Abstract
Bacterial surface polysaccharides are assembled by glycosyltransferase enzymes that typically use sugar nucleotide or polyprenyl-monophosphosugar activated donors. Characterized representatives exist for many monosaccharides but neither the donor nor the corresponding glycosyltransferases have been definitively identified for ribofuranose residues found in some polysaccharides. Klebsiella pneumoniae O-antigen polysaccharides provided prototypes to identify dual-domain ribofuranosyltransferase proteins catalyzing a two-step reaction sequence. Phosphoribosyl-5-phospho-d-ribosyl-α-1-diphosphate serves as the donor for a glycan acceptor-specific phosphoribosyl transferase (gPRT), and a more promiscuous phosphoribosyl-phosphatase (PRP) then removes the residual 5′-phosphate. The 2.5-Å resolution crystal structure of a dual-domain ribofuranosyltransferase ortholog from Thermobacillus composti revealed a PRP domain that conserves many features of the phosphatase members of the haloacid dehalogenase family, and a gPRT domain that diverges substantially from all previously characterized phosphoribosyl transferases. The gPRT represents a new glycosyltransferase fold conserved in the most abundant ribofuranosyltransferase family.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw mass spectrometry data supporting the presented figures have been uploaded to the MassIVE repository under the ID MSV000088832. The ORF4Tc structure has been deposited PDB (7SHG). PDB IDs used in the analysis of this work include 5ALN, 3VAY, 1YNS, 2OYC, 1DQN, 1FSG, 1X42 and 4LYY. Accession numbers for modeled ribosyltransferases and all orthologs are available in Supplementary Table 3. Accessions for the K. pneumoniae O4 and O7 clusters are KU310493.1 and MN173773.1, respectively.
References
Whitfield, C., Szymanski, C. M. Aebi, M. in Essentials of Glycobiology (eds Varki, A. et al.) 265–282 (Cold Spring Harbor Laboratory Press, 2017).
Whitfield, C., Wear, S. S. & Sande, C. Assembly of bacterial capsular polysaccharides and exopolysaccharides. Annu. Rev. Microbiol. 74, 521–543 (2020).
Whitfield, C. & Trent, M. S. Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 83, 199–128 (2014).
Rajagopal, M. & Walker, S. Envelope structures of Gram-positive bacteria. Curr. Top. Microbiol. Immunol. 404, 1–44 (2017).
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Aanensen, D. M., Mavroidi, A., Bentley, S. D., Reeves, P. R. & Spratt, B. G. Predicted functions and linkage specificities of the products of the Streptococcus pneumoniae biosynthetic loci. J. Bacteriol. 189, 7856–7876 (2007).
Morona, J. K., Morona, R. & Paton, J. C. Comparative genetics of capsular polysaccharide biosynthesis in Streptococcus pneumoniae types belonging to serogroup 19. J. Bacteriol. 181, 5355–5364 (1999).
Liu, B. et al. Structure and genetics of Escherichia coli O antigens. FEMS Microbiol. Rev. 44, 655–683 (2020).
Sarvas, M. & Nikaido, H. Biosynthesis of T1 antigen in Salmonella: origin of d-galactofuranose and d-ribofuranose residues. J. Bacteriol. 105, 1063–1072 (1971).
Senchenkova, S. N. et al. Structural and genetic characterization of the Shigella boydii type 10 and type 6 O antigens. J. Bacteriol. 187, 2551–2554 (2005).
Liu, B. et al. Structure and genetics of Shigella O antigens. FEMS Microbiol. Rev. 32, 627–653 (2008).
Hove-Jensen, B. et al. Phosphoribosyl diphosphate (PRPP): biosynthesis, enzymology, utilization, and metabolic significance. Microbiol. Mol. Biol. Rev. 81, e00040-16 (2017).
Kudo, F., Fujii, T., Kinoshita, S. & Eguchi, T. Unique O-ribosylation in the biosynthesis of butirosin. Bioorg. Medicinal Chem. 15, 4360–4368 (2007).
Wolucka, B. Biosynthesis of d-arabinose in mycobacteria—a novel bacterial pathway with implications for antimycobacterial therapy. FEBS J. 275, 2691–2711 (2008).
Harvey, H., Kus, J. V., Tessier, L., Kelly, J. & Burrows, L. L. Pseudomonas aeruginosa d-arabinofuranose biosynthetic pathway and its role in type IV pilus assembly. J. Biol. Chem. 286, 28128–28137 (2011).
Whitfield, C., Williams, D. M. & Kelly, S. D. Lipopolysaccharide O-antigens-bacterial glycans made to measure. J. Biol. Chem. 295, 10593–10609 (2020).
He, L. et al. ubiA (Rv8606c) encoding DPPR synthase involved in cell wall synthesis is associated with ethambutol resistance in Mycobacterium tuberculosis. Tuberculosis 95, 149–154 (2015).
Cai, L. et al. Prokaryotic expression, identification and bioinformatics analysis of the Mycobacterium tuberculosis Rv3807c gene encoding the putative enzyme committed to decaprenylphosphoryl-d-arabinose synthesis. Indian J. Microbiol. 54, 46–51 (2014).
Lu, S. et al. CDD/SPARCLE: the Conserved Domain Database in 2020. Nucleic Acids Res. 48, D265–D268 (2020).
Allen, K. N. & Dunaway-Mariano, D. Phosphoryl group transfer: evolution of a catalytic scaffold. Trends Biochem. Sci. 29, 495–503 (2004).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Read, R. J., Adams, P. D. & McCoy, A. J. Intensity statistics in the presence of translational noncrystallographic symmetry. Acta Crystallogr. Sect. D: Biol. Crystallogr. 69, 176 (2013).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Öster, L., Tapani, S., Xue, Y. & Käck, H. Successful generation of structural information for fragment-based drug discovery. Drug Discov. Today 20, 1104–1111 (2015).
Hou, Z., Zhang, H., Li, M., Chang, W. & IUCr Structure of 2-haloacid dehalogenase from Pseudomonas syringae pv. tomato DC3000. Acta Crystallogr. Sect. D: Biol. Crystallogr. 69, 1108–1114 (2013).
Wang, H., Pang, H., Bartlam, M. & Rao, Z. Crystal structure of human E1 enzyme and its complex with a substrate analog reveals the mechanism of its phosphatase/enolase activity. J. Mol. Biol. 348, 917–926 (2005).
Almo, S. C. et al. Structural genomics of protein phosphatases. J. Struct. Funct. Genomics 8, 121–140 (2007).
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Wuxian, Shi et al. Crystal structures of Giardia lamblia guanine phosphoribosyltransferase at 1.75 Å. Biochemistry 39, 6781–6790 (2000).
Héroux, A., White, E. L., Ross, L. J., Kuzin, A. P. & Borhani, D. W. Substrate deformation in a hypoxanthine-guanine phosphoribosyltransferase ternary complex: the structural basis for catalysis. Structure 8, 1309–1318 (2000).
Toukach, P. V. & Egorova, K. S. Carbohydrate structure database merged from bacterial, archaeal, plant and fungal parts. Nucleic Acids Res. 44, D1229–D1236 (2016).
Ovchinnikova, O. G. et al. Bacterial β-Kdo glycosyltransferases represent a new glycosyltransferase family (GT99). Proc. Natl Acad. Sci. USA 113, E3120–E3129 (2016).
Ovchinnikova, O. G. et al. Biochemical characterization of bifunctional 3-deoxy-β-d-manno-oct-2-ulosonic acid (β-Kdo) transferase KpsC from Escherichia coli involved in capsule biosynthesis. J. Biol. Chem. 291, 21519–21530 (2016).
Sinha, S. C. & Smith, J. L. The PRT protein family. Curr. Opin. Struct. Biol. 11, 733–739 (2001).
Brown, S., Maria, J. P. S. Jr & Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 67, 313–336 (2013).
Imperiali, B. Bacterial carbohydrate diversity—a brave new world. Curr. Opin. Chem. Biol. 53, 1–8 (2019).
Zhang, H. et al. The highly conserved domain of unknown function 1792 has a distinct glycosyltransferase fold. Nat. Commun. 5, 4339 (2014).
Kattke, M. D. et al. Structure and mechanism of TagA, a novel membrane-associated glycosyltransferase that produces wall teichoic acids in pathogenic bacteria. PLoS Pathog. 15, e1007723 (2019).
Sauvage, E., Kerff, F., Terrak, M., Ayala, J. A. & Charlier, P. The penicillin-binding proteins: structure and role in peptidoglycan biosynthesis. FEMS Microbiol. Rev. 32, 234–258 (2008).
Mann, E. & Whitfield, C. A widespread three-component mechanism for the periplasmic modification of bacterial glycoconjugates. Can. J. Chem. 94, 883–893 (2016).
Berti, F. & Adamo, R. Antimicrobial glycoconjugate vaccines: an overview of classic and modern approaches for protein modification. Chem. Soc. Rev. 47, 9015–9025 (2018).
Litschko, C., Budde, I., Berger, M. & Fiebig, T. Exploitation of capsule polymerases for enzymatic synthesis of polysaccharide antigens used in glycoconjugate vaccines. Methods Mol. Biol. 2183, 313–330 (2021).
Yates, L., Mills, D. & DeLisa, M. Bacterial glycoengineering as a biosynthetic route to customized glycomolecules. Adv. Biochemical Eng./Biotechnol. 175, 167–200 (2021).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Mirdita, M., Ovchinnikov, S. & Steinegger, M. ColabFold–making protein folding accessible to all. Preprint at bioRxiv https://doi.org/10.1101/2021.08.15.456425 (2021).
Gasteiger, E. et al. in The Proteomics Protocols Handbook (ed. Walker, J. M.) 571–607 (Humana Press Inc., 2005).
Kabsch, W. XDS. Acta Crystallogr. Sect. D: Biol. Crystallogr. 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. Sect. D: Biol. Crystallogr. 58, 1948–1954 (2002).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D: Biol. Crystallogr. 60, 2126–2132 (2004).
Wetter, M. et al. Engineering, conjugation, and immunogenicity assessment of Escherichia coli O121 O antigen for its potential use as a typhoid vaccine component. Glycoconj. J. 30, 511–522 (2012).
Acknowledgements
These studies were supported by individual Natural Sciences and Engineering Research Council Discovery Grants awarded to M.S.K. (no. RGPIN-2020-07113), T.L.L. (no. RGPIN-2018-04365) and C.W. (no. RGPIN-2020-03886). C.W. holds a Canada Research Chair, S.D.K. is a recipient of a Natural Sciences and Engineering Research Council Postgraduate Scholarship and D.M.W. received an Ontario Graduate Scholarship. We thank T. Forrester for assistance with crystal freezing and S. Colville at the Canadian Light Source for help with data collection.
Author information
Authors and Affiliations
Contributions
S.D.K. performed all of the bioinformatic and biochemical experiments, and crystallized ORF4Tc. D.M.W. contributed to the initial foundational identification and analysis of the Klebsiella enzymes. J.T.N. and T.K. synthesized the synthetic acceptors under the supervision of T.L.L. M.S.K. solved and analyzed the structure of ORF4Tc. C.W. conceived the project and oversaw the experimental work performed by S.D.K and D.M.W. S.D.K. M.S.K., T.L.L. and C.W. prepared the initial draft of the paper and all authors made contributions to the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
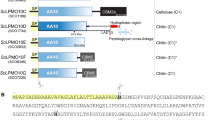
Extended Data Fig. 1 Alignment of the two domains from ORF5KpO4 and ORF7KpO7.
a, Multiple sequence alignment including PRPKpO4, PRPKpO7 and a structurally and functionally characterized HAD representative, human enolase-phosphatase E1 (PDB ID: 1ZS9)27. The predicted secondary structures of each are shown above the alignment. The PRP (HAD) domains in the O4 and O7 enzymes are provided by residues 1-214 and 1-216, respectively, and the breakpoint for creating the separated PRP-domain constructs is indicated by a vertical arrow. Active-site loops are indicated above the alignment, and an asterisk denotes the catalytic Asp residue. b, Multiple sequence alignment and secondary structures of gPRTKpO4 and gPRTKpO7. Secondary structures were predicted with JPred45, and then aligned using PROMALS3D47 and ClustalW46, respectively. Alignments in a and b were visualized with ESPript. Bars and arrows represent α-helices and β-strands, respectively.
Extended Data Fig. 2 Identification of ORF4Tc from Thermobacillus composti and biochemical analysis of its PRP activity.
a, Organization of the glycan biosynthesis gene cluster. The pathway defining wzm/wzt genes (orange) encode the subunits of the ABC transporter. orf4Tc (purple) encodes the dual-domain PRP-gPRT protein. The general functions encoded by the remaining orfs (gray) are predicted by BLAST analyses. orf3 encodes a putative methyltransferase, while orf5 and orf6 appear to encode glycosyltransferases belonging to the GT4 and GT2 families, respectively. b, In vitro reactions containing ORF5KpO4 D10A and ORF4Tc with 2 demonstrate the ability of ORF4Tc to dephosphorylate the ORF5KpO4 D10A product. Products were separated by HPLC, and the product structures are shown on chromatograms with a black square representing the methoxybenzamide tag in 2. The product identities were established by comparison of HPLC retention times to known compounds, authenticated by MS. The analysis was performed in triplicate with similar results.
Extended Data Fig. 3 Representative structural homologs of the PRP and gPRT domains.
All structures were superimposed on ORF4Tc. Models are colored using rainbow (N-terminus dark blue, C-terminus red). a, PRP domain of ORF4Tc. b, Protein of unknown function from Pyrococcus horikoshii (1 × 42). c, PRP domain of human soluble epoxide hydrolase (5aln). d, Pseudomonas syringae haloacid dehalogenase (3vay). e, Human enolase phosphatase 1 (1yns). f, Human chronophin (2oyc), a bifunctional pyridoxal phosphate and cofilin phosphatase (note that this HAD lacks a helical cap domain, and has a second Rossmann domain fused instead). g, The gPRT domain of ORF4Tc h, Giardia lamlia guanine phosphoribosyltransferase (1dqn) i, Shewanella pealeana hypoxanthine phosphoribosyltransferase (4lyy). j, Toxoplasma gondii hypoxanthine-guaninephosphoribosyltransferase (1fsg).
Extended Data Fig. 4 Metal ion-dependence of PRP activity in ORF7KpO7.
in vitro reactions demonstrating the Mg2+ requirements for activity of the ORF7KpO7 PRP and gPRT domains. To demonstrate the Mg2+ dependent activity of the gPRT, ORF7KpO7 D28A was tested in a reaction lacking added Mg2+. For the PRP, initial reactions were conducted with ORF7KpO7 D28A in the presence of 25 mM Mg2+ to allow the gPRT to transfer Rib-P, and the reactions were then spiked with 50 mM EDTA before PRPO7 was added. HPLC analysis detected only a trace amount of dephosphorylated substrate, consistent with PRP activity being dependent on Mg2+. All of the reactions were incubated for 15 mins and aliquots of the reaction mixtures were separated by HPLC on a GlycoSep N column eluted with a normal phase gradient of water. The structures of donors and products are shown on each chromatogram, and the black square in the structure represents the methoxybenzamide tag in 1. The structures were established by comparison to the HPLC retention times of products in validated control reactions; ORF7KpO7 and ORF7KpO7 D28A transfer of Ribf or Ribf-P to 1, respectively. The analysis was performed in triplicate with similar results.
Extended Data Fig. 5 Atomic displacement parameters (B-factors) mapped onto the structure.
The most ordered regions of the structure are blue, while those with elevated B-factors tend towards more red colours. Note that the hood domain of the gPRT has the highest overall B-factors, and that these increase towards the tip. This suggests that the hood domain may be mobile enough to close over the active site during catalysis.
Extended Data Fig. 6 Activities of gPRTKpO7 site-directed alanine-replacement mutants.
a, The chromatograms show the profiles of aliquots of reaction mixtures separated by HPLC on a GlycoSep N column eluted with a normal phase gradient of water. Each reaction was performed with the indicated mutant, PRPP donor and acceptor 1. After 1 h (left column) or 18 h (right column), reactions were stopped by the addition of acetonitrile. R284A and R311A mutant proteins were inactive, while D285A and D417A showed substantially reduced activity. D285A showed no detectable activity at 1 h but retained small amounts of activity evident after 18 h. D417A converted almost half of the substrate after 1-hour reaction and full conversion after 18 hours. The identities of the reaction products are shown on each chromatogram. The black box represents the methoxybenzamide tag in 1. The analysis was performed in triplicate with similar results. b, The identities of the reaction products from 18 h reactions for the active variants were confirmed by mass spectrometry. The extracted ion chromatogram (EIC) traces of m/z 704.345 (1 + Ribf) are shown, together with expanded regions of the corresponding ESI mass-spectrum at the indicated retention time.
Extended Data Fig. 7 Alphafold modeling of the gPRT domain of ORF7KpO7, ORF5KpO4 and selected orthologs (as indicated).
The source bacterium and the structure of the corresponding Ribf-containing glycans are shown. For details on accessions and keys to the glycose structures see Supplementary Table 3. Models are coloured with rainbow (N-terminus dark blue, C-terminus red).
Supplementary information
Supplementary Information
Supplementary Tables 1–4, Figs. 1–5, experimental procedures and synthetic schemes of 1 and 2.
Rights and permissions
About this article
Cite this article
Kelly, S.D., Williams, D.M., Nothof, J.T. et al. The biosynthetic origin of ribofuranose in bacterial polysaccharides. Nat Chem Biol 18, 530–537 (2022). https://doi.org/10.1038/s41589-022-01006-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01006-6
This article is cited by
-
Basket of sweets
Nature Chemical Biology (2023)
-
A multi-enzyme machine polymerizes the Haemophilus influenzae type b capsule
Nature Chemical Biology (2023)