Abstract
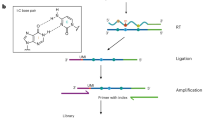
Transcriptome-wide mapping of N6-methyladenosine (m6A) at base resolution remains an issue, impeding our understanding of m6A roles at the nucleotide level. Here, we report a metabolic labeling method to detect mRNA m6A transcriptome-wide at base resolution, called ‘m6A-label-seq’. Human and mouse cells could be fed with a methionine analog, Se-allyl-l-selenohomocysteine, which substitutes the methyl group on the enzyme cofactor SAM with the allyl. Cellular RNAs could therefore be metabolically modified with N6-allyladenosine (a6A) at supposed m6A-generating adenosine sites. We pinpointed the mRNA a6A locations based on iodination-induced misincorporation at the opposite site in complementary DNA during reverse transcription. We identified a few thousand mRNA m6A sites in human HeLa, HEK293T and mouse H2.35 cells, carried out a parallel comparison of m6A-label-seq with available m6A sequencing methods, and validated selected sites by an orthogonal method. This method offers advantages in detecting clustered m6A sites and holds promise to locate nuclear nascent RNA m6A modifications.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The high-throughput sequencing data reported in this paper have been deposited in the Gene Expression Omnibus database at https://www.ncbi.nlm.nih.gov/geo/ (accession no. GSE131316). All data supporting the findings are available in the paper and Supplementary Information files.
References
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways—2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Helm, M. & Motorin, Y. Detecting RNA modifications in the epitranscriptome: predict and validate. Nat. Rev. Genet. 18, 275–291 (2017).
Li, X. Y., Xiong, X. S. & Yi, C. Q. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods 14, 23–31 (2017).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Zhao, B. X. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Bokar, J. A. et al. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N 6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997).
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N 6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N 6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Jia, G. et al. N 6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. Q. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Wang, X. et al. N 6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Xiao, W. et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61, 507–519 (2016).
Huang, H. L. et al. Recognition of RNA N 6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20, 285–295 (2018).
Edens, B. M. et al. FMRP modulates neural differentiation through m6A-dependent mRNA nuclear export. Cell Rep. 28, 845–854 (2019).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Meyer, K. D. & Jaffrey, S. R. The dynamic epitranscriptome: N 6-methyladenosine and gene expression control. Nat. Rev. Mol. Cell Biol. 15, 313–326 (2014).
Yang, Y., Hsu, P. J., Chen, Y. S. & Yang, Y. G. Dynamic transcriptomic m6A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 28, 616–624 (2018).
Shi, H. L., Wei, J. B. & He, C. Where, when and how: context-dependent functions of RNA methylation writers, readers and erasers. Mol. Cell 74, 640–650 (2019).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Molinie, B. et al. m6A-LAIC-seq reveals the census and complexity of the m6A epitranscriptome. Nat. Methods 13, 692–698 (2016).
Chen, K. et al. High-resolution N 6-methyladenosine (m6A) map using photo-crosslinking-assisted m6A sequencing. Angew. Chem. Int. Ed. 54, 1587–1590 (2015).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Ule, J. et al. CLIP identifies nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Zhang, Z. et al. Single-base mapping of m6A by an antibody-independent method. Sci. Adv. 5, eaax0250 (2019).
Garcia-Campos, M. A. et al. Deciphering the ‘m6A Code’ via antibody-independent quantitative profiling. Cell 178, 731–747 (2019).
Meyer, K. D. DART-seq: an antibody-free method for global m6A detection. Nat. Methods 16, 1275–1280 (2019).
Harcourt, E. M., Ehrenschwender, T., Batista, P. J., Chang, H. Y. & Kool, E. T. Identification of a selective polymerase enables detection of N 6-methyladenosine in RNA. J. Am. Chem. Soc. 135, 19079–19082 (2013).
Liu, N. et al. Probing N 6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA 19, 1848–1856 (2013).
Imanishi, M., Tsuji, S., Suda, A. & Futaki, S. Detection of N 6-methyladenosine based on the methyl-sensitivity of MazF RNA endonuclease. Chem. Commun. 53, 12930–12933 (2017).
Hong, T. T. et al. Precise antibody-independent m6A identification via 4SedTTP-involved and FTO-assisted strategy at single-nucleotide resolution. J. Am. Chem. Soc. 140, 5886–5889 (2018).
Liu, W. L. et al. Identification of a selective DNA ligase for accurate recognition and ultrasensitive quantification of N 6-methyladenosine in RNA at one-nucleotide resolution. Chem. Sci. 9, 3354–3359 (2018).
Xiao, Y. et al. An elongation- and ligation-based qPCR amplification method for the radiolabeling-free detection of locus-specific N 6-methyladenosine modification. Angew. Chem. Int. Ed. 57, 15995–16000 (2018).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Dalhoff, C., Lukinavicius, G., Klimasauskas, S. & Weinhold, E. Direct transfer of extended groups from synthetic cofactors by DNA methyltransferases. Nat. Chem. Biol. 2, 31–32 (2006).
Wang, R. et al. Profiling genome-wide chromatin methylation with engineered posttranslation apparatus within living cells. J. Am. Chem. Soc. 135, 1048–1056 (2013).
Shu, X. et al. N 6-allyladenosine: a new small molecule for RNA labeling identified by mutation assay. J. Am. Chem. Soc. 139, 17213–17216 (2017).
Tomkuviene, M., Mickute, M., Vilkaitis, G. & Klimasauskas, S. Repurposing enzymatic transferase reactions for targeted labeling and analysis of DNA and RNA. Curr. Opin. Biotechnol. 55, 114–123 (2019).
Hartstock, K. et al. Enzymatic or in vivo installation of propargyl groups in combination with click chemistry for the enrichment and detection of methyltransferase target sites in RNA. Angew. Chem. Int. Ed. 57, 6342–6346 (2018).
Mickute, M. et al. Animal Hen1 2′-O-methyltransferases as tools for 3′-terminal functionalization and labeling of single-stranded RNAs. Nucleic Acids Res. 46, e104 (2018).
Calabretta, A. & Leumann, C. J. Base pairing and miscoding properties of 1,N 6-ethenoadenine- and 3,N 4-ethenocytosine-containing RNA oligonucleotides. Biochemistry 52, 1990–1997 (2013).
Ottria, R., Casati, S., Baldoli, E., Maier, J. A. M. & Ciuffreda, P. N 6-alkyladenosines: synthesis and evaluation of in vitro anticancer activity. Biorg. Med. Chem. 18, 8396–8402 (2010).
Grammel, M., Luong, P., Orth, K. & Hang, H. C. A chemical reporter for protein AMPylation. J. Am. Chem. Soc. 133, 17103–17105 (2011).
Bothwell, I. R. & Luo, M. K. Large-scale, protection-free synthesis of Se-adenosyl-l-selenomethionine analogues and their application as cofactor surrogates of methyltransferases. Org. Lett. 16, 3056–3059 (2014).
Yue, Y. et al. VIRMA mediates preferential m6A mRNA methylation in 3′UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 4, 10 (2018).
Csepany, T., Lin, A., Baldick, C. J. & Beemon, K. Sequence specificity of messenger-RNA N 6-adenosine methyltransferase. J. Biol. Chem. 265, 20117–20122 (1990).
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N 6-methyladenosine by m6A-seq based on immunocapturing and massively parallel sequencing. Nat. Protoc. 8, 176–189 (2013).
Chen, S. F., Zhou, Y. Q., Chen, Y. R. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, 884–890 (2018).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ma, L. J. et al. Evolution of transcript modification by N 6-methyladenosine in primates. Genome Res. 27, 385–392 (2017).
Robinson, M. D. et al. Evaluation of affinity-based genome-wide DNA methylation data: effects of CpG density, amplification bias and copy number variation. Genome Res. 20, 1719–1729 (2010).
Li, X. Y. et al. Base-resolution mapping reveals distinct m1A methylome in nuclear- and mitochondrial-encoded transcripts. Mol. Cell 68, 993–1005 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general-purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We acknowledge support from the National Key Research and Development Program of China (2017YFA0506800), the National Natural Science Foundation of China (91853110, 21977087), the Fundamental Research Funds for the Central Universities, and Dabeinong Funds for Discipline Development and Talent Training in Zhejiang University. We thank B. Zhao at Life Sciences Institute of Zhejiang University for offering the H2.35 cell line. We thank L. Zhang at Shanghai Jiaotong University for providing FTO protein. We thank S.F. Reichard for editing the manuscript. C.H. is an investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.L. conceived the project. X.S. and J.C. designed and performed most experiments. J.C. worked on the bioinformatics analysis and Z.L., L.M. and X.C. offered guidance. M.C. and S.X. worked on the synthesis of Se-allyl-l-selenohomocysteine. Q.D., T.L. and X.Y. provided help with building in vitro biochemical assays. M.G. helped with subcloning. F.W. and Y.W. performed mass spectrometry measurements. J.L. supervised the project. J.L., X.S. and J.C. wrote the manuscript. Y.Y., C.H. and X.F. contributed to the manuscript. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.H. is a scientific founder of Accent Therapeutics and a member of its scientific advisory board. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–22 and Tables 1–4
Supplementary Data Set 1:
The detailed information of a6A peaks from the a6A-RIP sequencing in HeLa cells.
Supplementary Data Set 2:
The detailed information of a6A peaks from the a6A-RIP sequencing in HEK293T cells.
Supplementary Data Set 3:
The detailed information of a6A peaks from the a6A-RIP sequencing in H2.35 cells.
Supplementary Data Set 4:
The detailed information of m6A peaks from the m6A-RIP sequencing in HEK293T cells
Supplementary Data Set 5:
The detailed information of m6A peaks from the m6A-RIP sequencing in H2.35 cells
Supplementary Data Set 6:
The detailed information of mutation sites called from m6A-label-seq in HeLa cells.
Supplementary Data Set 7:
The detailed information of mutation sites called from m6A-label-seq in HEK293T cells.
Supplementary Data Set 8
The detailed information of mutation sites called from m6A-label-seq in H2.35 cells.
Supplementary Data Set 9
Differential gene expression results in HeLa cells under methionine versus Se-derived analog.
Rights and permissions
About this article
Cite this article
Shu, X., Cao, J., Cheng, M. et al. A metabolic labeling method detects m6A transcriptome-wide at single base resolution. Nat Chem Biol 16, 887–895 (2020). https://doi.org/10.1038/s41589-020-0526-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0526-9
This article is cited by
-
Decoding epitranscriptomic regulation of viral infection: mapping of RNA N6-methyladenosine by advanced sequencing technologies
Cellular & Molecular Biology Letters (2024)
-
RNA modifications in cellular metabolism: implications for metabolism-targeted therapy and immunotherapy
Signal Transduction and Targeted Therapy (2024)
-
GLORI for absolute quantification of transcriptome-wide m6A at single-base resolution
Nature Protocols (2024)
-
Transcriptome-wide profiling and quantification of N6-methyladenosine by enzyme-assisted adenosine deamination
Nature Biotechnology (2023)
-
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Nature Communications (2023)