Abstract
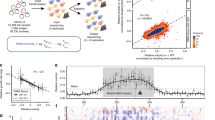
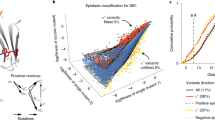
Defining the biologically active structures of proteins in their cellular environments remains challenging for proteins with multiple conformations and functions, where only a minor conformer might be associated with a given function. Here, we use deep mutational scanning to probe the structure and dynamics of α-synuclein, a protein known to adopt disordered, helical and amyloid conformations. We examined the effects of 2,600 single-residue substitutions on the ability of intracellularly expressed α-synuclein to slow the growth of yeast. Computational analysis of the data showed that the conformation responsible for this phenotype is a long, uninterrupted, amphiphilic helix with increasing dynamics toward the C terminus. Deep mutational scanning can therefore determine biologically active conformations in cellular environments, even for a highly dynamic multi-conformational protein.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw sequencing data are available at the NCBI Sequence Read Archive (PRJNA564806). Source data for Figs. 2–4 and Supplementary Fig. 2 are available online. Unprocessed images are available upon request.
Code availability
Programs developed to analyze the data reported are available at github.com/rnewberry17.
References
Socolich, M. et al. Evolutionary information for specifying a protein fold. Nature 437, 512–518 (2005).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Natl Acad. Sci. USA 108, E1293–E1301 (2011).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, e28766 (2011).
Toth-Petroczy, A. et al. Structured states of disordered proteins from genomic sequences. Cell 167, 158–170.e12 (2016).
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Bolognesi, B. et al. The mutational landscape of a prion-like domain. Nat. Commun. 10, 4162 (2019).
Gray, V. E. et al. Elucidating the molecular determinants of Aβ aggregation with deep mutational scanning. G3 (Bethesda) 9, 3683–3689 (2019).
Schmiedel, J. M. & Lehner, B. Determining protein structures using deep mutagenesis. Nat. Genet. 51, 1177–1186 (2019).
Rollins, N. J. et al. Inferring protein 3D structure from deep mutation scans. Nat. Genet. 51, 1170–1176 (2019).
Weinreb, P. H., Zhen, W., Poon, A. W., Conway, K. A. & Lansbury, P. T. Jr. NACP, a protein implicated in Alzheimer’s disease and learning, is natively unfolded. Biochemistry 35, 13709–13715 (1996).
Theillet, F.-X. et al. Structural disorder of monomeric α-synuclein persists in mammalian cells. Nature 530, 45–50 (2016).
Conway, K. A. et al. Acceleration of oligomerization, not fibrillization, is a shared property of both α-synuclein mutations linked to early-onset Parkinson’s disease: implications for pathogenesis and therapy. Proc. Natl Acad. Sci. USA 97, 571–576 (2000).
Bartels, T., Choi, J. G. & Selkoe, D. J. α-Synuclein occurs physiologically as a helically folded tetramer that resists aggregation. Nature 477, 107–110 (2011).
Ulmer, T. S., Bax, A., Cole, N. B. & Nussbaum, R. L. Structure and dynamics of micelle-bound human α-synuclein. J. Biol. Chem. 280, 9595–9603 (2005).
Jao, C. C., Hegde, B. G., Chen, J., Haworth, I. S. & Langen, R. Structure of membrane-bound α-synuclein from site-directed spin labeling and computational refinement. Proc. Natl Acad. Sci. USA 105, 19666–19671 (2008).
Fusco, G. et al. Direct observation of the three regions in α-synuclein that determine its membrane-bound behaviour. Nat. Commun. 5, 3827 (2014).
Tuttle, M. D. et al. Solid-state NMR structure of a pathogenic fibril of full-length human α-synuclein. Nat. Struct. Mol. Biol. 23, 409–415 (2016).
Guerrero-Ferreira, R. et al. Cryo-EM structure of alpha-synuclein fibrils. eLife 7, e36402 (2018).
Li, B. et al. Cryo-EM of full-length α-synuclein reveals fibril polymorphs with a common structural kernel. Nat. Commun. 9, 3609 (2018).
Nelson, R. et al. Structure of the cross-β spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Fernandez-Escamilla, A.-M., Rousseau, F., Schymkowitz, J. & Serrano, L. Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nat. Biotech. 22, 1302–1306 (2004).
Outeiro, T. F. & Lindquist, S. Yeast cells provide insight into alpha-synuclein biology and pathobiology. Science 302, 1772–1775 (2003).
Soper, J. H. et al. α-Synuclein-induced aggregation of cytoplasmic vesicles in Saccharomyces cerevisiae. Mol. Biol. Cell 19, 1093–1103 (2008).
Shahmoradian, S. H. et al. Lewy pathology in Parkinson’s disease consists of crowded organelles and lipid membranes. Nat. Neurosci. 22, 1099–1109 (2019).
Willingham, S., Outeiro, T. F., DeVit, M. J., Lindquist, S. L. & Muchowski, P. J. Yeast genes that enhance the toxicity of a mutant Huntingtin fragment or α-synuclein. Science 302, 1769–1772 (2003).
Cooper, A. A. et al. α-Synuclein blocks ER-Golgi traffic and Rab1 rescues neuron loss in Parkinson’s models. Science 313, 324–328 (2006).
Gitler, A. D. et al. α-Synuclein is part of a diverse and highly conserved interaction network that includes PARK9 and manganese toxicity. Nat. Genet. 41, 308–315 (2009).
Khurana, V. et al. Genome-scale networks link neurodegenerative disease genes to α-synuclein through specific molecular pathways. Cell Syst. 4, 157–170.e114 (2017).
Tardiff, D. F. et al. Yeast reveal a “druggable” Rsp5/Nedd4 network that ameliorates α-synuclein toxicity in neurons. Science 342, 979–983 (2013).
Fanning, S. et al. Lipidomic analysis of α-synuclein neurotoxicity identifies stearoyl CoA desaturase as a target for Parkinson treatment. Mol. Cell 73, 1–14 (2018).
Chung, C. Y. et al. Identification and rescue of α-synuclein toxicity in Parkinson patient-derived neurons. Science 342, 983–987 (2013).
Volles, M. J. & Lansbury, P. T. Jr. Relationships between the sequence of α-synuclein and its membrane affinity, fibrillization propensity, and yeast toxicity. J. Mol. Biol. 366, 1510–1522 (2007).
Bodner, C. R., Dobson, C. M. & Bax, A. Multiple tight phospholipid-binding modes of α-synuclein revealed by solution NMR spectroscopy. J. Mol. Biol. 390, 775–790 (2009).
Fakhree, M. A. A., Engelbertink, S. A. J., van Leijenhorst-Groener, K. A., Blum, C. & Claessens, M. M. A. E. Cooperation of helix insertion and lateral pressure to remodel membranes. Biomacromolecules 20, 1217–1223 (2019).
Burré, J., Sharma, M. & Südhof, T. C. Systematic mutagenesis of α-synuclein reveals distinct sequence requirements for physiological and pathological activities. J. Neurosci. 32, 15227–15242 (2012).
Bendor, J. T., Logan, T. P. & Edwards, R. H. The function of α-synuclein. Neuron 79, 1044–1066 (2013).
Segrest, J. P. et al. The amphipathic helix in the exchangeable apolipoproteins: a review of secondary structure and function. J. Lipid Res. 33, 141–166 (1992).
Bussell, R. Jr. & Eliezer, D. A structural and functional role for 11-mer repeats in α-synuclein and other exchangeable lipid binding proteins. J. Mol. Biol. 329, 763–778 (2003).
Fusco, G. et al. Structural basis of synaptic vesicle assembly promoted by α-synuclein. Nat. Commun. 7, 12563 (2016).
Snead, D. & Eliezer, D. Intrinsically disordered proteins in synaptic vesicle trafficking and release. J. Biol. Chem. 294, 3325–3342 (2019).
Galvagnion, C. et al. Lipid vesicles trigger α-synuclein aggregation by stimulating primary nucleation. Nat. Chem. Biol. 11, 229–234 (2015).
Fusco, G. et al. Structural basis of membrane disruption and cellular toxicity by α-synuclein oligomers. Science 358, 1440–1443 (2017).
Emberly, E. G., Mukhopadhyay, R., Wingreen, N. S. & Tang, C. Flexibility of α-helices: results of a statistical analysis of database protein structures. J. Mol. Biol. 327, 229–237 (2003).
Braun, A. R. et al. α-Synuclein induces both positive mean curvature and negative Gaussian curvature in membranes. J. Am. Chem. Soc. 134, 2613–2620 (2012).
Logan, T., Bendor, J., Toupin, C., Thorn, K. & Edwards, R. H. α-Synuclein promotes dilation of the exocytotic fusion pore. Nat. Neurosci. 20, 681–689 (2017).
Lautenschläger, J., Kaminski, C. F. & Kaminski Schierle, G. S. α-Synuclein—regulator of exocytosis, endocytosis, or both? Trends Cell Biol. 27, 468–479 (2017).
Starita, L. M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Matreyek, K. A. et al. Multiplex assessment of protein variant abundance by massively parallel sequencing. Nat. Genet. 50, 874–882 (2018).
Gietz, R. D. & Schiestl, R. H. Large-scale high-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 38–41 (2007).
Fowler, D. M., Stephany, J. J. & Fields, S. Measuring the activity of protein variants on a large scale using deep mutational scanning. Nat. Protoc. 9, 2267–2284 (2014).
Kloepper, K. D., Woods, W. S., Winter, K. A., George, J. M. & Rienstra, C. M. Preparation of α-synuclein fibrils for solid-state NMR: expression, purification, and incubation of wild-type and mutant forms. Protein Expr. Purif. 48, 112–117 (2006).
Huang, C., Ren, G., Zhou, H. & Wang, C.-C. A new method for purification of recombinant human α-synuclein in Escherichia coli. Protein Expr. Purif. 42, 173–177 (2005).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Acknowledgements
We thank D. Larsen for technical assistance. We acknowledge students of the Integrated Program in Quantitative Biology at UCSF for contributions to our analytical approach and initial models of toxicity; their findings regarding α-synuclein DMS under different environmental conditions will be described in a manuscript currently in preparation. We thank the laboratory of H. El-Samad (University of California, San Francisco) for yeast strains, the laboratory of V. M.-Y. Lee for plasmids (University of Pennsylvania) and Twist Bioscience for providing the dsDNA variant library in support of our educational efforts. This work was supported by grants from the NIH to M.K. (grant no. DP2 GM119139) and to W.F.D. (grant nos. R35-122603, P01-AG002132, R01-GM117593) and by the UCSF Program in Breakthrough Biomedical Research, which is funded in part by the Sandler Foundation, through grants to M.K. R.W.N. was supported by NIH training grant no. T32-HL007731 and a UCSF Program in Breakthrough Biomedical Research Postdoc Independent Research Grant, which is funded in part by the Sandler Foundation. Flow cytometry was supported by grant no. P30-CA082103 (NIH).
Author information
Authors and Affiliations
Contributions
R.W.N. and M.K. conceived the project. R.W.N., M.K. and W.F.D. formulated the hypotheses and designed the experiments. R.W.N., E.D.C. and M.K. designed the library and sequencing strategy. R.W.N. constructed and screened the variant library and yeast strains. R.W.N., J.T.L. and M. K. designed the cell sorting strategy. J.T.L. performed flow cytometry. E.D.C. collected next-generation sequencing data. R.W.N. purified proteins and performed circular dichroism spectroscopy and fluorescence microscopy. R.W.N. and W.F.D. developed structural models. R.W.N., M.K. and W.F.D analyzed the data and drafted the manuscript. All authors contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5
Source data
Rights and permissions
About this article
Cite this article
Newberry, R.W., Leong, J.T., Chow, E.D. et al. Deep mutational scanning reveals the structural basis for α-synuclein activity. Nat Chem Biol 16, 653–659 (2020). https://doi.org/10.1038/s41589-020-0480-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0480-6
This article is cited by
-
Deep mutational scan of a drug efflux pump reveals its structure–function landscape
Nature Chemical Biology (2023)
-
Genome-wide prediction of disease variant effects with a deep protein language model
Nature Genetics (2023)
-
Using human genetics to improve safety assessment of therapeutics
Nature Reviews Drug Discovery (2023)
-
Single residue modulators of amyloid formation in the N-terminal P1-region of α-synuclein
Nature Communications (2022)
-
From systems to structure — using genetic data to model protein structures
Nature Reviews Genetics (2022)