Abstract
T cell exhaustion presents one of the major hurdles to cancer immunotherapy. Among exhausted CD8+ tumor-infiltrating lymphocytes, the terminally exhausted subset contributes directly to tumor cell killing owing to its cytotoxic effector function. However, this subset does not respond to immune checkpoint blockades and is difficult to be reinvigorated with restored proliferative capacity. Here, we show that a half-life-extended interleukin-10–Fc fusion protein directly and potently enhanced expansion and effector function of terminally exhausted CD8+ tumor-infiltrating lymphocytes by promoting oxidative phosphorylation, a process that was independent of the progenitor exhausted T cells. Interleukin-10–Fc was a safe and highly efficient metabolic intervention that synergized with adoptive T cell transfer immunotherapy, leading to eradication of established solid tumors and durable cures in the majority of treated mice. These findings show that metabolic reprogramming by upregulating mitochondrial pyruvate carrier-dependent oxidative phosphorylation can revitalize terminally exhausted T cells and enhance the response to cancer immunotherapy.
Similar content being viewed by others
Main
Cancer immunotherapy represented by immune checkpoint blockades (ICBs) has achieved remarkable clinical success1. However, an outstanding challenge remains that a great majority of patients fail to respond to this therapy2,3,4. The low response rate is in part due to the fact that tumor-infiltrating lymphocytes (TILs) become exhausted and eventually incapable to control tumor progression5,6,7. Two distinct subsets of exhausted CD8+ TILs were recently identified with different functional properties8,9,10,11. One of the subsets, termed ‘progenitor exhausted’ (TCF-1+TIM-3−) CD8+ T cells, shows relatively high proliferative capacity and the capability to differentiate into ‘terminally exhausted’ (TCF-1−TIM-3+) CD8+ TILs, the other subpopulation that directly contributes to the killing of tumor cells owing to its superior cytotoxicity to the progenitor exhausted TILs. Progenitor exhausted CD8+ T cells can respond to anti-PD-1 checkpoint blockade therapy and mediate tumor growth control8,9,10. However, terminally exhausted CD8+ TILs, a subset with impaired proliferative capacity, do not respond to ICBs or most existing immunotherapies8,12,13,14. Therefore, it remains a major challenge to reinvigorate the terminally exhausted subpopulation of CD8+ TILs in the tumor microenvironment (TME) and exploit its therapeutic potential.
Metabolic restriction imposed in the TME, such as glucose deprivation and hypoxia, greatly alters the cell signaling of TILs leading to impaired anti-tumor immune responses15,16. It has been reported that exhausted T cells exhibit suppressed mitochondrial respiration and/or glycolysis and such poor metabolic fitness may reinforce T cell exhaustion17,18,19,20. Metabolic interventions that could enhance the effector function and proliferative capacity of exhausted T cells are thereby being actively pursued. We and others have shown that maintaining mitochondrial fitness restores the proliferation and effector function of exhausted T cells leading to enhanced anti-tumor immunity18,19,20. Interleukin-10 (IL-10) is a pleiotropic cytokine that can promote anti-tumor immunity in multiple murine tumor models21,22,23. Recently, patients treated with PEGylated IL-10 (pegilodecakin) showed increased numbers of PD-1- and LAG-3-positive CD8+ T cells in the circulation24. In addition, IL-10 has been shown to enhance the mitochondrial oxidative phosphorylation (OXPHOS) of macrophages25. However, whether IL-10 could also reprogram T cell metabolic profiles and restore the function of exhausted T cells remains unexplored.
Here, we report that a half-life-extended IL-10–Fc fusion protein directly expanded terminally exhausted CD8+ TILs and promoted their effector function in a way independent of progenitor exhausted CD8+ TILs, leading to eradication of established solid tumors and durable cures in a majority of treated mice when combined with adoptive T cell transfer (ACT) or ICB immunotherapy. Our results provide preclinical evidence that IL-10–Fc is a safe and highly effective therapy that acts on a specific subset of CD8+ TILs distinct from those responding to ICBs. Thus, IL-10–Fc could potentially complement and synergize with many existing cancer immunotherapies for enhanced efficacy and response rates. Furthermore, we found that IL-10–Fc reprogramed T cell metabolism by promoting OXPHOS through the mitochondrial pyruvate carrier (MPC) and such metabolic reprogramming was essential for reactivating terminally exhausted CD8+ TILs and enhancing the ultimate therapeutic outcome by IL-10–Fc. These findings provide insight into the crucial role of metabolic profiles in T cell exhaustion and reinvigoration and lay the foundation for further identification of metabolic switches for regulating T cell activities in the TME.
Results
IL-10–Fc reinvigorates terminally exhausted CD8+ TILs
We first produced a recombinant half-life-extended fusion protein of human IL-10 and IgG1 Fc (IL-10–Fc), which could cross-react with mouse IL-10 receptor (IL-10R)26 in a dose-dependent manner (Extended Data Fig. 1a–d). To treat subcutaneous (s.c.) B16F10 tumors, we transferred PMEL CD8+ T cells (5 × 106) that recognize the gp100 cognate antigen to mice through intravenous (i.v.) injection adjuvanted by peritumorally administered IL-10–Fc or phosphate-buffered saline (PBS) as control. B16F10 melanoma is a poorly immunogenic tumor with very few lymphocyte infiltrates27,28. ACT of tumor-antigen-specific PMEL CD8+ T cells greatly enhanced tumor infiltration of total CD45.2+ TILs and CD3+ T cells (Extended Data Fig. 1e,f), providing the basis for us to assess the effects of IL-10–Fc on exhausted TILs. We found that the treatment of IL-10–Fc combined with ACT markedly increased the number of CD3+ TILs, particularly CD8+ T cells, as compared with ACT alone (Fig. 1a). Interestingly, IL-10–Fc treatment induced no notable alteration in counts or inflammatory properties of other lymphocytes or myeloid cells in the TME except that inhibition of dendritic cell maturation was noticed (Extended Data Fig. 1g–i).
Thy1.2+ C57BL/6 mice were inoculated subcutaneously with B16F10 tumor cells (1 × 106) and received i.v. adoptive transfer of activated Thy1.1+ PMEL CD8+ T cells (5 × 106) on day 6 followed by peritumoral (p.t.) administration of IL-10–Fc (20 µg) or PBS control every other day until day 12. On day 14, mice were killed and tumors were processed and analyzed by flow cytometry. Data are one representative of three or four independent experiments and n = 4 independent animals unless otherwise noted. a, Counts of CD8+ TILs in tumors. Shown are pooled data of two independent experiments (n = 10 independent animals). b, Gating strategy for terminally exhausted CD8+ TILs using surface markers PD-1 and TIM-3. c, Representative flow cytometry plots showing the frequencies of PD-1+TIM-3+ terminally exhausted CD8+ T cells among all CD44+CD8+ TILs. d, Counts of three subpopulations among endogenous and PMEL CD8+ TILs (n = 5 independent animals). e, Frequencies of Ki67+BrdU+ T cells among each subpopulation of CD8+ TILs. All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test. NS, not significant (P > 0.05).
We next determined the specific subset(s) of CD8+ TILs that responded to IL-10–Fc treatment. Among all the CD8+ TILs, terminally exhausted CD8+ T cells were substantially expanded, whereas the frequencies and counts of progenitor exhausted CD8+ T cells (TCF-1+TIM-3−) remained unchanged or slightly decreased (Extended Data Fig. 2a–c). We noticed that the TCF-1−TIM-3+ subpopulation was completely in line with the PD-1+TIM-3+ double-positive subset (Fig. 1b), which also showed reduced polyfunctionality (Extended Data Fig. 2d). Such observation was consistent with that in chronic infections12,13,14. To simplify the staining procedures and analyses, we next used surface inhibitory markers, PD-1 and TIM-3, to define the terminally exhausted subpopulation in the following experiments. IL-10–Fc treatment markedly and selectively expanded the PD-1+TIM-3+ double-positive but not the PD-1+ single-positive subset, with substantially increased frequencies as well as 10.5- and 3.3-fold greater cell counts (combination treatment versus ACT alone) of adoptively transferred PMEL and endogenous CD8+ T cells, respectively (Fig. 1c,d). The PD-1+TIM-3+ T cells also exhibited enhanced BrdU incorporation and Ki67 expression when treated with IL-10–Fc, suggesting increased proliferative capacity (Fig. 1e).
Consistently, we observed the highest expression level of IL-10R subunit alpha (IL-10Rα) on the PD-1+TIM-3+CD8+ T cells among all the subsets of CD8+ TILs (Fig. 2a). In addition, IL-10Rα-knockout (IL-10Rα-KO) P14 CD8+ T cells failed to respond to IL-10–Fc treatment for cell expansion in vitro and in vivo (Fig. 2b and Extended Data Fig. 2e–g), suggesting IL-10–Fc signals directly through IL-10R on T cells. Importantly, the expanded PD-1+TIM-3+CD8+ TILs retained their superior capacity in producing granzyme B, a key cytotoxic molecule, and other effector molecules (Fig. 2c,d), and showed higher expression level of activation marker CD69 (Fig. 2e) as well as reduced expression of inhibitory marker PD-1 (Fig. 2f), a phenotype that was also observed in Tox-mutant exhausted CD8+ T cells29. Altogether, IL-10–Fc promoted proliferation and effector function of terminally exhausted CD8+ TILs through IL-10R on T cells.
The experimental setting was the same as descried in Fig. 1. Data are one representative of three or four independent experiments. a, Representative flow cytometry plots (upper) and mean fluorescence intensity (MFI, bottom) showing IL-10Rα expression on three subpopulations of CD8+ TILs (n = 6 independent animals). FMO, fluorescence minus one. b, CD45.2+ Rosa26-Cas9-knock-in (Cas9-KI) mice were inoculated subcutaneously with B16-gp33 tumor cells (2 × 105) and received an i.v. adoptive cotransfer of activated CD45.1+CD45.2+ P14 CD8+ T cells (Ctrl, 2 × 106) and activated CD45.1+ IL-10Rα-KO P14 CD8+ T cells (2 × 106) on day 10 followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 16. On day 18, mice were killed and tumors were processed and analyzed by flow cytometry. Data are pooled from two independent experiments (n = 9 independent animals). Shown are frequencies of Ctrl P14 T cells and IL-10Rα-KO P14 T cells among all CD45+ TILs. c, Frequencies of granzyme B+CD8+ T cells among each subpopulation of CD8+ TILs (n = 5 independent animals). d, Frequencies of granzyme B+IFNγ+TNFα+ polyfunctional cells among PD-1+TIM-3+CD8+ TILs (n = 7 independent animals). e, MFI of CD69 expressed on PD-1+TIM-3+CD8+ TILs (n = 5 independent animals). f, Representative flow cytometry histograms showing PD-1 expression level and MFI of PD-1 in endogenous and PMEL PD-1+TIM-3+CD8+ TILs (n = 5 independent animals). All data represent the mean ± s.e.m. and are analyzed by one-way ANOVA and Tukey’s test or two-sided Student’s t-test. NS, not significant (P > 0.05).
IL-10–Fc directly expands terminally exhausted CD8+ TILs
Notably, the expansion of PD-1+TIM-3+ double-positive CD8+ TILs by IL-10–Fc was antigen-dependent. In a cotransfer experiment, PMEL and OT-I (T cell receptor (TCR) transgenic T cells recognizing ovalbumin (OVA) antigen) CD8+ T cells were coadministered in mice bearing B16F10 tumors (Fig. 3a). The frequency of PD-1+TIM-3+ double-positive cells and total transferred cells of PMEL but not OT-1 T cells were markedly increased by IL-10–Fc (Fig. 3b–d). The results suggest that only the antigen-specific (PMEL) terminally exhausted CD8+ TILs respond to IL-10–Fc treatment. To examine whether terminally exhausted T cells responded to IL-10–Fc treatment directly, we next transferred the two subsets (PD-1+TIM-3− and PD-1+TIM-3+) of PMEL T cells sorted from CD8+ TILs to recipient mice bearing B16F10 tumors. We found the PD-1+TIM-3+CD8+ T cells were prominently expanded in vivo in the absence of PD-1+TIM-3−CD8+ T cells (Extended Data Fig. 3a,b). Ex vivo culture of the two sorted subsets separately confirmed that the increased PD-1+TIM-3+ population upon IL-10–Fc treatment was contributed mainly by direct expansion of the terminally exhausted T cells rather than the conversion from progenitor exhausted T cells (Extended Data Fig. 3c–f). In addition, the treatment of IL-10–Fc showed negligible effect on the apoptosis of CD8+ TILs, indicating the increased count of PD-1+TIM-3+CD8+ T cells was not due to reduced cell apoptosis either (Extended Data Fig. 3g).
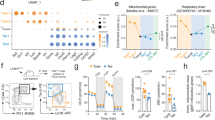
a–d, Thy1.2+ C57BL/6 mice were sublethally lymphodepleted and received adoptive cotransfer of Thy1.1+ naive PMEL and CD45.1+ naive OT-I CD8+ T cells. The mice were then inoculated with B16F10 tumor cells. On day 10, the mice were treated with adoptive transfer of activated Thy1.2+ PMEL CD8+ T cells followed by administration of IL-10–Fc or PBS control. On day 17, mice were killed for flow cytometry analyses of TILs (n = 7 independent animals). a, Experimental timeline. b, Representative flow cytometry plots showing the frequencies of the PD-1+TIM-3+ subpopulation among OT-I or PMEL (Thy1.1+) CD8+ TILs. c, Frequencies of transferred OT-I or PMEL (Thy1.1+) cells among total CD8+ TILs. d, Frequencies of PD-1+TIM-3+ subpopulation among total transferred OT-I or PMEL (Thy1.1+) CD8+ TILs. e,f, CD45.1+CD45.2+ C57BL/6 mice were inoculated with B16-gp33 tumor cells and sublethally lymphodepleted by total body irradiation. Mice received i.v. adoptive transfer of activated CD45.2+Tcf7DTR-GFP P14 CD8+ T cells followed by administration of IL-10–Fc or PBS control. To deplete TCF-1+ P14 CD8+ T cells, DT was given by intraperitoneal (i.p.) injection. On day 25, all mice were killed and tumors were processed and analyzed by flow cytometry (n = 5 independent animals). e, Experimental timeline. f, Frequencies of terminally exhausted (Tcf7DTR-GFP−PD-1+TIM-3+) P14 CD8+ T cells among all CD8+ TILs. g,h, C57BL/6 mice were inoculated with B16-gp33 tumor cells and received adoptive transfer of activated Tcf7DTR-GFP P14 CD8+ T cells followed by p.t. administration of IL-10–Fc or PBS control. DT was given intraperitoneally to deplete TCF-1+ P14 CD8+ T cells. g, Experimental timeline. h, Average tumor growth curves of each treatment group (n = 5 independent animals). Shown in parentheses is the number of long-term-surviving mice among the total number of mice in the group. All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test (c, d) or one-way ANOVA and Tukey’s test (f) or two-way ANOVA (h). NS, not significant (P > 0.05).
To confirm the above findings, we next exploited Tcf7DTR-GFP transgenic P14 T cells that allow a selective depletion of the progenitors (TCF-1+) by diphtheria toxin (DT) treatment9. With the progenitors depleted in vivo (Extended Data Fig. 3h,i), the terminally exhausted subset (Tcf7DTR-GFP−PD-1+TIM-3+ CD8+) in tumor still responded to the IL-10–Fc treatment and was expanded to a comparable level as that in mice without DT depletion, suggesting the expansion of terminally exhausted T cells by IL-10–Fc treatment is independent of the progenitors (Fig. 3e,f). To determine the direct contribution of terminally exhausted T cells in tumor growth control, we transferred Tcf7DTR-GFP transgenic P14 T cells recognizing gp33 antigen to mice bearing B16-gp33 tumors (Fig. 3g,h). With DT-mediated depletion of the progenitor exhausted T cells, the combination therapy of ACT and IL-10–Fc induced similar tumor regression and curative responses to that without depletion, implying the terminally exhausted T cells responded directly to IL-10–Fc for enhanced anti-tumor efficacy. Therefore, IL-10–Fc expanded the antigen-specific terminally exhausted CD8+ TILs in a progenitor exhausted cell-independent manner.
IL-10–Fc promotes OXPHOS of T cells
Inspired by the observation that IL-10 reprograms the metabolic profiles of macrophages25, we next examined whether IL-10/IL-10R signaling regulates CD8+ T cell metabolism to elucidate the underlying mechanism. Upon TCR stimulation with peptide-pulsed antigen presenting cells or a dimerized anti-CD3 antibody, both naive and primed CD8+ T cells upregulated OXPHOS in the presence of IL-10–Fc, evidenced by higher basal oxygen consumption rate (OCR) (Fig. 4a,b). To mimic the persistent tumor-antigen stimulation in the TME, we set up a coculture system of B16F10 cells and activated PMEL CD8+ T cells. Both basal and maximal OCRs of CD8+ T cells were elevated markedly upon IL-10–Fc treatment, whereas extracellular acidification rate (ECAR) remained stable (Fig. 4c,d and Extended Data Fig. 4a,b). The ratios of OCR to ECAR were also prominently increased (Fig. 4e). These results suggest that IL-10–Fc actively reprograms T cell metabolism toward higher OXPHOS dependence when there is persistent antigen stimulation. Such antigen dependence for metabolic reprogramming was consistent with the observed antigen-dependent CD8+ TIL expansion in vivo. Furthermore, the T cell proliferation was greatly enhanced by IL-10–Fc, and so was the killing efficiency against B16F10 target cells in the coculture experiment (Fig. 4f,g), providing evidence that the promoted OXPHOS may lead to enhanced proliferative capacity and cytotoxicity of CD8+ T cells.
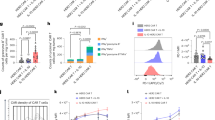
a, Basal OCR of naive PMEL CD8+ T cells from splenocytes activated with hgp100 peptide in the presence or absence of IL-10–Fc for 2 d. Data are one representative of three independent experiments (n = 4 independent samples). b, Basal OCR of primed PMEL CD8+ T cells in resting phase (day 7) restimulated by dimerized anti-CD3 (α-CD3) antibody in the presence or absence of IL-10–Fc for 1 d. Data are one representative of three independent experiments (n = 6 independent samples). c–g, Primed PMEL CD8+ T cells in the resting phase were cocultured with B16F10 tumor cells at a ratio of 1 to 1 in the presence or absence of IL-10–Fc for 2 d. CD8+ T cells were isolated for Seahorse assay and flow cytometry analyses. Data are one representative of at least three independent experiments (n = 3 independent samples). c, Real-time analysis of OCR. d, Average basal and maximal OCR. e, Ratios of OCR to ECAR. f,g, Counts of CD8+ T cells (f) and tumor cells (g) in the coculture assay. h–k, PMEL PD-1+TIM-3+CD8+ T cells were sorted from in vitro culture with restimulation by dimerized α-CD3 antibody for 1 d in the presence or absence of IL-10–Fc (n = 3 independent samples). h, Real-time analysis of OCR. i, Average maximal OCR. j, Ratios of maximal OCR to ECAR. k, Sorted PMEL PD-1+TIM-3+CD8+ T cells were cocultured with B16F10 cells at a ratio of 1:1 in the presence or absence of IL-10–Fc for 2 d. Shown are average PD-1+TIM-3+CD8+ T cell counts. Data are one representative of two independent experiments (n = 3 independent samples). l,m, Control P14 CD8+ T cells with WT IL-10Rα or IL-10Rα-KO P14 CD8+ T cells were restimulated by dimerized α-CD3 antibody in the presence or absence of IL-10–Fc and cultured for 2 d. Shown are average basal (l) and maximal (m) OCR. Data are one representative of two independent experiments (n = 5 independent samples). All data represent the mean ± s.e.m. and are analyzed by one-way ANOVA and Tukey’s test or two-sided Student’s t-test. NS, not significant (P > 0.05). FCCP, carbonyl cyanide 4- (trifluoromethoxy)phenylhydrazone; R/A, rotenone and antimycin A.
To recapitulate the terminal exhaustion phenotype in vivo, we next produced PD-1+TIM-3+CD8+ T cells ex vivo through over stimulation with a dimerized anti-CD3 antibody (Extended Data Fig. 5a). A sorted subpopulation of PD-1+TIM-3+CD8+ T cells with a good number could hence be obtained, which had been confirmed with the TCF-1−TIM-3+ phenotype (Extended Data Fig. 5b). Ex vivo produced and sorted PMEL PD-1+TIM-3+CD8+ T cells showed similarly elevated levels of OCR and ratio of OCR to ECAR as the whole CD8+ T cell population (Fig. 4h–j), and were markedly expanded by IL-10–Fc (Fig. 4k). Using IL-10Rα-KO CD8+ T cells over stimulated by the dimerized anti-CD3 antibody, we found the metabolic reprogramming of T cells by IL-10–Fc was indeed dependent on IL-10R on T cells (Fig. 4l,m).
To extend this observation to human T cells, we cocultured human chimeric antigen receptor (CAR)-T cells that targeted human epidermal growth factor receptor 2 (HER2) with ME275 human melanoma cells expressing HER2 (ME275-HER2) or HER2-positive SKOV3 human ovarian cancer cells (SKOV3-HER2). We found human CAR-T cells responded to IL-10–Fc treatment similarly to mouse T cells exhibiting enhanced OCR, proliferation and killing efficiency against target cells (Extended Data Fig. 6a–c). Human PD-1+LAG-3+CD8+ T cells induced by over stimulation of an anti-human CD3 antibody expressed the highest level of IL-10Rα among all the subsets of CD8+ T cells and responded to IL-10–Fc treatment in a similar way to mouse PD-1+TIM-3+CD8+ T cells (Extended Data Fig. 6d–f). Altogether, IL-10–Fc promoted OXPHOS of mouse and human T cells leading to enhanced proliferative capacity and cytotoxicity.
IL-10–Fc potentiates ACT and ICB immunotherapies
The discovery that IL-10–Fc expands the terminally exhausted CD8+ TILs while promoting their effector function motivated us to assess the anti-solid-tumor efficacy of ACT immunotherapies in combination with IL-10–Fc. In a therapeutic setting with B16F10 tumor, a poorly immunogenic and highly aggressive mouse melanoma model, the combination therapy of peritumorally administered IL-10–Fc and adoptive transfer of activated PMEL CD8+ T cells (5 × 106) without lymphodepletion preconditioning consistently induced complete tumor regression and durable cures in 90% of treated mice (Fig. 5a,b and Extended Data Fig. 7a). In contrast, ACT treatment alone transiently controlled tumor growth but failed to induce tumor regression. Administration of IL-10–Fc in the absence of ACT eradicated 30% of tumors, suggesting the need of exogenous tumor-reactive T cells to achieve superior efficacy. In line with the potent anti-tumor efficacy, combination therapy of ACT and IL-10–Fc induced a remarkably high ratio of CD8+ T cells to regulatory T (Treg) cells and high frequency of cytotoxic and polyfunctional cells among both endogenous and transferred CD8+ T cells (Extended Data Fig. 7b,c). Furthermore, close to 80% of long-term survivors treated with IL-10–Fc plus ACT rejected a rechallenge of B16F10 cells 2 months post cessation of therapy, indicating the induction of anti-tumor immune memory (Fig. 5c). Notably, IL-10–Fc administration with even 50% lower dosing frequency led to similar anti-tumor efficacy with 80% of B16F10 tumors cleared when combined with ACT (Extended Data Fig. 7d–h).
Thy1.2+ C57BL/6 mice were inoculated subcutaneously with B16F10 melanoma cells (5 × 105), YUMM1.7-OVA melanoma cells (1 × 106) or MC38-HER2 colon adenocarcinoma cells (1 × 106) and received i.v. adoptive cell transfers of activated Thy1.1+ PMEL CD8+ T cells (5 × 106), OT-I CD8+ T cells (5 × 106) or HER2 CAR T cells (5 × 106), respectively, on day 6, followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 20. Mice receiving injections of PBS control or IL-10–Fc (20 µg × 8) only served as controls in the B16F10 model. Mice receiving injections of PBS control or untransduced T cells served as controls in the MC38-HER2 model. Shown are average tumor growth curves (a, d, g) and survival curves (b, e, h) of each treatment group in different tumor models. Red arrows indicate the start of treatment. Shown are numbers of long-term-surviving mice among the total number of mice in the group (b, e, h). Survivors from treatment groups of combination of ACT therapies and IL-10–Fc in three different models were rechallenged subcutaneously with B16F10 (1 × 105), YUMM1.7-OVA (5 × 105) or MC38-HER2 (1 × 106) cells, respectively, on day 90 post primary inoculation. Naive WT mice (n = 5) were inoculated with the same number of tumor cells as controls. Shown are survival curves (c, f, i) and numbers of long-term-surviving mice against the re-challenges. a–c, Therapy study with B16F10 model. Shown are pooled data of two independent experiments (n = 10 independent animals). d–f, Therapy study with YUMM1.7-OVA model. Shown is one representative of three independent experiments (n = 8 independent animals). g–i, Therapy study with MC38-HER2 model. Shown are pooled data of two independent experiments (n = 6 independent animals). Data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test for tumor growth data and log-rank test for survival curves. NS, not significant (P > 0.05).
To test the robustness of this combination therapy, we next assessed whether large established solid tumors could be controlled. OVA-expressing YUMM1.7 mouse melanoma cells (YUMM1.7-OVA) were inoculated subcutaneously and allowed to grow to high tumor burden (size ≥ 50 mm2 or 150 mm3)30,31 before the initiation of therapy. Combined treatment of ACT of OT-I CD8+ T cells (5 × 106) and IL-10–Fc led to remarkable tumor regression and eventually elimination, while ACT alone showed only transient tumor growth inhibition without durable therapeutic effect (Fig. 5d). In addition, more than 60% of mice treated with IL-10–Fc and ACT combination therapy exhibited long-term survival and 100% of the survivors rejected a rechallenge of YUMM1.7-OVA cells (Fig. 5e,f). We next extended this combination strategy to CAR-T cell therapy, which is an important immunotherapy modality in the clinic32. We prepared mouse CAR-T cells that targeted HER2 and inoculated mice with MC38-HER2 tumor, a murine colon adenocarcinoma stably transfected with HER2. Transfer of HER2-targeting CAR-T cells (5 × 106) alone to mice resulted in minimum therapeutic effect (Fig. 5g,h). By contrast, ACT of CAR-T cells adjuvanted by IL-10–Fc completely eradicated tumors and led to durable cures in ~90% of treated mice. Notably, 100% of the cured animals that rejected primary tumors after CAR-T cells and IL-10–Fc combination therapy also rejected a second challenge of MC38-HER2 cells (Fig. 5i). Lymphoreplete models were used in all the ACT therapeutic studies above-mentioned, permitting the harnessing of the host anti-tumor immunity and overcoming the need for preconditioning, a procedure that is typically necessary for ACT therapy in the clinic but excludes many patients due to life-threatening toxicities33. Importantly, unlike other immune stimulatory cytokines, such as IL-2 (ref. 34) or IL-15 (ref. 35), IL-10–Fc treatment alone or in combination with ACT was safe and exhibited no overt toxicities. All the treated mice showed no body weight loss or elevation of serum levels of liver enzymes (Extended Data Fig. 7i–k). In addition, we found the combination of IL-10–Fc and anti-PD-1 eradicated established tumors and induced durable immune protection in a mouse CT26 colorectal tumor model (Extended Data Fig. 8). Thus, IL-10–Fc markedly and safely enhanced ACT and ICB immunotherapies, leading to robust and complete responses and durable cures in a majority of mice with established solid tumors.
IL-10–Fc enhances OXPHOS of terminally exhausted CD8+ TILs
To understand how IL-10–Fc treatment regulates the gene expression of terminally exhausted CD8+ TILs in vivo, we performed an RNA-sequencing (RNA-seq) analysis of sorted tumor-antigen-specific Thy1.1+PD-1+TIM-3+CD8+ TILs from B16F10 tumors treated with ACT in combination with IL-10–Fc or PBS control (Fig. 6a). IL-10–Fc treatment led to significant changes in 842 genes (adjusted P values < 0.05) in the PD-1+TIM-3+ subpopulation of CD8+ T cells in tumors (Fig. 6b and Extended Data Fig. 9a,b). The PD-1+TIM-3+CD8+ TILs in the IL-10–Fc-treated group showed higher expression of genes encoding complexes for electron transport chain (ETC), that is, OXPHOS related36, including Atp6v1g1, Cox5a, Cox8a, Ndufa5 and Ndufv3, as well as those encoding cytotoxic molecules including Gzmb, Klra2, Klrc2, Nkg7 and Tnfsf10, but lower expression of genes encoding inhibitory receptors and exhaustion transcription factors such as Cd200r1 and Nr4a2 compared with control cells (Fig. 6c). The results were consistent with the observed phenotypes of IL-10–Fc-treated PD-1+TIM-3+CD8+ TILs and their metabolic profiles in vitro. In addition, gene set enrichment analysis (GSEA) and ingenuity pathway analysis (IPA) of the transcriptional differences between IL-10–Fc and PBS control groups revealed strong enrichment of gene signatures and pathways associated with T cell OXPHOS and effector function (Fig. 6d,e and Extended Data Fig. 9c,d).
a–e, Thy1.2+ C57BL/6 mice were sublethally lymphodepleted and received adoptive transfer of Thy1.1+ naive PMEL CD8+ T cells. Mice were inoculated with B16F10 tumor cells and were treated with adoptive transfer of activated Thy1.2+ PMEL CD8+ T cells followed by administration of IL-10–Fc or PBS control. On day 18, tumor-infiltrating Thy1.1+PD-1+TIM-3+CD8+ T cells were sorted for profiling gene expression with RNA-seq (n = 2 for PBS group, n = 4 for IL-10–Fc group). a, Experimental timeline. b, Mean average plot of the RNA-seq dataset (graph shows fold change in expression (log2 (IL-10–Fc/PBS)) versus mean expression of genes (log10 (base mean)) with mean expression ≥10, and |log2(IL-10–Fc/PBS)| ≤ 7). Significantly upregulated or downregulated genes are shown in orange and blue, respectively (false discovery rate (FDR)-adjusted P value < 0.05 and |log2(fold change)| > 1). Two-tailed Wald statistic was used; adjustment for multiple comparisons (Padj): Benjamini–Hochberg (BH)-adjusted P values. c, Heat map illustrating the average transcript expression of the indicated genes. Rows represent averaged z-scores. Asterisks represent transcripts with significant differential expression by DESeq2 (FDR < 0.05). d,e, Enrichment of gene signatures from MSigDB by gene set permutation test. Enrichment score is calculated based on a weighted Kolmogorov–Smirnov-like statistic test. Adjustment for multiple comparison (FDR Q value): BH-adjusted P values. GSEAs of OXPHOS (d) and effector function (e) were performed to compare PD-1+TIM-3+CD8+ TILs treated with IL-10–Fc versus PBS. f, The experimental setting was the same as that shown in Fig. 1. Shown are frequencies of MitoSOX+CD8+ T cells among each subpopulation of CD8+ TILs. Data are representative of two independent experiments (n = 7 independent animals). g, Thy1.2+ C57BL/6 mice bearing YUMM1.7-OVA tumors received adoptive transfer of activated CD45.1+ OT-I CD8+ T cells followed by administration of IL-10–Fc or PBS control every other day until day 20. On day 22, CD8+ TILs from pooled samples were enriched and PD-1+TIM-3+CD8+ TILs were sorted. Shown are average basal OCRs. Data are one representative of two independent experiments (n = 3 independent samples). All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test. NS, not significant (P > 0.05). NES, normalized enrichment score.
To further characterize the metabolic regulation effect of IL-10–Fc in vivo, we analyzed the mitochondrial profiles of CD8+ TILs. In agreement with the RNA-seq data, we found IL-10–Fc treatment upregulated the levels of mitochondrial reactive oxygen species (ROS) in the PD-1+TIM-3+CD8+ TILs, indicating enhanced mitochondrial respiration in vivo (Fig. 6f). To directly measure the metabolic profile of sorted CD8+ TILs, we were able to isolate PD-1+TIM-3+CD8+ TILs of a high enough count from YUMM1.7-OVA tumors. Consistent with the RNA-seq data and in vitro results, IL-10–Fc treatment markedly increased the basal OCR level of the PD-1+TIM-3+CD8+ TILs (Fig. 6g). These results indicated that terminally exhausted CD8+ TILs underwent metabolic reprograming toward OXPHOS, remained highly cytotoxic and maintained effector function following exposure to IL-10–Fc.
IL-10–Fc promotes OXPHOS in an MPC-dependent manner
We next used several pathway-specific inhibitors to probe the molecular basis of metabolic regulation of T cells by IL-10–Fc (Fig. 7a–c). Surprisingly, the enhanced OXPHOS was not a result of increased activity of fatty acid oxidation (FAO) or glutaminolysis, as the treatment with FAO inhibitor etomoxir (ETO) and glutaminase inhibitor bis-2-(5-phenylacetamido-1,3,4-thiadiazol-2-yl)ethyl sulfide (BPTES) did not impair the IL-10–Fc-induced elevation of OXPHOS or CD8+ T cell proliferation. However, inhibiting glycolysis with 2-deoxy-d-glucose (2-DG) or blocking pyruvate transportation by inhibiting MPC with UK5099 completely abrogated the effect of IL-10–Fc. Given that MPC plays a central role in importing cytosolic pyruvate into the mitochondrial matrix37, our results suggest IL-10–Fc-induced metabolic regulation might rely on pyruvate generated from glycolysis. To further examine this postulate, we crossed the Mpc1-floxed mice (Mpc1fl/fl)38,39 with Cd4cre × OT-I transgenic mice to obtain the MPC1-deficient OT-I mice, in which Mpc1 gene was ablated in OT-I T cells. Compared with wild-type (WT) OT-I CD8+ T cells, MPC1-knockout (MPC1-KO) OT-I CD8+ T cells failed to respond to IL-10–Fc for promoting OXPHOS or cell expansion (Fig. 7d,e). In addition, the mitochondrial biomass, membrane potential and ROS level of the WT, but not MPC1-KO, PD-1+TIM-3+CD8+ OT-I TILs in the B16F10-OVA tumors were increased upon IL-10–Fc treatment (Extended Data Fig. 10a–c), indicating that IL-10–Fc enhanced the mitochondrial function of terminally exhausted CD8+ TILs in a pyruvate/MPC-dependent manner. Interestingly, we found the IL-10–Fc treatment showed minimum effect on either protein expression or RNA transcription level of MPC1 in CD8+ T cells (Extended Data Fig. 10d,e). Indeed, in restimulated CD8+ T cells IL-10–Fc treatment resulted in activation of STAT3 signaling (Extended Data Fig. 10f), which may interact with ETC complexes in the mitochondria and boost ETC activities for enhanced OXPHOS40. Altogether, IL-10–Fc promoted OXPHOS and mitochondrial function in T cells in an MPC-dependent manner.
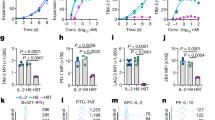
a–c, Resting PMEL CD8+ T cells were restimulated by dimerized α-CD3 antibody in the presence of the indicated inhibitors for 2 d with or without IL-10–Fc. Data are one representative of three independent experiments (n = 4 independent samples). a, Related metabolism pathways and inhibitors used. b, Basal OCR of PMEL CD8+ T cells. c, Fold change of CD8+ T cell counts. d,e, Resting WT or MPC1-KO OT-I CD8+ T cells were restimulated by dimerized α-CD3 antibody in the presence or absence of IL-10–Fc for 2 d. Shown are relative basal OCR of CD8+ T cells (d) and relative CD8+ T cell counts (e) (n = 5 independent samples). f, Resting PMEL CD8+ T cells were restimulated similarly as in a–c in the presence or absence of oligomycin for 2 d. MFI of intracellular granzyme B production was measured (n = 3 independent samples). g,h, Resting PMEL CD8+ T cells were cultured in low-glucose medium with different concentrations of sodium pyruvate for 2 d. g, Shown is fold expansion of PD-1+TIM-3+CD8+ T cells. h, Resting WT and MPC1-KO OT-I CD8+ T cells were cultured similarly as in g. Shown is fold expansion of PD-1+TIM-3+CD8+ T cells with sodium pyruvate (3 mM) versus control. i,j, CD45.1+CD45.2+ C57BL/6 mice bearing B16F10-OVA tumors received adoptive transfer of activated CD45.1+ WT or CD45.2+ MPC1-KO OT-I CD8+ T cells followed by administration of IL-10–Fc or PBS control every other day until day 13 (n = 5 independent animals). Frequencies of transferred OT-I CD8+ T cells among total CD45+ TILs (i) and relative counts of PD-1+TIM-3+CD8+ TILs (j) on day 15. k, C57BL/6 mice bearing YUMM1.7-OVA tumors were sublethally lymphodepleted and received adoptive transfer of activated WT or MPC1-KO OT-I CD8+ T cells on day 10 followed by administration of IL-10–Fc or PBS control every other day until day 24. Shown are average tumor growth curves of each treatment group (n = 4 independent animals). All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test or one-way ANOVA with Tukey’s test or two-way ANOVA (k). NS, not significant (P > 0.05).
Metabolic reprogramming is essential for T cell reinvigoration
It is worth noting that the effect of IL-10–Fc in enhancing proliferation and cytotoxicity (represented by granzyme B production) of the PD-1+TIM-3+ CD8+ T cells was abrogated by the treatment of oligomycin, an OXPHOS pan inhibitor41 (Fig. 7c,f), suggesting the induced metabolic reprogramming was necessary for reinvigorating terminally exhausted T cells by IL-10–Fc. Directly feeding WT OT-I CD8+ T cells, but not MPC1-KO OT-I CD8+ T cells, with sodium pyruvate as an alternative approach of metabolic reprogramming to the IL-10–Fc treatment similarly promoted the proliferation of PD-1+TIM-3+CD8+ T cells upon restimulation by dimerized anti-CD3 antibody (Fig. 7g,h), providing additional evidence that metabolic reprogramming could be an underlying mechanism for reinvigorating exhausted T cells.
To confirm that pyruvate/MPC-dependent OXPHOS is essential for in vivo expansion of terminally exhausted CD8+ TILs and ultimate therapeutic outcome, we next transferred MPC1-KO OT-I CD8+ T cells or WT OT-I CD8+ T cells to mice bearing s.c. B16F10-OVA tumors in combination with IL-10–Fc or PBS control treatment. IL-10–Fc treatment failed to expand the total CD8+ TILs or the PD-1+TIM-3+ subpopulation in mice receiving adoptive transfer of MPC1-KO OT-I CD8+ T cells as compared with that of WT OT-I CD8+ T cells (Fig. 7i,j). We next assessed the therapeutic efficacy of ACT therapy of MPC1-KO OT-I CD8+ T cells combined with IL-10–Fc in YUMM1.7-OVA tumor-bearing mice. The combination of adoptively transferred MPC1-KO OT-I CD8+ T cells and IL-10–Fc exhibited substantially reduced efficacy in tumor growth control compared with that of WT OT-I CD8+ T cells and IL-10–Fc, suggesting that promoting MPC-dependent OXPHOS was necessary for the enhanced therapeutic efficacy mediated by IL-10–Fc (Fig. 7k). These results indicated that MPC-dependent metabolic reprogramming was essential for revitalizing the terminally exhausted CD8+ TILs by IL-10–Fc.
Discussion
T cell exhaustion is one of the major barriers to an effective cancer immunotherapy5,6. While intratumoral progenitor exhausted T cells are known to respond to ICBs, terminally exhausted CD8+ T cells are currently believed to be difficult to reactivate with most existing immunotherapies. Here, we demonstrated IL-10–Fc was a safe and highly efficient metabolic intervention that directly expanded terminally exhausted CD8+ TILs and enhanced their cytotoxic functions, leading to curative cancer immunotherapies. Furthermore, we showed that IL-10–Fc promoted the pyruvate- and MPC-dependent OXPHOS of T cells, inducing reinvigoration of terminally exhausted T cells for enhanced anti-tumor immunity.
We found the therapeutic mode of IL-10–Fc was distinct from and complementary to that of ICBs. The effect of anti-PD-1 antibody therapy relies on the progenitor exhausted CD8+ TILs in the TME8,14. Anti-PD-1 antibody therapy induces the proliferation and differentiation of progenitor exhausted CD8+ TILs, which subsequently lead to the expansion of terminally exhausted CD8+ TILs exerting the cytotoxic function against cancer cells. In contrast, the therapeutic effect of IL-10–Fc is independent of progenitor exhausted CD8+ TILs. Although possessing superior cytotoxic function to the progenitor cells, terminally exhausted CD8+ TILs are short-lived and incapable of self-renewing. IL-10–Fc is a potent cytokine therapy that directly expands the terminally exhausted CD8+ TILs in the absence of the progenitor cells. We have examined the sources and found the increased terminally exhausted T cells upon the IL-10–Fc treatment were contributed mainly by the direct expansion of terminally exhausted CD8+ TILs (IL-10R-dependent), rather than the conversion from progenitor exhausted T cells or the reduced apoptosis of terminally exhausted T cells. The therapeutic outcome and duration of response to ICB therapy may be limited in patients with poor/no tumor infiltration of progenitor exhausted CD8+ TILs. Therefore, the treatment of IL-10–Fc provides a complementary strategy to the existing immunotherapies for enhancing patient response rate.
Metabolically fit effector T cells undergo a metabolic switch to relying on aerobic glycolysis upon activation to meet the high energetic and biosynthesis demands for proliferation and effector function42,43,44. By contrast, exhausted T cells exhibit noticeable metabolic depression of glycolysis and/or OXPHOS17. In particular, impaired mitochondrial OXPHOS was recently found to drive T cell exhaustion18,19,20. CD8+ TILs under persistent antigen stimulation and metabolic stress accumulated dysfunctional mitochondria, leading to reduced proliferative capacity and effector function18,19,20. In line with these observations, our results also imply that terminally exhausted T cells may undergo metabolic adaption in TME to reduce pyruvate-dependent OXPHOS. By directly feeding terminally exhausted T cells with sodium pyruvate to rewire T cell metabolism instead of using IL-10–Fc, we showed that elevated pyruvate-dependent OXPHOS was indeed sufficient to promote the proliferation of WT T cells but not the MPC1-KO T cells. Furthermore, the necessity of promoted OXPHOS for enhanced proliferation and effector function of terminally exhausted T cells as well as the ultimate therapeutic efficacy induced by IL-10–Fc treatment was confirmed by a set of experiments using an OXPHOS inhibitor (oligomycin) or MPC1-KO T cells, where the effects of IL-10–Fc were completely abrogated. Together, these observations implicate that elevation of pyruvate- and MPC-dependent OXPHOS is sufficient and necessary for revitalization of terminally exhausted CD8+ T cells by IL-10–Fc. This finding lays the foundation for further identification of metabolic stimulations that are needed for reinvigorating terminally exhausted T cells, currently a major limitation for cancer immunotherapy.
Substrates for mitochondrial OXPHOS include pyruvate, fatty acids and glutamine. Intriguingly, we found that the enhancement of pyruvate- and MPC-dependent OXPHOS rather than FAO or glutaminolysis was necessary for restoring the functionality of terminally exhausted CD8+ T cells by IL-10–Fc. Our discovery suggests that the effect of metabolic reprograming on anti-tumor immunity of T cells might be pathway and source specific. However, it remains elusive how IL-10–Fc boosts pyruvate- and MPC-dependent OXPHOS in terminally exhausted T cells. Several other pathways related to T cell functions, such as CD28 signaling and NFAT, were also found to be significantly upregulated by IL-10–Fc in the IPA analysis. Elucidating the role of metabolic reprogramming and related signaling pathways in regulating the differentiation, exhaustion and function of various immune cells, such as T cells and natural killer cells, will likely provide critical insight into the design of next-generation immunotherapy enabled by metabolic interventions. Besides cancer, one may expect IL-10–Fc to show similar effects on the terminally exhausted T cells in chronic infections.
Combination cancer immunotherapy involving IL-10–Fc was safe and highly effective. Treatment of solid tumors with ACT immunotherapy remains a major challenge and cures are typically rare. When combined with ACT therapy, IL-10–Fc eradicated a majority of established solid tumors leading to durable cures and protection in multiple mouse tumor models. IL-10–Fc-based metabolic therapy should be applicable in combinations with many existing immunotherapies for enhanced efficacy, such as ICBs, cancer vaccines and so on. TILs from patients who fail to respond to ICBs may be dominated by the terminally exhausted subpopulation8,14. Therefore, the combination of IL-10–Fc with anti-PD-1 therapy could potentially improve patients’ response rates. Indeed, some preclinical45 and clinical46 evidence has been reported recently showing improved therapeutic outcomes of such combination. Furthermore, IL-10–Fc similarly promoted OXPHOS and proliferation of human T cells. Importantly, IL-10–Fc selectively expanded the tumor-antigen-specific CD8+ TILs but not the bystander CD8+ T cells, showing good safety profiles which were confirmed by the measurements of body weight and serum levels of liver enzymes of treated mice. A recent phase I clinical trial showed encouraging results that combination of a PEGylated IL-10 with an anti-PD-1 monoclonal antibody was relatively safe and activated the anti-tumor immunity in patients with advanced solid tumors46, indicating the high translational potential of such combinations in the clinic.
Methods
Mice
Six-to-eight-week-old female CD45.2+Thy1.2+ C57BL/6 (C57BL/6J) mice, BALB/c (BALB/cByJ) mice and CD45.1 mice (B6.SJL-Ptprca Pepcb/BoyCrl) were purchased from Charles River Laboratories. TCR-transgenic Thy1.1+ pmel-1 (PMEL) mice (B6.Cg-Thy1a/Cy Tg(TcraTcrb)8Rest/J) and TCR-transgenic OT-I mice (C57BL/6-Tg(TcraTcrb)1100Mjb/J) were originally purchased from The Jackson Laboratory and maintained in the École Polytechnique Fédérale de Lausanne (EPFL) Center of PhenoGenomics animal facility. Transgenic mice that carried CD4 promoter-mediated Cre recombinase (Cd4cre mice, (B6.Cg-Tg(Cd4-cre)1Cwi/BfluJ)), TCR-transgenic P14 mice (B6;D2-Tg(TcrLCMV)327Sdz/JDvsJ), Rosa26-LSL-Cas9 knock-in mice47 (B6J.129(B6N)-Gt(ROSA)26Sortm1(CAG-cas9*,-EGFP)Fezh/J) and Rosa26-Cas9 knock-in mice (B6J.129(Cg)-Gt(ROSA)26Sortm1.1(CAG-cas9*,-EGFP)Fezh/J) were originally purchased from The Jackson Laboratory and maintained at the University of Lausanne’s pathogen-free facility. CRISPR–Cas9 knock-in P14 TCR-transgenic mice were obtained by crossing of Rosa26-LSL-Cas9 knock-in mice with Cd4cre mice on a P14 background. CD45.1 mice were crossed with CD45.2+ C57BL/6 mice to generate CD45.1+CD45.2+ mice. CRISPR–Cas9 knock-in P14 TCR-transgenic mice were crossed with CD45.1 mice to generate CRISPR–Cas9 knock-in P14 TCR-transgenic mice on either CD45.1+ background or CD45.1+CD45.2+ background. Thy1.1+ PMEL mice were crossed with Thy1.2+ C57BL/6 mice to generate Thy1.2+ PMEL mice. OT-I mice were crossed with CD45.1 mice to generate CD45.1+ OT-I mice. Mpc1fl/fl mice (Mpc1tm1a(EUCOMM)Wtsi) were obtained from J.-C. Martinou (University of Geneva) with permission from J. Rutter (University of Utah School of Medicine)48. Mpc1fl/fl mice were crossed to Cd4cre mice on an OT-I background to generate MPC1-KO OT-I mice. Tcf7DTR-GFP P14 mice on a CD45.2 background were generated as described9. The genotyping was confirmed by Transnetyx. All mice were housed in the EPFL Center of PhenoGenomics or a conventional animal facility of the University of Lausanne and were kept in individually ventilated cages, at 19–23 °C, with 45–65% humidity and with a 12-h dark/light cycle. Experimental procedures in mouse studies were approved by the Swiss authorities (Canton of Vaud, animal protocol IDs 3206, 3533, 3009, 2688.2 and 3040.1) and performed in accordance with the guidelines from the Center of PhenoGenomics of the EPFL and the animal facility of the University of Lausanne.
Human blood samples
All human blood samples (prepared as buffy coats) were purchased from Interregional Blood Transfusion SRC Ltd. with informed consent of anonymous healthy donors, and genetically engineered with ethics approval from the Federal Office of Public Health, Switzerland (Notification 4182209/2).
Cells and tumor models
B16F10 melanoma cells, CT26 mouse colorectal cancer cells, SKOV3-HER2 cells, HEK293T cells and Phoenix-Eco cells were originally acquired from the American Type Culture Collection. B16-gp33, B16F10-OVA and YUMM1.7-OVA mouse melanoma cell lines were generated as previously reported9,49. HER2-transduced MC38 mouse colon cancer cell line (MC38-HER2) and ME275 human melanoma cell line (ME275-HER2) were generated according to the previous publication50. All the mouse tumor cells and SKOV3-HER2 cells were cultured in complete DMEM, a DMEM (Gibco/Thermo Fisher Scientific) supplemented with fetal bovine serum (FBS) (10% v/v, Gibco/Thermo Fisher Scientific) and penicillin/streptomycin (1% v/v, Gibco/Thermo Fisher Scientific). ME275-HER2 human melanoma cells were cultured in RPMI-1640 (Gibco/Thermo Fisher Scientific) medium supplemented with FBS (10% v/v, Gibco/Thermo Fisher Scientific), HEPES (pH 7.2~7.5, 1% v/v, Gibco/Thermo Fisher Scientific) and penicillin/streptomycin (1% v/v, Gibco/Thermo Fisher Scientific). B16F10, YUMM1.7-OVA, B16F10-OVA, B16-gp33 or MC38-HER2 tumor cells (5 × 105 or 1 × 106 or as indicated) were implanted subcutaneously into the right flanks of Thy1.2+ C57BL/6 WT mice to establish the syngeneic tumor models. CT26 mouse colorectal cancer cells (3 × 105) were subcutaneously inoculated into the right flanks of BALB/c mice.
Production of IL-10–Fc protein
As reported previously51,52,53, the IL-10–Fc fusion protein containing a human IL-10 fused at the N terminus with a noncytolytic human IgG1 Fc was expressed by FreeStyle 293-F Cells (Gibco/Thermo Fisher Scientific) at the EPFL Protein Expression Core Facility. Supernatant of culture medium containing IL-10–Fc fusion protein was collected by centrifugation after a 7-d culture and was filtered through a 0.22-μm membrane to obtain a clear solution. The recombinant protein was first captured with a HiTrap Protein A affinity chromatography column on an AKTA pure 25 (GE Healthcare), and eluted with an elution buffer (0.05 M sodium citrate, 0.3 M sodium chloride, pH 3.0). The eluted protein was collected immediately in a neutralization buffer (1 M Tris–HCl, pH 10.0), followed by concentration with membrane ultrafiltration (molecular weight cut-off 10 kDa) in a Vivaspin (GE Healthcare). The concentrated protein solution was further purified with a Superdex 200 Increase size-exclusion chromatography column (GE Healthcare) at a flow rate of 1.0 ml min−1 with PBS buffer on an AKTA Pure 25 (Extended Data Fig. 1a,b). The purified protein was aliquoted and stored at −80 °C before use. The purity of IL-10–Fc was confirmed with SDS–PAGE (Extended Data Fig. 1c).
Preparation of PMEL, WT OT-I, MPC1-KO OT-I and Tcf7 DTR-GFP P14 T cells, and human CD8+ T cells
Spleens from PMEL or OT-I mice were mechanically disrupted and ground through a 70-μm strainer (Fisher Scientific). Red blood cells (RBCs) were lysed with ACK lysis buffer (2 ml per spleen, Gibco/Thermo Fisher Scientific) for 5 min at 25 °C. The splenocytes were washed twice with cold complete RPMI medium, which contained RPMI-1640 (Gibco), FBS (10% v/v), HEPES (pH 7.2~7.5, 1% v/v), penicillin/streptomycin (1% v/v), sodium pyruvate (1% v/v, Gibco/Thermo Fisher Scientific) and 2-mercaptoethanol (0.1% v/v, Gibco/Thermo Fisher Scientific). Splenocytes were then resuspended at a cell density of 2 × 106 per ml in complete RPMI medium supplemented with mouse IL-2 (10 ng ml−1) and IL-7 (1 ng ml−1, PeproTech), as well as human gp10025–33 or OVA257-264 peptide (1 μM, GenScript) for PMEL or OT-I T cells, respectively. After a 3-d culture, live cells were enriched by density gradient centrifugation against Ficoll-Paque PLUS (GE Healthcare), followed by another 2-d culture at a cell density of 0.5~1.0 × 106 per ml in complete RPMI medium supplemented with mouse IL-2 (10 ng ml−1) and IL-7 (10 ng ml−1) to afford activated CD8+ T cells with purity >95% (flow cytometry analyses). MPC1-KO OT-I T cells were prepared similarly as WT OT-I T cells. Peptide-primed CD8+ T cells were next used for ACT in mice. For in vitro experiments, the peptide-primed CD8+ T cells were cultured in complete RPMI medium supplemented with IL-2 (10 ng ml−1) and IL-7 (10 ng ml−1) for another 2 d to afford primed CD8+ T cells in resting phase (day 7). In the pyruvate feeding experiment, primed CD8+ T cells in resting phase were cultured in RPMI medium with low glucose (1 g l−1 d-glucose, Thermo Fisher Scientific), FBS (10% v/v), HEPES (pH 7.2~7.5, 1% v/v), penicillin/streptomycin (1% v/v) and 2-mercaptoethanol (0.1% v/v), which was further supplemented with IL-2 (10 ng ml−1) and sodium pyruvate at different concentrations for 2 d.
Tcf7DTR-GFP P14 T cells were isolated from splenocytes of Tcf7DTR-GFP P14 mice by mashing the spleen through a 100-µm nylon cell strainer (BD Falcon). RBCs were lysed with ACK lysis buffer. For Fig. 3e,f and Extended Data Fig. 3h,i, CD8+ T cells isolated using mouse CD8+ T cell negative enrichment kit (StemCell Technologies) were activated on plates coated with anti-mouse CD3 antibody (17A2, eBioscience/Thermo Fisher Scientific) and anti-CD28 antibody (37.51, eBioscience/Thermo Fisher Scientific) plus in complete RPMI medium containing IL-2 (50 ng ml−1) for 2 d before adoptive transfer. For Fig. 3g,h, Tcf7DTR-GFP P14 T cells were primed by LCMV gp33-41 peptide (1 µM, GenScript) and prepared similarly as described above for PMEL T cells.
Human peripheral blood mononuclear cells from anonymous healthy donors (prepared as buffy coats) were activated in vitro with anti-human CD3 (OKT3, BioLegend) and CD28 (CD28.2, BioLegend) antibodies in the presence of human IL-2 (10 ng ml−1) for 2 d. Activated human CD8+ T cells were isolated by Ficoll density gradient separation and magnetic-activated cell sorting (MACS) using a human CD8+ T cell isolation kit (Miltenyi Biotec) for in vitro assays.
Preparation of control and IL-10Rα-KO P14 T cells
WT P14 CD8+ T cells were isolated from splenocytes of Cas9 knock-in P14 TCR-transgenic mice using CD8 negative selection kit (Miltenyi Biotec) and activated using mouse T-activator CD3/CD28 Dynabeads (Gibco/Thermo Fisher Scientific) for 24 h followed by 4-d expansion in complete RPMI medium containing IL-2 (10 ng ml−1) before use. To generate control and IL-10Rα-KO P14 CD8+ T cells, WT P14 CD8+ T cells were isolated and activated for 24 h similarly, followed by transduction with retroviruses containing scramble control guide RNA (gRNA) or IL-10Rα-targeting gRNA on plates coated with RetroNectin (5 µg cm−2, Clonetech/Takara). Transduced CD8+ T cells were then expanded similarly for an additional 4 d before use. The pool of gRNAs targeting IL-10Rα (IL-10Rα1: 5′-TCTGGCTTCAAACCACACAT-3′, IL-10Rα2: 5′-GCAGTGTTTACTTATCACGA-3′, IL-10Rα3: 5′-GTGGGGACAACACGGACAGT-3′) and a scramble control gRNA control (5′-GCGAGGTATTCGGCTCCGCG-3′) were designed using the publicly available online gRNA design tool CRISPick54.
Analyses of tumor-infiltrating immune cells
Thy1.2+ C57BL/6 mice were inoculated subcutaneously with B16F10 tumor cells (1 × 106) and received i.v. adoptive transfer of PMEL CD8+ T cells (5 × 106) on day 6 post tumor inoculation, followed by four doses (or as indicated) of peritumoral (p.t.) administration of IL-10–Fc (20 µg) or PBS control every other day starting from day 6. For BrdU experiments, mice were administered BrdU (0.8 mg, Sigma-Aldrich) via i.p. injection 1 d before tumor collection. On day 14 (or as indicated), tumors were dissected from the surrounding tissues, weighed, mechanically minced and stirred at 1,000 r.p.m. in RPMI-1640 medium with collagenase Type IV (1 mg ml−1, Gibco/Thermo Fisher Scientific), dispase II (100 μg ml−1, Sigma-Aldrich), hyalurondase (100 μg ml−1, Sigma-Aldrich) and DNase I (100 μg ml−1, Sigma-Aldrich) at 37 °C for 60 min for digestion. RBC lysis was performed on the digested tumor samples with ACK lysing buffer. Tumor-infiltrating leukocytes were then enriched by density gradient centrifugation against Percoll (GE Healthcare), resuspended in PBS with BSA albumin (0.2%, wt/v, Sigma-Aldrich), stained with indicated antibodies and analyzed with flow cytometry. For mice bearing B16F10-OVA tumors and receiving adoptive transfer of either MPC1-KO or WT OT-I T cells, or mice bearing B16-gp33 tumors and receiving adoptive transfer of WT control P14, IL-10Rα-KO P14 or Tcf7DTR-GFP P14 T cells, immune cell infiltrates in tumor were analyzed in a similar way as described above.
Cotransfer of PMEL and OT-I CD8+ T cells
Thy1.2+ C57BL/6 mice were sublethally lymphodepleted by total body irradiation (4 Gy) on day −4, followed by i.v. adoptive cotransfer of Thy1.1+ naive PMEL (2 × 106) and CD45.1+ naive OT-I CD8+ T cells (2 × 106) on day −3 (Fig. 3a–d). On day 0, mice were inoculated with B16F10 tumor cells (5 × 105) subcutaneously and then treated with i.v. adoptive transfer of activated Thy1.2+PMEL CD8+ T cells (5 × 106) on day 10, followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day starting from day 10 (four doses in total). On day 17, mice were killed and TILs were analyzed by flow cytometry as described above.
Cotransfer of WT and MPC1-KO OT-I CD8+ T cells
CD45.1+CD45.2+ mice were inoculated subcutaneously with B16F10-OVA tumor cells (5 × 105) and received i.v. adoptive cotransfer of activated WT CD45.1+ OT-I CD8+ T cells and MPC1-KO CD45.2+ OT-I CD8+ T cells (1:1, 5 × 106 for each) on day 6, followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 12. On day 13, mice were killed and TILs were analyzed by flow cytometry as described above.
Selective depletion of progenitor exhausted T cells
CD45.1+CD45.2+ C57BL/6 mice were inoculated subcutaneously with B16-gp33 tumor cells (2.5 × 105) and received lymphodepletion treatment to deplete the endogenous lymphocytes 1 d before i.v. adoptive transfer of CD45.2+ Tcf7DTR-GFP P14 T cells (1 × 105) on day 11. Mice were peritumorally administrated with IL-10–Fc (20 µg) or PBS control every other day starting from day 11 (seven doses in total). To deplete the diphtheria toxin receptor (DTR)-expressing cells, DT (Sigma-Aldrich) was injected intraperitoneally twice (days 15 and 16) at a dose of 50 µg kg−1. TILs were analyzed on day 25 similarly as described above. In a therapeutic study (Fig. 3g,h), C57BL/6 mice were inoculated subcutaneously with B16-gp33 tumor cells (5 × 105) and received i.v. adoptive transfer of Tcf7DTR-GFP P14 T cells (2.5 × 106) on day 6 followed by p.t. administration of IL-10–Fc (20 µg) or PBS control through every other day starting from day 6 (four doses in total). DT was injected intraperitoneally on days 6, 8 and 10 at a dose of 50 µg kg−1. Tumor area and body weight were monitored every other day.
Flow cytometry analyses
For surface marker staining, cells were collected in U-bottom 96-well plates (Thermal Fischer Scientific), blocked with anti-mouse CD16/32 antibody (BioLegend) and incubated with indicated antibodies at 4 °C for 20 min, followed by live/dead staining by 4,6-diamidino-2-phenylindole (DAPI, Sigma-Aldrich) or Zombie Aqua Fixable Dye (BioLegend). Cells were then washed with PBS containing BSA (0.2%, wt/v) and resuspended in the same buffer for flow cytometry analyses. For intracellular staining, cells were first stained for surface markers and Aqua Fixable Dye as described above. Next, cells were fixed and permeabilized with a Cytofix/Cytoperm Fixation/Permeabilization Solution Kit (BD Biosciences) for cytokine staining or a Foxp3/Transcription Factor Staining Buffer Set (eBioscience) for BrdU, active caspase-3 and transcription factors staining per the manufacturer’s instructions, followed by incubation with indicated antibodies for intracellular staining (temperature and time per the manufacturer’s instructions). For intracellular cytokine staining, cells were first stimulated by a Cell Stimulation Cocktail (protein transport inhibitors included, Invitrogen/Thermo Fisher Scientific) at 37 °C for 4–6 h. Cells were then processed similarly for surface marker staining and intracellular staining as described above. Data were collected using an Attune NxT Flow Cytometer with Attune NxT Software v.3 (Invitrogen/Thermal Fischer Scientific). Analyses were performed using FlowJo 10.6.1 (Tree Star). Gate margins were determined by isotype controls and fluorescence-minus-one controls.
Antibodies and reagents for flow cytometry
The following antibodies or staining reagents were purchased from BioLegend: CD16/32 (93, 101302), Thy1.1 (OX-7, 202529), Thy1.2 (30-H12, 105343), CD45.1 (A20, B218971), CD45.2 (104, 109814), CD8α (53-6.7, 100714), CD8β (YTS256.7.7, 126606), CD4 (RM4-5, 100526), NK1.1 (PK136, 108740), F4/80 (BM8,123108), CD3ε (17A2, 100306), CD19 (6D5,115520), CD44 (IM7, 103006), CD11c (N418, 117348), I-A/I-E (MHC-II, M5/114.15.2, 107643), Siglec-F (S17007L, 155508), CD80 (16-10A1, 104734), CD86 (GL-1, 105006), Foxp3 (MF-14, 126406), CD11b (M1/70, 101228), Ki67 (16A8, 652424), BrdU (3D4, 364104), Granzyme B (GB11, 515403), IFN-γ (XMG1.2, 505826), TNF-α (MP6-XT22, 506308), IL-2 (JES6-5H4, 503822), IL-10Rα (1B1.3a, 112705), CD69 (H1.2F3, 104512), Gr-1 (RB6-8C5, 202519), PD-1 (29F.1A12, 135216), TIM-3 (RMT3-23, 119706). TCF-1 (C63D9, 2203S) was obtained from Cell Signaling Technology. Active caspase-3 (C92-605) was obtained from BD Biosciences. Alexa Fluor 488-conjugated goat anti-rabbit secondary antibody (A-11008), MitoTracker Green FM, MitoTracker Deep Red FM and MitoSOX Red mitochondrial superoxide indicators were obtained from Thermo Fisher Scientific.
Immunofluorescence staining
For tumor tissue section staining, tumors were fixed in paraformaldehyde (4%, Sigma-Aldrich) overnight followed by embedding in paraffin blocks. Paraffin slides were sectioned with a thickness of 4 µm. Tissue sections were rehydrated at the EPFL Histology Core Facility and stained with anti-CD3 antibody (SP7, Abcam, 1:150), followed by staining with an Alexa Fluor 488-conjugated goat anti-rabbit secondary antibody (Invitrogen/Thermo Fisher Scientific) and DAPI. Images were collected using an LSM700 confocal microscope (Zeiss) at the EPFL Bioimaging and Optics Platform.
Cell sorting
CD8+ TILs from B16F10 or YUMM1.7-OVA tumors or CD8+ T cells from in vitro culture were first enriched by MACS using mouse CD8 (TIL) MicroBeads (Miltenyi Biotec), and then stained with surface markers and DAPI followed by sorting with an Aria II sorter (BD Biosciences) at the EPFL Flow Cytometry Core Facility.
In vitro restimulation of CD8+ T cells
Activated PMEL or OT-I CD8+ T cells in resting phase (day 7 in culture) were restimulated with dimerized anti-CD3 antibody (prepared by mixing anti-CD3 antibody (17A2, Bio X Cell) with goat anti-rat IgG (Invitrogen/Thermo Fisher Scientific) at the molar ratio of 2:1) in complete RPMI medium containing IL-2 (10 ng ml−1) for 2 d (Extended Data Fig. 5a). Collected cells were phenotyped with flow cytometry based on the expression level of surface inhibitory receptors (PD-1 and TIM-3). The live CD8+ T cells were analyzed by flow cytometry, or isolated by Ficoll density gradient separation for Seahorse assay. The PD-1+TIM-3+CD8+ T cell subset was sorted for in vitro coculture assay or Seahorse assay.
Preparation of mouse and human CAR-T cells
Constructs for mouse and human CAR targeting HER2 were generated as previously described50. Mouse and human HER2 CAR-T cells were prepared according to a published report55. For mouse CAR-T cell preparation, Phoenix-Eco cells were transfected with HER2 CAR-carrying plasmid and pCL-Eco-packaging plasmid using the calcium phosphate method. Virus-containing supernatant was collected by centrifuge every 24 h post transfection until 72 h. The splenocytes from WT mice were stimulated by mouse T-activator CD3/CD28 Dynabeads for 1 d, and then transduced with the virus-containing supernatant using spin transduction with centrifugation at 2,000g for 2 h at 32 °C. After overnight incubation at 37 °C, the supernatant was replaced with fresh complete RPMI medium supplemented with mouse IL-2 (10 ng ml−1). Cells were passaged every 24 h at a ratio of 1:2 for another 3 d before use. Transduction efficiency was determined by staining surface HER2 CAR using biotinylated human Her2/ErbB2 Protein (Acro Biosystems) and avidin-Alexa Fluor 488 conjugate (Invitrogen/Thermo Fisher Scientific) 48 h post transduction. Untransduced T cells activated by Dynabeads were used as control.
To prepare human CAR-T cells, the replication-defective human HER2 CAR lentivirus was produced in HEK293T cells, which were transfected with plasmids of pVSV-G, Delta 8.9 and human HER2 CAR using the calcium phosphate method. Primary human T cells were isolated from the peripheral blood mononuclear cells of healthy donors and activated as described above, followed by transduction with the supernatant containing human HER2 CAR lentivirus particles on day 1 post activation using spin transduction for 2 h at 32 °C. Cells were passaged at a 1:2 ratio every day with fresh complete RPMI medium supplemented with human IL-2 (10 ng ml−1, BioLegend) for another 5 d before use. Transduction efficiency was evaluated as described above.
In vitro coculture of T cells and tumor cells
B16F10 tumor cells were cultured in complete DMEM as described above. Collected B16F10 tumor cells were seeded in 24- or 96-well microplates or flasks in complete DMEM at 37 °C for 2 d. Following aspiration of tumor culture medium, activated PMEL CD8+ T cells in resting phase as described above were suspended in complete RPMI medium supplemented with IL-2 (10 ng ml−1) and added to the tumor cell culture at a T cell/tumor cell ratio of 1:1. After another 2-d coculture, CD8+ T cells were isolated by Ficoll density gradient separation and MACS with mouse CD8 MicroBeads (Miltenyi Biotec) for Seahorse assay and flow cytometry analyses. To determine the lysis of target cells, the viability of tumor cells from the coculture was measured with DAPI staining and flow cytometry. Similarly, human HER2 CAR-T cells were cocultured with SKOV3-HER2 or ME275-HER2 tumor cells in vitro at an effector/target ratio of 1:1 in the presence or absence of IL-10–Fc (200 ng ml−1) for 2 d. Human CD8+ HER2 CAR-T cells were isolated by Ficoll density gradient separation and MACS using the human CD8+ T cell isolation kit (Miltenyi Biotec) for Seahorse assay and flow cytometry analyses.
Seahorse assay
Seahorse assay was performed to measure OCR and ECAR of T cells. Pan CD8+ T cells or sorted subsets of mouse or human CD8+ T cells (3 × 105 per well) with different treatment conditions were seeded in a Seahorse culture plate (Seahorse Bioscience) in a non-CO2 incubator at 37 °C for 40 min. OCR and ECAR were measured by an XF96 Seahorse Extracellular Flux Analyzer (Seahorse Bioscience) following the manufacturer’s instructions. During a Seahorse assay, cells were treated with oligomycin (1 μM, Sigma-Aldrich), carbonyl cyanide-4-(trifluoromethoxy)phenylhydrazone (FCCP, 2 μM, Sigma-Aldrich), rotenone (0.5 μM, Sigma-Aldrich), antimycin A (0.5 μM, Sigma-Aldrich), glucose (10 mM, Sigma-Aldrich) and 2-DG (50 mM, Sigma-Aldrich). Each condition was performed with 3–6 replicates in a single experiment. OXPHOS and glycolysis were calculated according to the previous report56.
Metabolic inhibitor treatments
Activated PMEL CD8+ T cells in resting phase (day 7 in culture) as described above were restimulated with dimerized anti-CD3 antibody in complete RPMI medium containing IL-2 (10 ng ml−1) and indicated inhibitors (2-DG, 0.5 mM; oligomycin, 1 µM; etomoxir (Sigma-Aldrich), 200 µM; BPTES (Sigma-Aldrich), 20 µM; or UK5099 (Sigma-Aldrich), 100 µM) for 2 d in the presence or absence of IL-10–Fc (200 ng ml−1). Live CD8+ T cell counts were determined by flow cytometry analyses. Pan CD8+ T cells or sorted subsets of CD8+ T cells were prepared for Seahorse assay as described above.
Anti-tumor therapy and rechallenging experiments
Mice bearing established tumors with area around 25–60 mm2 (day 6 post inoculation or as indicated) were treated with adoptive transfer of activated PMEL T cells, OT-I T cells, MPC1-KO OT-I T cells, Tcf7DTR-GFP P14 T cells, or HER2 CAR T cells (5 × 106 or as indicated), followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day (or as indicated) starting from day 6 (four or eight doses in total as indicated). Mice receiving p.t. administration of PBS control only, IL-10–Fc only or untransduced T cells served as controls. Tumor area and body weight were measured every other day. Tumor area was calculated by the formula Area = Length × Width from caliper measurements of two orthogonal diameters. Mice were euthanized when body weight loss was beyond 15% of predosing weight, or tumor area reached 150 mm2 (as a predetermined endpoint), or the animal had become moribund. In tumor cell rechallenging experiments, B16F10 (1 × 105), YUMM1.7-OVA (5 × 105), MC38-HER2 (1 × 106) or CT26 (3 × 105) cells were re-implanted subcutaneously into the left flanks of survived mice from treatment groups at day 90 post primary inoculation. Age-matched naive WT mice were subcutaneously inoculated with the same number of tumor cells as control. Survival of rechallenged mice was monitored for at least another 60 d.
Combination therapy of anti-PD-1 and IL-10–Fc
BALB/c mice were subcutaneously inoculated with CT26 mouse colon cancer cells (3 × 105). The mice bearing established tumors with area around 25–30 mm2 (day 6 post inoculation) were injected peritumorally with anti-PD-1 antibody (100 µg per injection, RMP1-14, Bio X Cell) on days 6, 9 and 12 along with p.t. administration of IL-10–Fc (20 µg per injection) every other day from days 6 to 14. Mice receiving administration of PBS only, IL-10–Fc only or anti-PD-1 antibody only served as controls. Tumor area and body weight were measured every other day.
Measurement of liver enzymes
Thy1.2+ C57BL/6 mice were inoculated subcutaneously with B16F10 tumor cells (1 × 106) and received i.v. adoptive transfer of activated Thy1.1+ PMEL CD8+ T cells (5 × 106) on day 6 followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 12. On day 14, mice were killed and serum samples were collected for analyses. The levels of alanine transaminase and aspartate transaminase in serum were measured using Stanbio Chemistry Reagents (Stanbio) per the manufacturer’s instructions.
RNA-seq and bioinformatics analysis
Mice were sublethally lymphodepleted by total body irradiation (4 Gy) on day −4, followed by i.v. adoptive transfer of Thy1.1+ naive PMEL T cells (2 × 106) on day −3 and inoculation with B16F10 tumor cells on day 0. On day 10, mice were treated with i.v. adoptive transfer of Thy1.2+-activated PMEL CD8+ T cells (5 × 106) prepared as described above, followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day starting from day 10 (four doses in total). On day 18, mice were killed and TILs were isolated as described above. Thy1.1+PD-1+TIM-3+CD8+ TILs (500 cells, purity >99%) were sorted into Hard-Shell 96-Well PCR Plates (Bio-Rad) containing Triton X-100 (0.2%) with RNase inhibitor (Takara) and stored at −80 °C until library preparation. Libraries were sequenced based on a published protocol57. RNA-seq data were collected on a HiSeq3000 (Illumina) with the Real-Time Analysis (RTA) software. The RNA-seq raw data were processed through the standard RNA-seq analysis pipeline. Briefly, raw read alignment was performed using tophat2 v.2.1.0, with the parameters ‘–no-novel-junctions’ and ‘–G’ when specifying the genome file. The reference genome and GFF annotation correspond to the Mus musculus GRCm38.p4 genome version. Following the tophat2 alignment, read counts mapped to each gene were generated using HTseq count. The differential expression analysis of the raw counts was performed based on the DESeq2 R library. The false discovery rate-adjusted P values after Benjamini–Hochberg correction for multiple-testing were used as the statistics to define the differential expression. The adjusted P value cut-offs for the final lists of differentially expressed genes were as follows: IL-10–Fc versus PBS: 842 differentially expressed genes with adjusted P value smaller than 0.05. Heat maps were created using log2 fold change of genes identified as differentially expressed by DESeq2 (adjusted P value < 0.05). Rows were centered and scaled using z-scores. GSEAs were done using GSEA_4.0.2 and gene sets in the MSigDB database (C2 and C7) on z-scores against a gene set permutation. Canonical pathway analysis was performed using QIAGEN IPA.
Immunoblot analysis
Briefly, in vitro cultured PMEL CD8+ T cells (day 7) were starved in FBS-free RPMI medium overnight and restimulated as described above for the indicated time. Cells were lysed in RIPA lysis buffer (Cell Signaling Technology) with 1× protease inhibitor cocktails (Cell Signaling Technology). Before electrophoresis, total cell lysates were diluted with 4× protein sample loading buffer (Invitrogen/Thermo Fisher Scientific) containing 1× reducing reagents (Invitrogen/Thermo Fisher Scientific) and incubated at 72 °C for 10 min. Electrophoresis of proteins from total cell lysates was performed with NuPAGE electrophoresis system (Invitrogen/Thermo Fisher Scientific). Proteins were then transferred to PVDF membrane (Invitrogen/Thermo Fisher Scientific) by Trans-Blot SD semidry transfer cell (Bio-Rad) at 18 V for 30 min. The PVDF membranes were blocked with 5% w/v nonfat dry milk (Sigma-Aldrich) and then incubated with the indicated primary antibodies at 4 °C overnight. The bands were visualized using SignalFire ECL reagent (Cell Signaling Technology) after incubation with horseradish peroxidase (HRP)-conjugated antibodies. The primary antibodies of p-STAT3 (Tyr705, D3A7, 9145 S, 1:2,000) and MPC1 (D2L9I, 14462S, 1:1,000) and the secondary antibodies of HRP-linked anti-rabbit IgG (7074S, 1:3,000) were obtained from Cell Signaling Technology. The HRP-conjugated beta actin monoclonal antibody (BA3R, MA5-15739-HRP, 1:2,000) was obtained from Invitrogen/Thermo Fisher Scientific.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 8 (GraphPad Software). Data are presented as mean ± s.e.m. unless otherwise indicated. Comparisons of two groups were performed by using two-tailed unpaired Student’s t-test. Comparisons of multiple groups at a single time point were performed by using one-way analysis of variance (ANOVA) and Tukey’s test. Survival data were analyzed using the log-rank test. No statistically significant differences were considered when P values were larger than 0.05.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Gene sets in the MSigDB database (C2 and C7) were used for gene set enrichment analysis. All data generated and supporting the findings of this study are available within the paper. The RNA-seq data for tumor-infiltrating lymphocytes are available in the Gene Expression Omnibus database under accession code GSE168990. Source data are provided with this paper. Additional information and materials will be made available upon reasonable request.
References
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer–immune set point. Nature 541, 321–330 (2017).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 372, 2521–2532 (2015).
Page, D. B., Postow, M. A., Callahan, M. K., Allison, J. P. & Wolchok, J. D. Immune modulation in cancer with antibodies. Annu. Rev. Med. 65, 185–202 (2014).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Thommen, D. S. & Schumacher, T. N. T cell dysfunction in cancer. Cancer Cell 33, 547–562 (2018).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Siddiqui, I. et al. Intratumoral Tcf1+ PD-1+ CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Kurtulus, S. et al. Checkpoint blockade immunotherapy induces dynamic changes in PD-1− CD8+ tumor-infiltrating T cells. Immunity 50, 181–194.e6 (2019).
LaFleur, M. W. et al. PTPN2 regulates the generation of exhausted CD8+ T cell subpopulations and restrains tumor immunity. Nat. Immunol. 20, 1335–1347 (2019).
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 338, 1220–1225 (2012).
He, R. et al. Follicular CXCR5-expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–416 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Franco, F., Jaccard, A., Romero, P., Yu, Y. R. & Ho, P. C. Metabolic and epigenetic regulation of T-cell exhaustion. Nat. Metab. 2, 1001–1012 (2020).
Zhang, L. & Romero, P. Metabolic control of CD8+ T cell fate decisions and antitumor immunity. Trends Mol. Med. 24, 30–48 (2018).
Bengsch, B. et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8+ T cell exhaustion. Immunity 45, 358–373 (2016).
Scharping, N. E. et al. Mitochondrial stress induced by continuous stimulation under hypoxia rapidly drives T cell exhaustion. Nat. Immunol. 22, 205–215 (2021).
Yu, Y. R. et al. Disturbed mitochondrial dynamics in CD8+ TILs reinforce T cell exhaustion. Nat. Immunol. 21, 1540–1551 (2020).
Vardhana, S. A. et al. Impaired mitochondrial oxidative phosphorylation limits the self-renewal of T cells exposed to persistent antigen. Nat. Immunol. 21, 1022–1033 (2020).
Fujii, S. I., Shimizu, K., Shimizu, T. & Lotze, M. T. Interleukin-10 promotes the maintenance of antitumor CD8+ T-cell effector function in situ. Blood 98, 2143–2151 (2001).
Mumm, J. B. et al. IL-10 elicits IFNγ-dependent tumor immune surveillance. Cancer Cell 20, 781–796 (2011).
Tanikawa, T. et al. Interleukin-10 ablation promotes tumor development, growth, and metastasis. Cancer Res. 72, 420–429 (2012).
Naing, A. et al. PEGylated IL-10 (pegilodecakin) induces systemic immune activation, CD8+ T cell invigoration and polyclonal T cell expansion in cancer patients. Cancer Cell 34, 775–791.e3 (2018).
Ip, W. K. E., Hoshi, N., Shouval, D. S., Snapper, S. & Medzhitov, R. Anti-inflammatory effect of IL-10 mediated by metabolic reprogramming of macrophages. Science 356, 513–519 (2017).
Tan, J. C., Indelicato, S. R., Narula, S. K., Zavodny, P. J. & Chou, C. C. Characterization of interleukin-10 receptors on human and mouse cells. J. Biol. Chem. 268, 21053–21059 (1993).
Wang, J., Saffold, S., Krauss, J., Chen, W. & Cao, X. Eliciting T cell immunity against poorly immunogenic tumors by immunization with dendritic cell-tumor fusion vaccines. J. Immunol. 161, 5516–5524 (1998).
Lechner, M. G. et al. Immunogenicity of murine solid tumor models as a defining feature of in vivo behavior and response to immunotherapy. J. Immunother. 36, 477–489 (2013).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Moynihan, K. D. et al. Eradication of large established tumors in mice by combination immunotherapy that engages innate and adaptive immune responses. Nat. Med. 22, 1402–1410 (2016).
Pai, C. C. S. et al. Clonal deletion of tumor-specific T cells by interferon-γ confers therapeutic resistance to combination immune checkpoint blockade. Immunity 50, 477–492.e8 (2019).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Santos, J. M. et al. Adenovirus coding for interleukin-2 and tumor necrosis factor alpha replaces lymphodepleting chemotherapy in adoptive T cell therapy. Mol. Ther. 26, 2243–2254 (2018).
Klapper, J. A. et al. High-dose interleukin-2 for the treatment of metastatic renal cell carcinoma: a retrospective analysis of response and survival in patients treated in the Surgery Branch at the National Cancer Institute between 1986 and 2006. Cancer 113, 293–301 (2008).
Floros, T. & Tarhini, A. A. Anticancer cytokines: biology and clinical effects of interferon-α2, interleukin (IL)-2, IL-15, IL-21, and IL-12. Semin. Oncol. 42, 539–548 (2015).
Buck, M. D. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Herzig, S. et al. Identification and functional expression of the mitochondrial pyruvate carrier. Science 336, 93–96 (2012).
Gray, L. R. et al. Hepatic mitochondrial pyruvate carrier 1 is required for efficient regulation of gluconeogenesis and whole-body glucose homeostasis. Cell Metab. 22, 669–681 (2015).
Grenell, A. et al. Loss of MPC1 reprograms retinal metabolism to impair visual function. Proc. Natl Acad. Sci. USA 116, 3530–3535 (2019).
Wegrzyn, J. et al. Function of mitochondrial Stat3 in cellular respiration. Science 323, 793–797 (2009).
Hamaidi, I. et al. Sirt2 inhibition enhances metabolic fitness and effector functions of tumor-reactive T cells. Cell Metab. 32, 420–436 (2020).
Lim, A. R., Rathmell, W. K. & Rathmell, J. C. The tumor microenvironment as a metabolic barrier to effector T cells and immunotherapy. Elife 9, e55185 (2020).
Li, X. et al. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat. Rev. Clin. Oncol. 16, 425–441 (2019).
Chapman, N. M., Boothby, M. R. & Chi, H. Metabolic coordination of T cell quiescence and activation. Nat. Rev. Immunol. 20, 55–70 (2020).
Qiao, J. et al. Targeting tumors with IL-10 prevents dendritic cell-mediated CD8+ T cell apoptosis. Cancer Cell 35, 901–915.e4 (2019).
Naing, A. et al. Pegilodecakin combined with pembrolizumab or nivolumab for patients with advanced solid tumours (IVY): a multicentre, multicohort, open-label, phase 1b trial. Lancet Oncol. 20, 1544–1555 (2019).
Platt, R. J. et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Vanderperre, B. et al. Embryonic lethality of mitochondrial pyruvate carrier 1 deficient mouse can be rescued by a ketogenic diet. PLoS Genet. 12, e1006056 (2016).
Cheng, W. C. et al. Uncoupling protein 2 reprograms the tumor microenvironment to support the anti-tumor immune cycle. Nat. Immunol. 20, 206–217 (2019).
Tschumi, B. O. et al. CART cells are prone to Fas- and DR5-mediated cell death. J. Immunother. Cancer 6, 71 (2018).
Guo, Y. et al. Purification and characterization of human IL-10/Fc fusion protein expressed in Pichia pastoris. Protein Expr. Purif. 83, 152–156 (2012).
Armour, K. L., Clark, M. R., Hadley, A. G. & Williamson, L. M. Recombinant human IgG molecules lacking Fcγ receptor I binding and monocyte triggering activities. Eur. J. Immunol. 29, 2613–2624 (1999).
Steele, A. W., Nickerson, P. W., Steurer, W., Steiger, J. & Strom, T. B. Administration of noncytolytic IL-10/Fc in murine models of lipopolysaccharide-induced septic shock and allogeneic islet transplantation. J. Immunol. 154, 5590–5600 (1995).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Ma, L. et al. Enhanced CAR-T cell activity against solid tumors by vaccine boosting through the chimeric receptor. Science 365, 162–168 (2019).
Michelet, X. et al. Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat. Immunol. 19, 1330–1340 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Acknowledgements
We thank D. Trono and B.E. Correia for providing plasmids of Delta 8.9 and pVSV-G; J.-C. Martinou (University of Geneva) for providing Mpc1fl/fl mice; and A. Donda (University of Lausanne) for the technical support on human CAR T cells. We acknowledge the EPFL Center of PhenoGenomics, Flow Cytometry Core Facility and Protein Expression Core Facility for technical assistance. This work was supported in part by the Swiss National Science Foundation (SNSF project grant no. 315230_173243), the ISREC Foundation with a donation from the Biltema Foundation, the Swiss Cancer League (grant no. KFS-4600-08-2018), the European Research Council under the ERC grant agreement MechanoIMM (grant no. 805337), the Kristian Gerhard Jebsen Foundation, Fondation Pierre Mercier pour la science, an Anna Fuller Fund grant and the EPFL (L.T.). P.-C.H. was supported in part by the Swiss Institute for Experimental Cancer Research (ISREC grant no. 26075483), SNSF project grants (grant nos. 31003A_163204 and 31003A_182470), the Cancer Research Institute Lloyd J. Old STAR award and the European Research Council Starting Grant (grant no. 802773-MitoGuide). P.R. was supported in part by grants from the SNSF (grant nos. 310030_182735 and 310030E-164187). W.H. was supported in part by the Swiss Cancer League (grant no. KFS-4407-02-2018) and the SNSF (grant no. 310030B_179570). W.X. was supported in part by the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB29030000) and the Ministry of Science and Technology of China (grant no. 2016YFC1303503). M.G. was supported by the Chinese Scholarship Council (grant no. 201808320453).
Author information
Authors and Affiliations
Contributions
Y.G., Y.-Q.X. and L.T. conceived the study. Y.G., Y.-Q.X., L.T. and P.-C.H. designed the experiments. Y.G., Y.-Q.X., M.G., Y.Z., F.F., M.W., I.S., A.B., H.W., H.Y., B.F., X.X., C.M.S., B.T., A.C., Y.W., W.L., W.X., W.H. and P.R. performed the experiments. Y.G., Y.-Q.X., L.T. and P.-C.H. analyzed the data. Y.G., Y.-Q.X. and L.T. wrote the manuscript. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.G., L.T. and Y.-Q.X. are inventors on the patent (International Publication Number WO 2021053134) related to the technology described in this manuscript. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IL-10–Fc promotes tumor infiltration of T cells but shows less effects on other immune cells.
a, Schematic diagram of the production and purification of IL-10–Fc. b, Representative size-exclusion chromatographic traces. Peak 3 (P3) was collected and analyzed. c, Representative sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis of purified IL-10–Fc. DTT, dithiothreitol. Results are one representative of three independent experiments. d, Splenocytes from PMEL mice were cultured with human gp10025–33 (hgp100) peptide (1 μM) and IL-2 (10 ng ml−1) for 3 d in the presence of IL-10–Fc or recombinant IL-10 at a series of concentrations. Shown are fold changes of CD8+ T cell counts (normalized by that in PBS control) at each concentration of cytokine. e-i, The experimental setting was the same as described in Fig. 1. Data are one representative of three or four independent experiments. e, Counts of CD45.2+ tumor infiltrating lymphocytes (TILs) and CD3+ TILs in B16F10 tumors. Shown are pooled data of two independent experiments (n = 10 independent animals). f, Representative immunofluorescence images of samples from each group. Green, CD3; blue, DAPI. Scale bar: 100 μm. Results are one representative of two independent experiments. g, Counts of various immune cell subsets in B16F10 tumors. For endogenous CD8+ T cells, PMEL CD8+ T cells, and total CD4+ T cells, shown are pooled data of two independent experiments (n = 10 independent animals). For other immune cells, data are representative of three independent experiments (n = 5 independent animals). NK: natural killer; DCs: dendritic cells; TAMs: tumor associated macrophages; MDSCs: myeloid-derived suppressive cells. h, Mean fluorescence intensity (MFI) of maturation markers on CD11b+CD11c+ tumor infiltrating DCs (n = 5 independent animals). i, Frequencies of M1 phenotype (CD206–MHC-II+) among CD11b+F4/80+ TAMs (n = 5 independent animals). All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test or one-way ANOVA and Tukey’s test; NS, not significant (P > 0.05).
Extended Data Fig. 2 Tumor infiltrating PD-1+TIM-3+CD8+ T cells are PD-1+TCF-1–TIM-3+ terminally exhausted CD8+ T cells, which respond to IL-10–Fc through IL-10 receptor.
a-d, The experimental setting was the same as described in Fig. 1. a-c, Data are representative of two independent experiments (n = 4 independent animals). a, Representative flow cytometry plots showing the frequencies of progenitor exhausted (TCF-1+TIM-3–) and terminally exhausted (TCF-1–TIM-3+) CD8+ TILs among total CD44+PD-1+CD8+ TILs. b,c, Frequencies and counts of three subpopulations of CD8+ TILs based on TCF-1/TIM-3 four-quadrant gating as shown in a. d, Frequencies of IFNγ+TNFα+ polyfunctional and IL-2+ T cells among PD-1+TIM-3–CD8+ TILs or PD-1+TIM-3+CD8+ TILs from untreated tumors (n = 5 independent animals). Memory PMEL CD8+ T cells (n = 5 independent animals) in spleens from cured mice (schedule shown in Fig. 5c) were stimulated in the same way as positive controls. Data are one representative of two independent experiments. e-g, Activated CD8+ T cells from CRISPR-Cas9 KI P14 T cell receptor (TCR) transgenic mice were transfected with control gRNA or IL-10 receptor-α (IL-10Rα) allele specific gRNA to generate wild type control P14 T cells (Ctrl) or IL-10Rα-KO P14 T cells (IL-10Rα-KO), respectively. e, MFI of IL-10Rα expression. Data are one representative of two independent experiments (n = 6 independent samples). f, Ctrl P14 or IL-10Rα-KO P14 CD8+ T cells were restimulated by dimerized anti-CD3 (α-CD3) antibody in the presence or absence of IL-10–Fc and cultured for 2 d. Cell counts were analyzed by flow cytometry. Shown are relative T cell counts treated with IL-10–Fc versus that with PBS control. Data are one representative of two independent experiments (n = 3 independent samples). g, Experimental timeline of Fig. 2b. All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test or one-way ANOVA and Tukey’s test; NS, not significant (P > 0.05).
Extended Data Fig. 3 IL-10–Fc expands terminally exhausted CD8+ T cells in a progenitor exhausted cell-independent manner.
a,b, Thy1.2+ C57BL/6 mice were sublethally lymphodepleted and received adoptive transfer of Thy1.1+ naïve PMEL CD8+ T cells. Mice were inoculated with B16F10 tumor cells and were treated with adoptive transfer of Thy1.2+ activated PMEL CD8+ T cells. Subset [1] (PD-1+TIM-3–) and subset [2] (PD-1+TIM-3+) were sorted from pooled Thy1.1+CD8+ TILs (n = 10 independent animals). Sorted subsets (5 × 104 for each) were transferred separately into Thy1.2+ C57BL/6 recipient mice bearing B16F10 tumors followed by administration of IL-10–Fc or PBS control. On day 13, mice were killed (n = 5 independent animals for groups receiving subset [1]; n = 3 independent animals for groups receiving subset [2]). a, Experimental timeline. b, Counts of tumor infiltrating Thy1.1+ donor T cells. c-e, Thy1.2+ C57BL/6 mice bearing B16F10 tumors received adoptive transfer of activated PMEL Thy1.1+CD8+ T cells and were killed. TILs were pooled from 3 ~ 6 mice and two subsets of Thy1.2+ endogenous CD8+ T cells were sorted. Sorted subsets were cultured separately ex vivo in the presence of dimerized α-CD3 and IL-2 with or without IL-10–Fc for 3 d. Data are one representative of two independent experiments (n = 3 independent samples). c, Experimental timeline. d, Fold change of T cell counts after 3-d ex vivo culture (n = 3 independent samples). e, Frequencies of PD-1+TIM-3+ terminally exhausted CD8+ T cells after 3-d ex vivo culture (n = 3 independent samples). f, Stacked bar graphs show the putative origins of PD-1+TIM-3+ cells in IL-10–Fc-treated tumors based on experimental results in d and e. g, The experimental setting was the same as that described in Fig. 1. MFI of active caspase-3 among three subpopulations (n = 4 independent animals). Data are one representative of two independent experiments. h,i, The experimental setting was the same as that shown in Fig. 3e. (n = 4 independent animals). h, Frequencies of Tcf7DTR-GFP+ P14 CD8+ T cells among all CD8+ TILs before (day 15) and after (day 17) the DT treatment. i, Representative flow cytometry plots. All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test; NS, not significant (P > 0.05).
Extended Data Fig. 4 IL-10–Fc shows minimal effects on T cell glycolysis.
Primed PMEL CD8+ T cells in resting phase (day 7) were co-cultured with B16F10 tumor cells at a ratio of 1 to 1 in the presence or absence of IL-10–Fc. After a 2-d co-culture, CD8+ T cells were isolated for Seahorse assay and flow cytometry analyses, respectively. Data are representative of three independent experiments (n = 3 independent samples). a, Real-time analysis of extracellular acidification rate (ECAR) of PMEL CD8+ T cells from the co-culture assay. b, Basal and maximal ECAR levels of PMEL CD8+ T cells. All data represent the mean ± s.e.m. and are analyzed by one-way ANOVA and Tukey’s test; NS, not significant (P > 0.05).
Extended Data Fig. 5 Ex vivo generated PD-1+TIM-3+CD8+ T cells by restimulation exhibit exhaustion phenotypes similarly as terminally exhausted CD8+ TILs.
a, Dimerized α-CD3 antibody was produced by mixing rat α-CD3 antibody with goat α-rat IgG Fc antibody at the mole ratio of 2:1. Primed PMEL CD8+ T cells in resting phase (day 7) were restimulated by dimerized α-CD3 at indicated concentrations and cultured in complete RPMI 1640 medium supplemented with IL-2 (10 ng ml−1) for 2 d. Shown are representative flow cytometry plots and frequencies of PD-1+TIM-3+CD8+ T cells among total PMEL CD8+ T cells. Data are representative of three independent experiments (n = 3 independent samples). b, Activated PMEL CD8+ T cells in resting phase were restimulated by dimerized α-CD3 antibody and IL-2 (10 ng ml−1) for 2 d. Freshly isolated naïve PMEL CD8+ T cells from splenocytes and restimulated PMEL CD8+ T cells were compared for the expression of PD-1, TIM-3 surface markers and TCF-1 transcription factor.
Extended Data Fig. 6 IL-10–Fc enhances OXPHOS and proliferation of human CD8+ T cells as well as their killing efficiency of target cells.
a-c, Human HER2 CAR-T cells were co-cultured with SKOV3-HER2 (a) or ME275-HER2 (b,c) tumor cells in vitro at a ratio of 1 to 1 in the presence or absence of IL-10–Fc. After a 2-d co-culture, counts of tumor cells and CD8+ T cells were measured by flow cytometry. CD8+ T cells from the co-culture were further isolated for Seahorse assay (a). Data are one representative of two independent experiments (n = 3 independent samples). a, Average basal OCR of the CD8+ HER2 CAR-T cells. b, Relative counts of PD-1+LAG-3+CD8+ CAR-T cells treated with IL-10–Fc versus that with PBS. c, Percentage of target cell killing by CAR-T cells. d-f, Activated human CD8+ T cells in resting phase were restimulated by coated α-CD3 antibody (0.05 μg mL−1) for 2 d. PD-1+LAG-3+CD8+ T cells were sorted for a Seahorse assay. d, Average basal OCR of PD-1+LAG-3+CD8+ T cells. e, Relative counts of PD-1+LAG-3+CD8+ T cells treated with IL-10–Fc versus that with PBS. f, MFI of IL-10Rα expression on three different subpopulations. Data are one representative of two independent experiments (n = 3 independent samples). All data represent the mean ± s.e.m. and are analyzed by one-way ANOVA and Tukey’s test or two-sided Student’s t-test; NS, not significant (P > 0.05).
Extended Data Fig. 7 IL-10–Fc potentiates adoptive T cell transfer therapies to eradicate established solid tumors without overt toxicity.
a, Shown are individual tumor growth curves of the therapy experiments in Fig. 5a. Indicated are numbers of long-term surviving mice among the total number of mice in the group. b-c, The experimental setting was the same as that described in Fig. 1. b, Ratio of counts of CD8+ T cells to regulatory T cells (Tregs) in tumors in different treatment groups. c, Frequencies of granzyme B+IFNγ+TNFα+ polyfunctional CD8+ T cells among total CD8+ TILs. Data are one representative of three independent experiments (n = 5 independent animals). d-h, Thy1.2+ C57BL/6 mice were inoculated s.c. with B16F10 melanoma cells (5 × 105) on day 0, and received i.v. adoptive transfer of activated Thy1.1+ PMEL CD8+ T cells (5 × 106) on day 6, followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every 4 days until day 18. d, Experimental timeline. e-h, Shown are average tumor growth curves (e), survival curves (f), and individual tumor growth curves (g, h) of each treatment group. Indicated are numbers of long-term surviving mice out of the total numbers in the group. Data are representative of two independent experiments (n = 10 independent animals). i-k, The experimental setting was the same as that described in Fig. 1. On day 14, mice were killed and serum samples were collected for analysis. Shown are relative body weight (i), serum alanine transaminase (ALT, j) and aspartate transaminase (AST, k) levels of the treated mice. Shown are pooled data of two independent experiments (n = 10 independent animals). All data represent the mean ± s.e.m. and are analyzed by one-way ANOVA and Tukey’s test; NS, not significant (P > 0.05).
Extended Data Fig. 8 IL-10–Fc synergizes with immune checkpoint blockade therapy to eradicate established tumors.
BALB/c mice were inoculated subcutaneously with CT26 colon adenocarcinoma cells (3 × 105) and received p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 14, together with p.t. administration of α-PD-1 (RMP1-14, 100 µg) every 3 days until day 12. Data are one representative of three independent experiments (n = 5 or 10 independent animals). a, Experimental timeline. b, Shown are survival curves of each treatment group. Indicated are numbers of long-term surviving mice among the total number of mice in the group. c, Shown are individual tumor growth curves. d, Cured mice from treatment group of combination of α-PD-1 and IL-10–Fc were re-challenged subcutaneously with CT26 (3 × 105) cells at day 90 post primary inoculation. Naïve wild type mice were inoculated with the same number of tumor cells as controls. Shown are survival curves and numbers of long-term surviving mice against the re-challenges. All data represent the mean ± s.e.m. and are analyzed by Log-rank test for survival curves; NS, not significant (P > 0.05).
Extended Data Fig. 9 IL-10–Fc upregulates OXPHOS and effector function related pathways in terminally exhausted CD8+ TILs in vivo.
The experimental setting for RNA-sequencing (RNA-seq) was shown in Fig. 6a. a, Heatmap of top 100 differentially expressed genes (IL-10–Fc versus PBS) were generated using z-scores derived from log2 (fold change). RNA-seq datasets were generated from two independent animals from PBS treatment group and four independent animals from IL-10–Fc treatment group. b, Volcano plot of differentially expressed genes (IL-10–Fc versus PBS). Transcripts with a false discovery rate (FDR) value < 0.05 and log2 (fold change) > 1 are highlighted in red; transcripts with an FDR value < 0.05 and log2 (fold change) < -1 are highlighted in blue. c, Canonical pathway analysis was performed by using QIAGEN Ingenuity Pathway Analysis (IPA) based on differentially expressed genes (DEGs) from RNA-seq dataset. Enrichment Z-score is generated based on hypergeometric distribution, where the negative logarithm of the significance level (P value) is obtained by Fisher’s exact test at the right tail. Shown are top 15 pathways that were significantly upregulated by IL-10–Fc treatment (-log (P value) > 6). Numbers on the right of each bar indicates the significance -log (P value) of each pathway. d, Bean plot of log2 (fold change) of genes involved in oxidative phosphorylation pathway shown in c.
Extended Data Fig. 10 IL-10–Fc promotes mitochondrial fitness and function of terminally exhausted CD8+ TILs in vivo in an MPC dependent manner.
a-c, CD45.1+CD45.2+ mice were inoculated subcutaneously with B16F10-OVA tumor cells (5 × 105) and received i.v. adoptive co-transfer of activated CD45.1+ WT OT-I CD8+ T cells and CD45.2+ MPC1-KO OT-I CD8+ T cells (1:1, 5 × 106 for each) on day 6 followed by p.t. administration of IL-10–Fc (20 µg) or PBS control every other day until day 12. On day 13, mice were killed and tumors were processed and analyzed by flow cytometry. Data are one representative of two independent experiments (n = 7 independent animals). a, b, Relative MFI of MitoTracker Green FM (MitoGreen) (a) and MitoTracker Deep Red FM (MitoDeepRed) (b) of WT or MPC1-KO PD-1+TIM-3+CD8+ OT-I TILs treated with IL-10–Fc versus that with PBS control. c, Frequencies of MitoSOX+CD8+ T cells among total WT or MPC1-KO PD-1+TIM-3+CD8+ OT-I TILs. d, f, Activated PMEL CD8+ T cells were starved overnight and restimulated by dimerized α-CD3 antibody in the presence or absence of IL-10–Fc for overnight (d) or indicated time (f). Proteins from total cell lysates were separated by SDS-PAGE and MPC1 (d) or pSTAT3 (Tyr705) (f) were detected by Western blot. Results are one representative of three independent experiments. e, Transcript expression of Mpc1, Mpc1-ps, and Mpc2. Data are extracted from analyses of RNA-seq. All data represent the mean ± s.e.m. and are analyzed by two-sided Student’s t-test; NS, not significant (P > 0.05).
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed SDS–PAGE.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Guo, Y., Xie, YQ., Gao, M. et al. Metabolic reprogramming of terminally exhausted CD8+ T cells by IL-10 enhances anti-tumor immunity. Nat Immunol 22, 746–756 (2021). https://doi.org/10.1038/s41590-021-00940-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00940-2
This article is cited by
-
IL-10-expressing CAR T cells resist dysfunction and mediate durable clearance of solid tumors and metastases
Nature Biotechnology (2024)
-
An oncolytic virus–T cell chimera for cancer immunotherapy
Nature Biotechnology (2024)
-
Comparative transcriptional analysis identifies genes associated with the attenuation of Theileria parva infected cells after long-term in vitro culture
Scientific Reports (2024)
-
Metabolically enhanced CAR T cells efficiently clear solid tumors in mice
Nature Biotechnology (2024)
-
From complexity to clarity: unravelling tumor heterogeneity through the lens of tumor microenvironment for innovative cancer therapy
Histochemistry and Cell Biology (2024)