Abstract
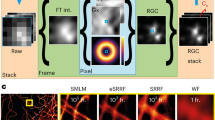
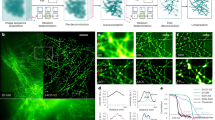
Various super-resolution imaging techniques have been developed to break the diffraction-limited resolution of light microscopy. However, it still remains challenging to obtain three-dimensional (3D) super-resolution information of structures and dynamic processes in live cells at high speed. We recently developed high-speed single-point edge-excitation sub-diffraction (SPEED) microscopy and its two-dimensional (2D)-to-3D transformation algorithm to provide an effective approach to achieving 3D sub-diffraction-limit information in subcellular structures and organelles that have rotational symmetry. In contrast to most other 3D super-resolution microscopy or 3D particle-tracking microscopy approaches, SPEED microscopy does not depend on complex optical components and can be implemented onto a standard inverted epifluorescence microscope. SPEED microscopy is specifically designed to obtain 2D spatial locations of individual immobile or moving fluorescent molecules inside sub-micrometer biological channels or cavities at high spatiotemporal resolution. After data collection, post-localization 2D-to-3D transformation is applied to obtain 3D super-resolution structural and dynamic information. The complete protocol, including cell culture and sample preparation (6–7 d), SPEED imaging (4–5 h), data analysis and validation through simulation (5–13 h), takes ~9 d to complete.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data used to generate example data figures can be found on our lab’s GitHub repository: https://github.com/YangLab-Temple/Data.
Code availability
The code used in this work is available at https://github.com/YangLab-Temple/Master under the GNU General Public License v3.0. Specifically, the code for the reproducibility rate can be found at https://github.com/YangLab-Temple/Master/tree/master/reproducibility%20rate.
References
Abbe, E. Beiträge zur Theorie des Mikroskops und der mikroskopischen Wahrnehmung. Archiv für Mikroskopische Anatomie 9, 413–468 (1873).
Mockl, L., Lamb, D. C. & Brauchle, C. Super-resolved fluorescence microscopy: Nobel Prize in Chemistry 2014 for Eric Betzig, Stefan Hell, and William E. Moerner. Angew. Chem. Int. Ed. Engl. 53, 13972–13977 (2014).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Biol. 190, 165–175 (2010).
Schermelleh, L. et al. Super-resolution microscopy demystified. Nat. Cell Biol. 21, 72–84 (2019).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Shroff, H., Galbraith, C. G., Galbraith, J. A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nat. Methods 5, 417–423 (2008).
Schnitzbauer, J., Strauss, M. T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Kner, P., Chhun, B. B., Griffis, E. R., Winoto, L. & Gustafsson, M. G. L. Super-resolution video microscopy of live cells by structured illumination. Nat. Methods 6, 339–342 (2009).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Babcock, H., Sigal, Y. M. & Zhuang, X. A high-density 3D localization algorithm for stochastic optical reconstruction microscopy. Opt. Nanoscopy https://doi.org/10.1186/2192-2853-1-6 (2012).
von Diezmann, A., Shechtman, Y. & Moerner, W. E. Three-dimensional localization of single molecules for super-resolution imaging and single-particle tracking. Chem. Rev. 117, 7244–7275 (2017).
Fuchs, J. et al. A photoactivatable marker protein for pulse-chase imaging with superresolution. Nat. Methods 7, 627–630 (2010).
Zhu, L., Zhang, W., Elnatan, D. & Huang, B. Faster STORM using compressed sensing. Nat. Methods 9, 721–723 (2012).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Hoze, N. et al. Heterogeneity of AMPA receptor trafficking and molecular interactions revealed by superresolution analysis of live cell imaging. Proc. Natl Acad. Sci. USA 109, 17052–17057 (2012).
Yang, T. T., Tran, M. N. T., Chong, W. M., Huang, C. E. & Liao, J. C. Single-particle tracking localization microscopy reveals nonaxonemal dynamics of intraflagellar transport proteins at the base of mammalian primary cilia. Mol. Biol. Cell 30, 828–837 (2019).
Izeddin, I. et al. Wavelet analysis for single molecule localization microscopy. Opt. Express 20, 2081–2095 (2012).
Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Heintzmann, R. & Cremer, C. G. in Optical Biopsies and Microscopic Techniques III 185–196 (International Society for Optics and Photonics, 1999).
Gustafsson, M. G. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008).
Schermelleh, L. et al. Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320, 1332–1336 (2008).
Heintzmann, R. & Ficz, G. Breaking the resolution limit in light microscopy. Brief. Funct. Genomic. Proteomic. 5, 289–301 (2006).
Vicidomini, G., Bianchini, P. & Diaspro, A. STED super-resolved microscopy. Nat. Methods 15, 173–182 (2018).
Westphal, V., Lauterbach, M. A., Di Nicola, A. & Hell, S. W. Dynamic far-field fluorescence nanoscopy. N. J. Phys. 9, 435–435 (2007).
Willig, K., Keller, J., Bossi, M. & Hell, S. W. STED microscopy resolves nanoparticle assemblies. N. J. Phys. 8, 106 (2006).
Willig, K. I., Rizzoli, S. O., Westphal, V., Jahn, R. & Hell, S. W. STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis. Nature 440, 935–939 (2006).
Dyba, M. & Hell, S. W. Focal spots of size lambda/23 open up far-field florescence microscopy at 33 nm axial resolution. Phys. Rev. Lett. 88, 163901 (2002).
Dyba, M., Jakobs, S. & Hell, S. W. Immunofluorescence stimulated emission depletion microscopy. Nat. Biotechnol. 21, 1303–1304 (2003).
Dyba, M., Keller, J. & Hell, S. W. Phase filter enhanced STED-4Pi fluorescence microscopy: theory and experiment. N. J. Phys. 7, 134–134 (2005).
Klar, T. A., Jakobs, S., Dyba, M., Egner, A. & Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl Acad. Sci. USA 97, 8206–8210 (2000).
Sidenstein, S. C. et al. Multicolour multilevel STED nanoscopy of actin/spectrin organization at synapses. Sci. Rep. 6, 26725 (2016).
Kilian, N. et al. Assessing photodamage in live-cell STED microscopy. Nat. Methods 15, 755–756 (2018).
Lauterbach, M. A. et al. Comparing video-rate STED nanoscopy and confocal microscopy of living neurons. J. Biophotonics 3, 417–424 (2010).
Pellett, P. A. et al. Two-color STED microscopy in living cells. Biomed. Opt. Express 2, 2364–2371 (2011).
Gwosch, K. C. et al. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nat. Methods 17, 217–224 (2020).
Ma, J. & Yang, W. Three-dimensional distribution of transient interactions in the nuclear pore complex obtained from single-molecule snapshots. Proc. Natl Acad. Sci. USA 107, 7305–7310 (2010).
Goryaynov, A., Ma, J. & Yang, W. Single-molecule studies of nucleocytoplasmic transport: from one dimension to three dimensions. Integr. Biol. 4, 10–21 (2012).
Goryaynov, A. & Yang, W. Role of molecular charge in nucleocytoplasmic transport. PLoS ONE 9, e88792 (2014).
Junod, S. L., Kelich, J. M., Ma, J. & Yang, W. Nucleocytoplasmic transport of intrinsically disordered proteins studied by high-speed super-resolution microscopy. Protein Sci. 29, 1459–1472 (2020).
Luo, W. et al. Axonemal lumen dominates cytosolic protein diffusion inside the primary cilium. Sci. Rep. 7, 1–11 (2017).
Ma, J., Goryaynov, A., Sarma, A. & Yang, W. Self-regulated viscous channel in the nuclear pore complex. Proc. Natl Acad. Sci. USA 109, 7326–7331 (2012).
Ma, J., Goryaynov, A. & Yang, W. Super-resolution 3D tomography of interactions and competition in the nuclear pore complex. Nat. Struct. Mol. Biol. 23, 239–247 (2016).
Ma, J. et al. High-resolution three-dimensional mapping of mRNA export through the nuclear pore. Nat. Commun. 4, 1–9 (2013).
Mudumbi, K. C. et al. Nucleoplasmic signals promote directed transmembrane protein import simultaneously via multiple channels of nuclear pores. Nat. Commun. 11, 1–14 (2020).
Westphal, V. et al. Video-rate far-field optical nanoscopy dissects synaptic vesicle movement. Science 320, 246–249 (2008).
Li, Y., Luo, W. & Yang, W. Nuclear transport and accumulation of smad proteins studied by single-molecule microscopy. Biophys. J. 114, 2243–2251 (2018).
Sun, C., Yang, W., Tu, L.-C. & Musser, S. M. Single-molecule measurements of importin α/cargo complex dissociation at the nuclear pore. Proc. Natl Acad. Sci. USA 105, 8613–8618 (2008).
Yang, W. & Musser, S. M. Visualizing single molecules interacting with nuclear pore complexes by narrow-field epifluorescence microscopy. Methods 39, 316–328 (2006).
Solovei, I. et al. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152, 584–598 (2013).
Mudumbi, K. C., Yang, W., Ma, J. & Schirmer, E. C. Single-point frap distinguishes inner and outer nuclear membrane protein distribution. Biophys. J. 110, 596a (2016).
Daigle, N. et al. Nuclear pore complexes form immobile networks and have a very low turnover in live mammalian cells. J. Cell Biol. 154, 71–84 (2001).
Feldherr, C. M., Kallenbach, E. & Schultz, N. Movement of a karyophilic protein through the nuclear pores of oocytes. J. Cell Biol. 99, 2216–2222 (1984).
Kubitscheck, U., Wedekind, P., Zeidler, O., Grote, M. & Peters, R. Single nuclear pores visualized by confocal microscopy and image processing. Biophys. J. 70, 2067–2077 (1996).
Tokunaga, M., Imamoto, N. & Sakata-Sogawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat. Methods 5, 159–161 (2008).
Mudumbi, K. C., Schirmer, E. C. & Yang, W. Single-point single-molecule FRAP distinguishes inner and outer nuclear membrane protein distribution. Nat. Commun. 7, 1–6 (2016).
Tingey, M., Mudumbi, K. C., Schirmer, E. C. & Yang, W. Casting a wider net: differentiating between inner nuclear envelope and outer nuclear envelope transmembrane proteins. Int. J. Mol. Sci. 20, 5248 (2019).
Ott, C. & Lippincott-Schwartz, J. Visualization of live primary cilia dynamics using fluorescence microscopy. Curr. Protoc. Cell Biol. Chapter 4, Unit 4 26 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Grünwald, D. & Singer, R. H. In vivo imaging of labelled endogenous β-actin mRNA during nucleocytoplasmic transport. Nature 467, 604–607 (2010).
Herbert, A. Single Molecule Light Microscopy ImageJ Plugins. http://www.sussex.ac.uk/gdsc/intranet/pdfs/SMLM.pdf (2014).
Punge, A. et al. 3D reconstruction of high-resolution STED microscope images. Microsc. Res. Tech. 71, 644–650 (2008).
Banterle, N., Bui, K. H., Lemke, E. A. & Beck, M. Fourier ring correlation as a resolution criterion for super-resolution microscopy. J. Struct. Biol. 183, 363–367 (2013).
Nieuwenhuizen, R. P. et al. Measuring image resolution in optical nanoscopy. Nat. Methods 10, 557–562 (2013).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Agasti, S. S. et al. DNA-barcoded labeling probes for highly multiplexed Exchange-PAINT imaging. Chem. Sci. 8, 3080–3091 (2017).
Bates, M., Blosser, T. R. & Zhuang, X. Short-range spectroscopic ruler based on a single-molecule optical switch. Phys. Rev. Lett. 94, 108101 (2005).
Bates, M. et al. in Single Molecule Spectroscopy in Chemistry, Physics and Biology 399–415 (Springer, 2010).
Gordon, M. P., Ha, T. & Selvin, P. R. Single-molecule high-resolution imaging with photobleaching. Proc. Natl Acad. Sci. USA 101, 6462–6465 (2004).
Heilemann, M., Margeat, E., Kasper, R., Sauer, M. & Tinnefeld, P. Carbocyanine dyes as efficient reversible single-molecule optical switch. J. Am. Chem. Soc. 127, 3801–3806 (2005).
Kubitscheck, U. et al. Nuclear transport of single molecules: dwell times at the nuclear pore complex. J. Cell Biol. 168, 233–243 (2005).
Ober, R. J., Ram, S. & Ward, E. S. Localization accuracy in single-molecule microscopy. Biophys. J. 86, 1185–1200 (2004).
Qu, X., Wu, D., Mets, L. & Scherer, N. F. Nanometer-localized multiple single-molecule fluorescence microscopy. Proc. Natl Acad. Sci. USA 101, 11298–11303 (2004).
Owen, D. M., Williamson, D., Magenau, A., Rossy, J. & Gaus, K. in Methods in Enzymology Vol. 504 221–235 (Elsevier, 2012).
York, A. G., Ghitani, A., Vaziri, A., Davidson, M. W. & Shroff, H. Confined activation and subdiffractive localization enables whole-cell PALM with genetically expressed probes. Nat. Methods 8, 327–333 (2011).
Temprine, K., York, A. G. & Shroff, H. Three-dimensional photoactivated localization microscopy with genetically expressed probes. Methods Mol. Biol. 1251, 231–261 (2015).
Sharonov, A. & Hochstrasser, R. M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl Acad. Sci. USA 103, 18911–18916 (2006).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Iinuma, R. et al. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 344, 65–69 (2014).
Schueder, F. et al. Multiplexed 3D super-resolution imaging of whole cells using spinning disk confocal microscopy and DNA-PAINT. Nat. Commun. 8, 1–9 (2017).
Lin, R., Clowsley, A. H., Lutz, T., Baddeley, D. & Soeller, C. 3D super-resolution microscopy performance and quantitative analysis assessment using DNA-PAINT and DNA origami test samples. Methods 174, 56–71 (2020).
Strauss, S. et al. Modified aptamers enable quantitative sub-10-nm cellular DNA-PAINT imaging. Nat. Methods 15, 685–688 (2018).
Nieves, D. J., Gaus, K. & Baker, M. A. DNA-based super-resolution microscopy: DNA-PAINT. Genes 9, 621 (2018).
Chung, E., Kim, D. & So, P. T. Extended resolution wide-field optical imaging: objective-launched standing-wave total internal reflection fluorescence microscopy. Opt. Lett. 31, 945–947 (2006).
Richter, V., Piper, M., Wagner, M. & Schneckenburger, H. Increasing resolution in live cell microscopy by structured illumination (SIM). Appl. Sci. 9, 1188 (2019).
Förster, R. et al. Simple structured illumination microscope setup with high acquisition speed by using a spatial light modulator. Opt. Express 22, 20663–20677 (2014).
Strohl, F. & Kaminski, C. F. Speed limits of structured illumination microscopy. Opt. Lett. 42, 2511–2514 (2017).
Schmidt, R. et al. Spherical nanosized focal spot unravels the interior of cells. Nat. Methods 5, 539–544 (2008).
Hell, S. W. Toward fluorescence nanoscopy. Nat. Biotechnol. 21, 1347–1355 (2003).
Manley, S., Gillette, J. M. & Lippincott-Schwartz, J. in Methods in Enzymology Vol. 475 109–120 (Elsevier, 2010).
Subach, F. V., Patterson, G. H., Renz, M., Lippincott-Schwartz, J. & Verkhusha, V. V. Bright monomeric photoactivatable red fluorescent protein for two-color super-resolution sptPALM of live cells. J. Am. Chem. Soc. 132, 6481–6491 (2010).
Ruba, A., Luo, W., Kelich, J., Tingey, M. & Yang, W. 3D tracking-free approach for obtaining 3D super-resolution information in rotationally symmetric biostructures. J. Phys. Chem. B 123, 5107–5120 (2019).
Acknowledgements
The authors acknowledge S. J. Junod for his assistance during the preparation of this manuscript. We also thank the many collaborators who have greatly helped us complete the research included in this manuscript: E. C. Schirmer (University of Edinburgh), K. J. Verhey (University of Michigan), N. G. Walter (University of Michigan) and R. Y. H. Lim (University of Basel). The project was supported by grants from the National Institutes of Health (GM094041, GM097037, GM116204 and GM22552 to W.Y.).
Author information
Authors and Affiliations
Contributions
Y.L., M.T. and A.R. produced the figures and tables and wrote and edited the manuscript. W.Y. edited the manuscript and provided appropriate guidance.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Alex Clowsley, Christian Soeller and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Ma, J. & Yang, W. Proc. Natl Acad. Sci. USA 107, 7305–7310 (2010): https://doi.org/10.1073/pnas.0908269107
Ma, J. et al. Nat. Struct. Mol. Biol. 23, 239–247 (2016): https://doi.org/10.1038/nsmb.3174
Luo, W. et al. Sci. Rep. 7, 15793 (2017): https://doi.org/10.1038/s41598-017-16103-z
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Equations 1–4.
Rights and permissions
About this article
Cite this article
Li, Y., Tingey, M., Ruba, A. et al. High-speed super-resolution imaging of rotationally symmetric structures using SPEED microscopy and 2D-to-3D transformation. Nat Protoc 16, 532–560 (2021). https://doi.org/10.1038/s41596-020-00440-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00440-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.