Abstract
The lack of knowledge about extreme conservation in genomes remains a major gap in our understanding of the evolution of gene regulation. Here, we reveal an unexpected role of extremely conserved 5′ untranslated regions (UTRs) in noncanonical translational regulation that is linked to the emergence of essential developmental features in vertebrate species. Endogenous deletion of conserved elements within these 5′ UTRs decreased gene expression, and extremely conserved 5′ UTRs possess cis-regulatory elements that promote cell-type-specific regulation of translation. We further developed in-cell mutate-and-map (icM2), a new methodology that maps RNA structure inside cells. Using icM2, we determined that an extremely conserved 5′ UTR encodes multiple alternative structures and that each single nucleotide within the conserved element maintains the balance of alternative structures important to control the dynamic range of protein expression. These results explain how extreme sequence conservation can lead to RNA-level biological functions encoded in the untranslated regions of vertebrate genomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw sequencing data (related to Figs. 4, 5 and 6) are deposited to GEO with accession code GSE155656. Processed reactivity data have been deposited in the RNA Mapping Database (RMDB) with accession codes CSDE1_DMS_0000 and CSDE1_DMS_0001. Sources for publicly available data are described in the Methods.
Code availability
All software used to analyze the study data are listed in the Methods and in the Nature Research Reporting Summary and are publicly available. All codes used to analyze icM2 data are available through a Github repository: github.com/barnalab/icm2p.
References
Dermitzakis, E. T., Reymond, A. & Antonarakis, S. E. Conserved non-genic sequences – an unexpected feature of mammalian genomes. Nat. Rev. Genet. 6, 151–157 (2005).
Harmston, N., Baresic, A. & Lenhard, B. The mystery of extreme non-coding conservation. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 368, 20130021 (2013).
Halligan, D. L. et al. Positive and negative selection in murine ultraconserved noncoding elements. Mol. Biol. Evol. 28, 2651–2660 (2011).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Dimitrieva, S. & Bucher, P. Genomic context analysis reveals dense interaction network between vertebrate ultraconserved non-coding elements. Bioinformatics 28, i395–i401 (2012).
Boffelli, D., Nobrega, M. A. & Rubin, E. M. Comparative genomics at the vertebrate extremes. Nat. Rev. Genet. 5, 456–465 (2004).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819 (2005).
Sandelin, A. et al. Arrays of ultraconserved non-coding regions span the loci of key developmental genes in vertebrate genomes. BMC Genomics 5, 99 (2004).
de la Calle-Mustienes, E. et al. A functional survey of the enhancer activity of conserved non-coding sequences from vertebrate Iroquois cluster gene deserts. Genome Res. 15, 1061–1072 (2005).
Sakuraba, Y. et al. Identification and characterization of new long conserved noncoding sequences in vertebrates. Mamm. Genome 19, 703–712 (2008).
Dermitzakis, E. T. et al. Comparison of human chromosome 21 conserved nongenic sequences (CNGs) with the mouse and dog genomes shows that their selective constraint is independent of their genic environment. Genome Res. 14, 852–859 (2004).
Katzman, S. et al. Human genome ultraconserved elements are ultraselected. Science 317, 915 (2007).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006).
Visel, A. et al. Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nat. Genet. 40, 158–160 (2008).
Visel, A. et al. A high-resolution enhancer atlas of the developing telencephalon. Cell 152, 895–908 (2013).
Ahituv, N. et al. Deletion of ultraconserved elements yields viable mice. PLoS Biol. 5, e234 (2007).
McLean, C. & Bejerano, G. Dispensability of mammalian DNA. Genome Res. 18, 1743–1751 (2008).
Dickel, D. E. et al. Ultraconserved enhancers are required for normal development. Cell 172, 491–499.e15 (2018).
Osterwalder, M. et al. Enhancer redundancy provides phenotypic robustness in mammalian development. Nature 554, 239–243 (2018).
Lareau, L. F., Inada, M., Green, R. E., Wengrod, J. C. & Brenner, S. E. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007).
Ni, J. Z. et al. Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay. Genes Dev. 21, 708–718 (2007).
Thomas, J. D. et al. RNA isoform screens uncover the essentiality and tumor-suppressor activity of ultraconserved poison exons. Nat. Genet. 52, 84–94 (2020).
Calin, G. A. et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12, 215–229 (2007).
Liz, J. et al. Regulation of pri-miRNA processing by a long noncoding RNA transcribed from an ultraconserved region. Mol. Cell 55, 138–147 (2014).
Xue, S. et al. RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 517, 33–38 (2015).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Jiang, L. et al. A quantitative proteome map of the human body. Cell 183, 269–283.e19 (2020).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Steri, M., Idda, M. L., Whalen, M. B. & Orrù, V. Genetic variants in mRNA untranslated regions. Wiley Interdiscip. Rev. RNA 9, e1474 (2018).
Blanco-Suarez, E., Liu, T.-F., Kopelevich, A. & Allen, N. J. Astrocyte-secreted chordin-like 1 drives synapse maturation and limits plasticity by increasing synaptic GluA2 AMPA receptors. Neuron 100, 1116–1132.e13 (2018).
Sakuta, H. et al. Ventroptin: a BMP-4 antagonist expressed in a double-gradient pattern in the retina. Science 293, 111–115 (2001).
Webb, T. R. et al. X-linked megalocornea caused by mutations in CHRDL1 identifies an essential role for ventroptin in anterior segment development. Am. J. Hum. Genet. 90, 247–259 (2012).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Liu, T. et al. Chordin-like 1 improves osteogenesis of bone marrow mesenchymal stem cells through enhancing BMP4-SMAD pathway. Front. Endocrinol. 10, 360 (2019).
Pei, Y.-F. et al. Hypermethylation of the CHRDL1 promoter induces proliferation and metastasis by activating Akt and Erk in gastric cancer. Oncotarget 8, 23155–23166 (2017).
Osório, C. et al. Growth differentiation factor 5 is a key physiological regulator of dendrite growth during development. Development 140, 4751–4762 (2013).
O’Keeffe, G. W. et al. Region-specific role of growth differentiation factor-5 in the establishment of sympathetic innervation. Neural Dev. 11, 4 (2016).
Wu, H., Li, J., Xu, D., Zhang, Q. & Cui, T. Growth differentiation factor 5 improves neurogenesis and functional recovery in adult mouse hippocampus following traumatic brain injury. Front. Neurol. 9, 592 (2018).
Buxton, P., Edwards, C., Archer, C. W. & Francis-West, P. Growth/differentiation factor-5 (GDF-5) and skeletal development. J. Bone Joint Surg. Am. 83-A, S23–S30 (2001).
Panganiban, G. & Rubenstein, J. L. R. Developmental functions of the Distal-less/Dlx homeobox genes. Development 129, 4371–4386 (2002).
Depew, M. J., Simpson, C. A., Morasso, M. & Rubenstein, J. L. R. Reassessing the Dlx code: the genetic regulation of branchial arch skeletal pattern and development. J. Anat. 207, 501–561 (2005).
Polleux, F., Morrow, T. & Ghosh, A. Semaphorin 3A is a chemoattractant for cortical apical dendrites. Nature 404, 567–573 (2000).
Serini, G. et al. Class 3 semaphorins control vascular morphogenesis by inhibiting integrin function. Nature 424, 391–397 (2003).
Shelly, M. et al. Semaphorin 3A regulates neuronal polarization by suppressing axon formation and promoting dendrite growth. Neuron 71, 433–446 (2011).
Polleux, F., Giger, R. J., Ginty, D. D., Kolodkin, A. L. & Ghosh, A. Patterning of cortical efferent projections by semaphorin–neuropilin interactions. Science 282, 1904–1906 (1998).
Good, P. F. et al. A role for semaphorin 3A signaling in the degeneration of hippocampal neurons during Alzheimer’s disease. J. Neurochem. 91, 716–736 (2004).
Galan-Caridad, J. M. et al. Zfx controls the self-renewal of embryonic and hematopoietic stem cells. Cell 129, 345–357 (2007).
Lee, A. S. Y., Kranzusch, P. J. & Cate, J. H. D. eIF3 targets cell-proliferation messenger RNAs for translational activation or repression. Nature 522, 111–114 (2015).
Gilbert, W. V., Zhou, K., Butler, T. K. & Doudna, J. A. Cap-independent translation is required for starvation-induced differentiation in yeast. Science 317, 1224–1227 (2007).
Martin, F. et al. Cap-assisted internal initiation of translation of histone H4. Mol. Cell 41, 197–209 (2011).
Legnini, I. et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 66, 22–37.e9 (2017).
Pamudurti, N. R. et al. Translation of circRNAs. Mol. Cell 66, 9–21.e7 (2017).
Leppek, K. et al. Gene- and species-specific Hox mRNA translation by ribosome expansion segments. Mol. Cell 80, 980–995.e13 (2020).
Hershey, J. W. B., Sonenberg, N. & Mathews, M. B. Principles of translational control: an overview. Cold Spring Harb. Perspect. Biol. 4, a011528. (2012).
Weingarten-Gabbay, S. et al. Comparative genetics. Systematic discovery of cap-independent translation sequences in human and viral genomes. Science 351, aad4939 (2016).
Xiao, Z.-S., Simpson, L. G. & Quarles, L. D. IRES-dependent translational control of Cbfa1/Runx2 expression. J. Cell. Biochem. 88, 493–505 (2003).
Jang, G. M. et al. Structurally distinct elements mediate internal ribosome entry within the 5′-noncoding region of a voltage-gated potassium channel mRNA. J. Biol. Chem. 279, 47419–47430 (2004).
Holcik, M. & Sonenberg, N. Translational control in stress and apoptosis. Nat. Rev. Mol. Cell Biol. 6, 318–327 (2005).
El-Naggar, A. M. & Sorensen, P. H. Translational control of aberrant stress responses as a hallmark of cancer. J. Pathol. 244, 650–666 (2018).
Spriggs, K. A., Bushell, M. & Willis, A. E. Translational regulation of gene expression during conditions of cell stress. Mol. Cell 40, 228–237 (2010).
Washietl, S., Hofacker, I. L., Lukasser, M., Hüttenhofer, A. & Stadler, P. F. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat. Biotechnol. 23, 1383–1390 (2005).
Torarinsson, E. et al. Comparative genomics beyond sequence-based alignments: RNA structures in the ENCODE regions. Genome Res. 18, 242–251 (2008).
Parker, B. J. et al. New families of human regulatory RNA structures identified by comparative analysis of vertebrate genomes. Genome Res. 21, 1929–1943 (2011).
Smith, M. A., Gesell, T., Stadler, P. F. & Mattick, J. S. Widespread purifying selection on RNA structure in mammals. Nucleic Acids Res. 41, 8220–8236 (2013).
Eddy, S. R. Computational analysis of conserved RNA secondary structure in transcriptomes and genomes. Annu. Rev. Biophys. 43, 433–456 (2014).
Rivas, E., Clements, J. & Eddy, S. R. Estimating the power of sequence covariation for detecting conserved RNA structure. Bioinformatics 36, 3072–3076 (2020).
Homan, P. J. et al. Single-molecule correlated chemical probing of RNA. Proc. Natl Acad. Sci. USA 111, 13858–13863 (2014).
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75–82 (2017).
Mustoe, A. M., Lama, N. N., Irving, P. S., Olson, S. W. & Weeks, K. M. RNA base-pairing complexity in living cells visualized by correlated chemical probing. Proc. Natl Acad. Sci. USA 116, 24574–24582 (2019).
Beaudoin, J.-D. et al. Analyses of mRNA structure dynamics identify embryonic gene regulatory programs. Nat. Struct. Mol. Biol. 25, 677–686 (2018).
Patalano, S., Mihailovich, M., Belacortu, Y., Paricio, N. & Gebauer, F. Dual sex-specific functions of Drosophila Upstream of N-ras in the control of X chromosome dosage compensation. Development 136, 689–698 (2009).
Elatmani, H. et al. The RNA-binding protein Unr prevents mouse embryonic stem cells differentiation toward the primitive endoderm lineage. Stem Cells 29, 1504–1516 (2011).
Mitchell, S. A., Brown, E. C., Coldwell, M. J., Jackson, R. J. & Willis, A. E. Protein factor requirements of the Apaf-1 internal ribosome entry segment: roles of polypyrimidine tract binding protein and upstream of N-ras. Mol. Cell. Biol. 21, 3364–3374 (2001).
Schepens, B. et al. A role for hnRNP C1/C2 and Unr in internal initiation of translation during mitosis. EMBO J. 26, 158–169 (2007).
Guo, A.-X., Cui, J.-J., Wang, L.-Y. & Yin, J.-Y. The role of CSDE1 in translational reprogramming and human diseases. Cell Commun. Signal. 18, 14 (2020).
Moore, K. S. et al. Csde1 binds transcripts involved in protein homeostasis and controls their expression in an erythroid cell line. Sci. Rep. 8, 2628 (2018).
Wurth, L. et al. UNR/CSDE1 drives a post-transcriptional program to promote melanoma invasion and metastasis. Cancer Cell 30, 694–707 (2016).
Horos, R. et al. Ribosomal deficiencies in Diamond-Blackfan anemia impair translation of transcripts essential for differentiation of murine and human erythroblasts. Blood 119, 262–272 (2012).
Guo, H. et al. Disruptive variants of CSDE1 associate with autism and interfere with neuronal development and synaptic transmission. Sci. Adv. 5, eaax2166 (2019).
Saltel, F. et al. Unr defines a novel class of nucleoplasmic reticulum involved in mRNA translation. J. Cell Sci. 130, 1796–1808 (2017).
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Kladwang, W., VanLang, C. C., Cordero, P. & Das, R. A two-dimensional mutate-and-map strategy for non-coding RNA structure. Nat. Chem. 3, 954–962 (2011).
Cordero, P. & Das, R. Rich RNA structure landscapes revealed by mutate-and-map analysis. PLoS Comput. Biol. 11, e1004473 (2015).
Bhaskaran, H. & Russell, R. Kinetic redistribution of native and misfolded RNAs by a DEAD-box chaperone. Nature 449, 1014–1018 (2007).
Alexa, A., Rahnenführer, J. & Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22, 1600–1607 (2006).
Motenko, H., Neuhauser, S. B., O’Keefe, M. & Richardson, J. E. MouseMine: a new data warehouse for MGI. Mamm. Genome 26, 325–330 (2015).
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Yoon, A. et al. Impaired control of IRES-mediated translation in X-linked dyskeratosis congenita. Science 312, 902–906 (2006).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Acknowledgements
We thank the members of the Barna laboratory for constructive criticism of the manuscript. This work was supported by New York Stem Cell Foundation grant NYSCF-R-I36 (M.B.), NIH grant 1R01HD086634 (M.B.), Alfred P. Sloan Research Fellowship (M.B.), Pew Scholars Award (M.B.), Mallinckrodt Foundation Award (M.B.), Benchmark Stanford Graduate Fellowship (G.W.B.) and Walter and Idun Berry Foundation (E.S.C.). M.B. is a New York Stem Cell Robertson Investigator.
Author information
Authors and Affiliations
Contributions
M.B., G.W.B. and E.S.C. conceived the project. M.B. supervised the project. L.J. and H.T. provided the GTEx data and critical feedback on its analysis. R.D. provided critical feedback on the development and analysis of icM2. E.S.C. carried out the large-scale reporter screens. G.W.B. performed all other experiments and data analysis. G.W.B. and M.B. wrote the manuscript in consultation with all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Jean-Denis Beaudoin, Philip Bevilacqua and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Hyperconserved 5’UTRs in vertebrate genomes.
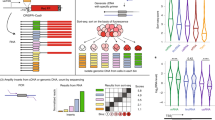
a, Left: heatmap of the positions of LOD ≥ 500 PhastCons elements in each h5UTR. Middle: heatmap of the relative positions (calculated in 100 bins across the h5UTRs) of the elements. Right: plot of average element overlap across the 100 bins to illustrate the positional preference. b, Histogram of the length of h5UTRs. Average length is 674nt. c, Histogram of the number of nucleotides overlap between LOD ≥ 500 PhastCons elements and h5UTRs. Average overlap is 389nt. d, Distributions of cross-tissue transcriptome-proteome correlations for all genes, genes with h5UTRs, or genes with variance-matched non-conserved 5’UTRs. Indicated p-values are from two-sided Wilcoxon rank sum tests for cross-tissue correlation values between h5UTR genes and all genes or between h5UTR genes and variance-matched non-conserved controls. e, Distributions of the number of annotated alternative 5’UTRs for all genes, genes with h5UTRs, or genes with size-matched non-conserved 5’UTRs. Indicated p-values are from two-sided Wilcoxon rank sum tests for the number of alternative 5’UTRs between h5UTR genes and all genes or between h5UTR genes and size-matched non-conserved controls. f, Scatter plot illustrating the lack of significant term enrichments for a size-matched set of non-conserved 5’ UTRs. X-axis and y-axis plots expected and the observed number of genes for each term. Blue dashed line indicates the minimum observed/expected ratio cutoff of 3. Green line indicates expected and observed counts where Fisher’s test p-value (pf) is estimated to have FDR = 0.05. Neighbor-weighted test p-value (pfw) ≤0.05 is further used as an additional cutoff. The final set of enriched terms passing filter is colored by pf and sized by pfw.
Extended Data Fig. 2 Non-canonical translation activation by hyperconserved 5’UTRs across cell types.
a, Density plots of non-canonical translation initiation activities from h5UTRs by bicistronic reporter assay. X-axis is the luciferase reporter activity ratios. Jittered dots mark individual reporter ratios for each h5UTR in each cell type. b, Summarized plot of ribosome load (sum of % mRNA times the ribosome number for each fraction) differential ratio between NSCs and ESCs calculated from polysome profiles for each gene shown in Extended Data Fig. 2c-l. Red indicates significant increase in NSCs and black indicates significant decrease (two-sided t-test p ≤ 0.05, n = 3, marked by asterisk). c-l, Endogenous polysome profiles of NSCs versus ESCs for genes with h5UTRs that show high non-canonical translation reporter activities in NSCs compared to ESCs. Distribution of mRNAs across sucrose gradient fractions are plotted. Y-axis plots the mean percent mRNA. Error bars indicate standard error. Asterisk indicates two-sided t-test p ≤ 0.05 for each fraction between the two cell types. n = 3 for each cell type. Indicated p-value (pf) is calculated by Fisher’s method across all fractions. Note that Extended Data Fig. 2c shows the profile of 18 S rRNA, which indicates lower global translation in NSCs compared to ESCs.
Extended Data Fig. 3 Non-canonical activation by hyperconserved 5’UTRs substantially contributes to translation.
a, Scatter plot of luciferase activity versus RNA level ratios (mean from n = 3) observed for the bicistronic reporters of 90 h5UTRs measured in 10T1/2 cells. Dashed line marks the 10% FDR used in Fig. 3a. Spearman correlation indicated on top left. b, The effect of various truncations of the h5UTRs on non-canonical initiation and total translation efficiency (also see Fig. 3d). Left: positions of truncations. Dashed lines indicate truncations. Purple horizontal lines indicate uORFs; yellow and red lines indicate in-frame and out-of-frame uAUGs, respectively. Middle: non-canonical initiation efficiency. Right: total translation efficiency. X-axis indicates the mean of luciferase reporter ratios relative to the wild-type. Error bars indicate standard error. Dashed line marks the wild-type 5’UTR activity. Asterisk indicates two-sided t-test p ≤ 0.05 for each truncation versus the full-length. The numbers to the left of the bars indicate n and p-values. c, Comparison of translational activities between the full-length long, non-conserved 5’UTRs versus the only first 300nt truncation. 11 different pairs are tested. X-axis indicates the mean log2 luciferase reporter ratios of each truncation relative to its full-length wild-type. Error bars indicate standard error. Bars colored in red indicate significantly reduced translation in the shorter, truncated 300nt fragment; black indicates significant increase (two-sided t-test, paired n = 3, p ≤ 0.05, marked by asterisk). The numbers to the left of the bars indicate p-values. d, Violin plot of full-length/truncated reporter activity ratios (log2) from hyperconserved and non-conserved 5’UTRs. p indicates two-sided Wilcoxon rank sum test p-value. Box hinges: 25% quantile, median, 75% quantile, respectively from left to right. Whiskers: lower or upper hinge ±1.5*IQR. e, Scatter plot of change in translation efficiency between full-length and truncated h5UTRs shown in Fig. 3e versus change in uAUG density (change in number of AUGs / change in length between each pair of full-length and truncated h5UTRs). r indicates pearson’s correlation coefficient and p indicates two-tailed p-value.
Extended Data Fig. 4 Cellular remodeling of hyperconserved 5’UTR RNA structures.
a, Stacked bar plots showing proportions of significant (FDR ≤ 0.05) or not significant windows that overlap uAUG in black versus that do not overlap uAUG in red. OR indicates odds ratio for overlaps uAUG / does not overlap uAUG, and p indicates Fisher’s test p-value (one-sided, Ha = odds ratio>0). b, Stacked bar plots showing proportions of significant (FDR ≤ 0.05) or not significant windows that overlap uORF in black versus that do not overlap uORF in red. OR indicates odds ratio for overlaps uORF / does not overlap uORF, and p indicates Fisher’s test p-value (one-sided, Ha = odds ratio>0). c, Zoomed-in view of differential accessibilities along h5UTRs with one or more significantly different windows under ATP depletion. Top plot shows -log10 p-value for each window. Highlighted boxes mark significantly different windows, above the dashed line indicating 5% FDR. Middle plot shows differential accessibility on the y-axis, where greater than zero indicates increased accessibility upon ATP depletion and less than zero indicates decreased accessibility. Bottom plot shows differential accessibility for in vitro refolded RNA. Error bars in each plot show standard error, n = 3. The three profiled regions shown on the left side exhibit discordant profiles between accessibility changes observed in cells following ATP depletion and accessibility changes observed for in cell versus in vitro refolded RNA. The other three on the right side exhibit concordant profiles.
Extended Data Fig. 6 In-vitro M2 analysis of Csde1 5’UTR.
a, Heatmap of in-vitro M2 accessibility matrix for Csde1 5’UTR from position 190 to 386. For each row, the chemical mapping profile of a single-nucleotide variant of the RNA is plotted across the columns, where the colors indicate z-scaled accessibility change values from the wild-type RNA. 1D data from each mutant are vertically stacked to display a 2D matrix. White boxes mark perturbation signals that support the model shown in Extended Data Fig. 6b; color bars at the bottom indicate the nucleotide positions of the stems that match the same color in the model. b, The model for the in-vitro structure of Csde1 5’UTR from position 190 to 386. Also see Extended Data Fig. 6a.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and Notes
Supplementary Tables
Supplementary Tables 1–9
Rights and permissions
About this article
Cite this article
Byeon, G.W., Cenik, E.S., Jiang, L. et al. Functional and structural basis of extreme conservation in vertebrate 5′ untranslated regions. Nat Genet 53, 729–741 (2021). https://doi.org/10.1038/s41588-021-00830-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00830-1
This article is cited by
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
Amplifying gene expression with RNA-targeted therapeutics
Nature Reviews Drug Discovery (2023)
-
Trait Improvement of Solanaceae Fruit Crops for Vertical Farming by Genome Editing
Journal of Plant Biology (2023)
-
Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells
Nature Communications (2022)
-
Perfect and imperfect views of ultraconserved sequences
Nature Reviews Genetics (2022)