Abstract
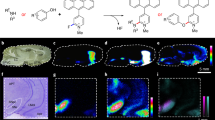
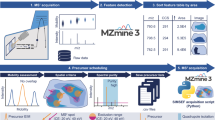
Molecule-specific techniques such as MALDI and desorption electrospray ionization mass spectrometry imaging enable direct and simultaneous mapping of biomolecules in tissue sections in a single experiment. However, neurotransmitter imaging in the complex environment of biological samples remains challenging. Our covalent charge-tagging approach using on-tissue chemical derivatization of primary and secondary amines and phenolic hydroxyls enables comprehensive mapping of neurotransmitter networks. Here, we present robust and easy-to-use chemical derivatization protocols that facilitate quantitative and simultaneous molecular imaging of complete neurotransmitter systems and drugs in diverse biological tissue sections with high lateral resolution. This is currently not possible with any other imaging technique. The protocol, using fluoromethylpyridinium and pyrylium reagents, describes all steps from tissue preparation (~1 h), chemical derivatization (1–2 h), data collection (timing depends on the number of samples and lateral resolution) and data analysis and interpretation. The specificity of the chemical reaction can also help users identify unknown chemical identities. Our protocol can reveal the cellular locations in which signaling molecules act and thus shed light on the complex responses that occur after the administration of drugs or during the course of a disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw datasets for both MALDI and DESI-MSI experiments are too large to be publicly shared but are available for research purposes from the corresponding author upon reasonable request. Source data for Table 1 and Supplementary Table 1 are provided as Supplementary Data.
References
Kandel, E. R., Schwartz, J. H. & Jessell, T. M. Principles of Neural Science 4th edn (McGraw-Hill: 2000).
Ng, J., Papandreou, A., Heales, S. J. & Kurian, M. A. Monoamine neurotransmitter disorders—clinical advances and future perspectives. Nat. Rev. Neurol. 11, 567–584 (2015).
Ganesana, M., Lee, S. T., Wang, Y. & Venton, B. J. Analytical techniques in neuroscience: recent advances in imaging, separation, and electrochemical methods. Anal. Chem. 89, 314–341 (2017).
Robinson, D. L., Hermans, A., Seipel, A. T. & Wightman, R. M. Monitoring rapid chemical communication in the brain. Chem. Rev. 108, 2554–2584 (2008).
Shariatgorji, M., Svenningsson, P. & Andren, P. E. Mass spectrometry imaging, an emerging technology in neuropsychopharmacology. Neuropsychopharmacology 39, 34–49 (2014).
Caprioli, R. M., Farmer, T. B. & Gile, J. Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal. Chem. 69, 4751–4760 (1997).
Wiseman, J. M., Ifa, D. R., Song, Q. & Cooks, R. G. Tissue imaging at atmospheric pressure using desorption electrospray ionization (DESI) mass spectrometry. Angew. Chem. Int. Ed. Engl. 45, 7188–7192 (2006).
Tareke, E., Bowyer, J. F. & Doerge, D. R. Quantification of rat brain neurotransmitters and metabolites using liquid chromatography/electrospray tandem mass spectrometry and comparison with liquid chromatography/electrochemical detection. Rapid Commun. Mass Spectrom. 21, 3898–3904 (2007).
Norris, J. L. & Caprioli, R. M. Analysis of tissue specimens by matrix-assisted laser desorption/ionization imaging mass spectrometry in biological and clinical research. Chem. Rev. 113, 2309–2342 (2013).
Bagley, M. C., Ekelof, M., Rock, K., Patisaul, H. & Muddiman, D. C. IR-MALDESI mass spectrometry imaging of underivatized neurotransmitters in brain tissue of rats exposed to tetrabromobisphenol A. Anal. Bioanal. Chem. 410, 7979–7986 (2018).
Fernandes, A. M. et al. Direct visualization of neurotransmitters in rat brain slices by desorption electrospray ionization mass spectrometry imaging (DESI–MS). J. Am. Soc. Mass Spectrom. 27, 1944–1951 (2016).
Passarelli, M. K. et al. The 3D OrbiSIMS-label-free metabolic imaging with subcellular lateral resolution and high mass-resolving power. Nat. Methods 14, 1175–1183 (2017).
Pace, C. L., Horman, B., Patisaul, H. & Muddiman, D. C. Analysis of neurotransmitters in rat placenta exposed to flame retardants using IR-MALDESI mass spectrometry imaging. Anal. Bioanal. Chem. 412, 3745–3752 (2020).
Esteve, C., Tolner, E. A., Shyti, R., van den Maagdenberg, A. M. & McDonnell, L. A. Mass spectrometry imaging of amino neurotransmitters: a comparison of derivatization methods and application in mouse brain tissue. Metabolomics 12, 30 (2016).
Kaya, I. et al. On-tissue chemical derivatization of catecholamines using 4-(N-Methyl)pyridinium boronic acid for ToF-SIMS and LDI-ToF mass spectrometry imaging. Anal. Chem. 90, 13580–13590 (2018).
Zenobi, R. & Knochenmuss, R. Ion formation in MALDI mass spectrometry. Mass Spectrom. Rev. 17, 337–366 (1998).
Shariatgorji, M. et al. Direct targeted quantitative molecular imaging of neurotransmitters in brain tissue sections. Neuron 84, 697–707 (2014).
Shariatgorji, M. et al. Simultaneous imaging of multiple neurotransmitters and neuroactive substances in the brain by desorption electrospray ionization mass spectrometry. Neuroimage 136, 129–138 (2016).
Ito, T. & Hiramoto, M. Use of mTRAQ derivatization reagents on tissues for imaging neurotransmitters by MALDI imaging mass spectrometry: the triple spray method. Anal. Bioanal. Chem. 411, 6847–6856 (2019).
Manier, M. L., Spraggins, J. M., Reyzer, M. L., Norris, J. L. & Caprioli, R. M. A derivatization and validation strategy for determining the spatial localization of endogenous amine metabolites in tissues using MALDI imaging mass spectrometry. J. Mass Spectrom. 49, 665–673 (2014).
Shariatgorji, M. et al. Comprehensive mapping of neurotransmitter networks by MALDI-MS imaging. Nat. Methods 16, 1021–1028 (2019).
Hulme, H. et al. Microbiome-derived carnitine mimics as previously unknown mediators of gut-brain axis communication. Sci. Adv. 6, eaax6328 (2020).
Jiang, S. H., Hu, L. P., Wang, X., Li, J. & Zhang, Z. G. Neurotransmitters: emerging targets in cancer. Oncogene 39, 503–515 (2020).
Falck, B., Thieme, G., Hillarp, N. A. & Torp, A. Fluorescence of catechol amines and related compounds condensed with formaldehyde. J. Histochem. Cytochem. 10, 348–354 (1962).
Falck, B. & Torp, A. New evidence for the localization of noradrenalin in the adrenergic nerve terminals. Med. Exp. Int. J. Exp. Med. 6, 169–172 (1962).
Pradhan, T. et al. Chemical sensing of neurotransmitters. Chem. Soc. Rev. 43, 4684–4713 (2014).
Fritschy, J. M. Is my antibody-staining specific? How to deal with pitfalls of immunohistochemistry. Eur. J. Neurosci. 28, 2365–2370 (2008).
Kenigsberg, R. L. & Cuello, A. C. Role of immunology in defining transmitter-specific neurons. Immunol. Rev. 100, 279–306 (1987).
Lee, G. J., Park, J. H. & Park, H. K. Microdialysis applications in neuroscience. Neurol. Res. 30, 661–668 (2008).
Bucher, E. S. & Wightman, R. M. Electrochemical analysis of neurotransmitters. Annu. Rev. Anal. Chem. 8, 239–261 (2015).
Ametamey, S. M., Honer, M. & Schubiger, P. A. Molecular imaging with PET. Chem. Rev. 108, 1501–1516 (2008).
Elsinga, P. H., Hatano, K. & Ishiwata, K. PET tracers for imaging of the dopaminergic system. Curr. Med. Chem. 13, 2139–2153 (2006).
Novotny, E. J. Jr., Fulbright, R. K., Pearl, P. L., Gibson, K. M. & Rothman, D. L. Magnetic resonance spectroscopy of neurotransmitters in human brain. Ann. Neurol. 54, S25–S31 (2003).
Bowman, A. P. et al. Ultra-high mass resolving power, mass accuracy, and dynamic range MALDI mass spectrometry imaging by 21-T FT-ICR MS. Anal. Chem. 92, 3133–3142 (2020).
Kompauer, M., Heiles, S. & Spengler, B. Atmospheric pressure MALDI mass spectrometry imaging of tissues and cells at 1.4-μm lateral resolution. Nat. Methods 14, 90–96 (2017).
Ràfols, P. et al. Signal preprocessing, multivariate analysis and software tools for MA(LDI)-TOF mass spectrometry imaging for biological applications. Mass Spectrom. Rev. 37, 281–306 (2018).
Alexandrov, T. Spatial metabolomics and imaging mass spectrometry in the age of artificial intelligence. Annu. Rev. Biomed. Data Sci. 3, 61–87 (2020).
Van de Plas, R. et al. Prospective exploration of biochemical tissue composition via imaging mass spectrometry guided by principal component analysis. Pac. Symp. Biocomput. 2007, 458–469 (2007).
Verbeeck, N., Caprioli, R. M. & Van de Plas, R. Unsupervised machine learning for exploratory data analysis in imaging mass spectrometry. Mass Spectrom. Rev. 39, 245–291 (2020).
Goodwin, R. J., Iverson, S. L. & Andren, P. E. The significance of ambient-temperature on pharmaceutical and endogenous compound abundance and distribution in tissues sections when analyzed by matrix-assisted laser desorption/ionization mass spectrometry imaging. Rapid Commun. Mass Spectrom. 26, 494–498 (2012).
Sköld, K. et al. The significance of biochemical and molecular sample integrity in brain proteomics and peptidomics: stathmin 2-20 and peptides as sample quality indicators. Proteomics 7, 4445–4456 (2007).
Nelson, K. A., Daniels, G. J., Fournie, J. W. & Hemmer, M. J. Optimization of whole-body zebrafish sectioning methods for mass spectrometry imaging. J. Biomol. Tech. 24, 119–127 (2013).
Schwartz, S. A., Reyzer, M. L. & Caprioli, R. M. Direct tissue analysis using matrix-assisted laser desorption/ionization mass spectrometry: practical aspects of sample preparation. J. Mass Spectrom. 38, 699–708 (2003).
Dannhorn, A. et al. Universal sample preparation unlocking multimodal molecular tissue imaging. Anal. Chem. 92, 11080–11088 (2020).
Kahn, J. & Schemmer, P. Comprehensive review on Custodiol-N (HTK-N) and its molecular side of action for organ preservation. Curr. Pharm. Biotechnol. 18, 1237–1248 (2017).
Shariatgorji, M. et al. Pyrylium salts as reactive matrices for MALDI-MS imaging of biologically active primary amines. J. Am. Soc. Mass Spectrom. 26, 934–939 (2015).
Chumbley, C. W. et al. Absolute quantitative MALDI imaging mass spectrometry: a case of rifampicin in liver tissues. Anal. Chem. 88, 2392–2398 (2016).
Hamm, G. et al. Quantitative mass spectrometry imaging of propranolol and olanzapine using tissue extinction calculation as normalization factor. J. Proteom. 75, 4952–4961 (2012).
Nilsson, A. et al. Fine mapping the spatial distribution and concentration of unlabeled drugs within tissue micro-compartments using imaging mass spectrometry. PLoS ONE 5, e11411 (2010).
Prentice, B. M., Chumbley, C. W. & Caprioli, R. M. Absolute quantification of rifampicin by MALDI imaging mass spectrometry using multiple TOF/TOF events in a single laser shot. J. Am. Soc. Mass Spectrom. 28, 136–144 (2017).
Takai, N., Tanaka, Y., Inazawa, K. & Saji, H. Quantitative analysis of pharmaceutical drug distribution in multiple organs by imaging mass spectrometry. Rapid Commun. Mass Spectrom. 26, 1549–1556 (2012).
Kallback, P. et al. Cross-validated matrix-assisted laser desorption/ionization mass spectrometry imaging quantitation protocol for a pharmaceutical drug and its drug-target effects in the brain using time-of-flight and Fourier transform ion cyclotron resonance analyzers. Anal. Chem. 92, 14676–14684 (2020).
Kallback, P., Shariatgorji, M., Nilsson, A. & Andren, P. E. Novel mass spectrometry imaging software assisting labeled normalization and quantitation of drugs and neuropeptides directly in tissue sections. J. Proteom. 75, 4941–4951 (2012).
Pirman, D. A. & Yost, R. A. Quantitative tandem mass spectrometric imaging of endogenous acetyl-L-carnitine from piglet brain tissue using an internal standard. Anal. Chem. 83, 8575–8581 (2011).
Porta, T., Lesur, A., Varesio, E. & Hopfgartner, G. Quantification in MALDI-MS imaging: what can we learn from MALDI-selected reaction monitoring and what can we expect for imaging? Anal. Bioanal. Chem. 407, 2177–2187 (2015).
Kilkenny, C., Browne, W. J., Cuthill, I. C., Emerson, M. & Altman, D. G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Abbassi-Ghadi, N. et al. Repeatability and reproducibility of desorption electrospray ionization-mass spectrometry (DESI-MS) for the imaging analysis of human cancer tissue: a gateway for clinical applications. Anal. Methods 7, 71–80 (2015).
Takats, Z., Wiseman, J. M., Gologan, B. & Cooks, R. G. Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science 306, 471–473 (2004).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Race, A. M., Styles, I. B. & Bunch, J. Inclusive sharing of mass spectrometry imaging data requires a converter for all. J. Proteom. 75, 5111–5112 (2012).
Kallback, P., Nilsson, A., Shariatgorji, M. & Andren, P. E. msIQuant—quantitation software for mass spectrometry imaging enabling fast access, visualization, and analysis of large data sets. Anal. Chem. 88, 4346–4353 (2016).
Wishart, D. S. et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 46, D608–D617 (2018).
Palmer, A. et al. FDR-controlled metabolite annotation for high-resolution imaging mass spectrometry. Nat. Methods 14, 57–60 (2017).
Shariatgorji, R. et al. Bromopyrylium derivatization facilitates identification by mass spectrometry imaging of monoamine neurotransmitters and small molecule neuroactive compounds. J. Am. Soc. Mass Spectrom. 31, 2551–2557 (2020).
Fridjonsdottir, E. et al. Mass spectrometry imaging identifies abnormally elevated brain L-DOPA levels and extra-striatal monoaminergic dysregulation in L-DOPA-induced dyskinesia. Sci. Adv. 7, eabe5948 (2021).
Acknowledgements
This work was financially supported by the Swedish Research Council (Medicine and Health Grant 2018–03320 and Natural and Engineering Science Grants 2018–05501 and 2018-05133), the Swedish Brain Foundation (FO2018-0292), the Swedish Foundation for Strategic Research (RIF14-0078) and the Science for Life Laboratory (SciLifeLab).
Author information
Authors and Affiliations
Contributions
R.S. and A.N. conceived the methodology; designed experiments; acquired, analyzed and interpreted data; and wrote the manuscript. E.F. analyzed data and wrote part of the manuscript. N.S. and A.D. designed experiments; acquired, analyzed and interpreted data; and wrote part of the manuscript. P.S. provided tissue samples, supervised animal experiments and edited the manuscript. R.J.A.G. designed experiments, analyzed and interpreted data and wrote part of the manuscript. L.R.O. conceived the methodology, designed experiments, synthesized the reactive matrices, interpreted data and edited the manuscript. P.E.A. conceived the methodology, designed experiments, interpreted data, wrote the manuscript and is principal investigator for the grants that fund this research.
Corresponding author
Ethics declarations
Competing interests
N.S, A.D. and R.J.A.G. are full-time salaried employees, own stocks of AstraZeneca and performed this study as part of their regular duties. None of the authors received any form of royalty or further financial compensation from AstraZeneca for publishing these protocols. L.R.O., A.N., R. S. and P.E.A. are co-founders and shareholders in Tag-ON AB and have filed a patent application ‘Reactive desorption and/or laser ablation ionization matrices and use thereof’, no. PCT/SE2019/050197 (patent pending).
Additional information
Peer review information Nature Protocols thanks Axel Karl Walch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Shariatgorji, M. et al. Nat. Methods 16, 1021–1028 (2019): https://doi.org/10.1038/s41592-019-0551-3
Shariatgorji, M. et al. Neuron 84, 697–707 (2014): https://doi.org/10.1016/j.neuron.2014.10.011
Shariatgorji, M. et al. Neuroimage 136, 129–38 (2016): https://doi.org/10.1016/j.neuroimage.2016.05.004
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Supplementary Tables 1–3 and Supplementary References.
Supplementary Data
Mass accuracy calculations, red phosphorus mass reference table and source data for LOD calculations
Rights and permissions
About this article
Cite this article
Shariatgorji, R., Nilsson, A., Fridjonsdottir, E. et al. Spatial visualization of comprehensive brain neurotransmitter systems and neuroactive substances by selective in situ chemical derivatization mass spectrometry imaging. Nat Protoc 16, 3298–3321 (2021). https://doi.org/10.1038/s41596-021-00538-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00538-w
This article is cited by
-
Two-photon fluorescence imaging and specifically biosensing of norepinephrine on a 100-ms timescale
Nature Communications (2023)
-
Visualization of metabolites and microbes at high spatial resolution using MALDI mass spectrometry imaging and in situ fluorescence labeling
Nature Protocols (2023)
-
Prosaposin maintains lipid homeostasis in dopamine neurons and counteracts experimental parkinsonism in rodents
Nature Communications (2023)
-
Chemical tagging mass spectrometry: an approach for single-cell omics
Analytical and Bioanalytical Chemistry (2023)
-
Mass spectrometry imaging: new eyes on natural products for drug research and development
Acta Pharmacologica Sinica (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.