Ferritin Nanoparticle vaccination in rhesus macaques
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.