Abstract
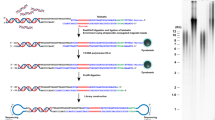
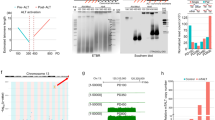
Here we provide a detailed protocol for the enrichment of telomeric repeats from mouse and human cells. The procedure consists of two successive rounds of digestion with frequently cutting restriction enzymes followed by size fractionation. Around 2 mg of genomic DNA is required, and the procedure lasts 5–6 d and yields preparations enriched >800-fold in telomeres. The purified material is suitable for single-molecule analysis of telomere structure, visualizing telomere replication and recombination intermediates by electron microscopy or performing molecular combing at telomeric repeats. No special skills are required for the enrichment procedure, while some assistance is needed in harvesting a large number of plates in a timely fashion at the beginning of the procedure. A smaller-scale version of the protocol that involves one round of digestion and purification requires 200 µg of DNA and enriches telomeres ~50-fold in 4 d is also provided. The latter can be combined with specific labeling for single-molecule analysis of replicating DNA or for long-read sequencing analysis of telomeric repeats. The procedure described here can be adapted to the enrichment of other repetitive elements, based on the use of restriction enzymes that do not cut into the repeat of interest.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data generated or analyzed during this work are included in the published article. Source data are provided with this paper.
References
Lansdorp, P. M. et al. Heterogeneity in telomere length of human chromosomes. Hum. Mol. Genet 5, 685–691 (1996).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Miller, K. M., Rog, O. & Cooper, J. P. Semi-conservative DNA replication through telomeres requires Taz1. Nature 440, 824–828 (2006).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Allshire, R. C. et al. Telomeric repeat from T. thermophila cross hybridizes with human telomeres. Nature 332, 656–659 (1988).
Moyzis, R. K. et al. A highly conserved repetitive DNA sequence, (TTAGGG)n, present at the telomeres of human chromosomes. Proc. Natl Acad. Sci. USA. 85, 6622–6626 (1988).
de Lange, T. et al. Structure and variability of human chromosome ends. Mol. Cell Biol. 10, 518–527 (1990).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Tomáška, Ľ., Cesare, A. J., AlTurki, T. M. & Griffith, J. D. Twenty years of t-loops: a case study for the importance of collaboration in molecular biology. DNA Repair 94, 102901 (2020).
Munoz-Jordan, J. L., Cross, G. A., de Lange, T. & Griffith, J. D. t-loops at trypanosome telomeres. EMBO J. 20, 579–588 (2001).
Cesare, A. J., Quinney, N., Willcox, S., Subramanian, D. & Griffith, J. D. Telomere looping in P. sativum (common garden pea). Plant J. 36, 271–279 (2003).
Cesare, A. J. & Griffith, J. D. Telomeric DNA in ALT cells is characterized by free telomeric circles and heterogeneous t-loops. Mol. Cell Biol. 24, 9948–9957 (2004).
Cesare, A. J., Groff-Vindman, C., Compton, S. A., McEachern, M. J. & Griffith, J. D. Telomere loops and homologous recombination-dependent telomeric circles in a Kluyveromyces lactis telomere mutant strain. Mol. Cell Biol. 28, 20–29 (2008).
Raices, M. et al. C. elegans telomeres contain G-strand and C-strand overhangs that are bound by distinct proteins. Cell 132, 745–757 (2008).
Drosopoulos, W. C., Kosiyatrakul, S. T., Yan, Z., Calderano, S. G. & Schildkraut, C. L. Human telomeres replicate using chromosome-specific, rather than universal, replication programs. J. Cell Biol. 197, 253–266 (2012).
Dilley, R. L. et al. Break-induced telomere synthesis underlies alternative telomere maintenance. Nature 539, 54–58 (2016).
Mateos-Gomez, P. A. et al. Mammalian polymerase theta promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Baur, J. A., Wright, W. E. & Shay, J. W. Analysis of mammalian telomere position effect. Methods Mol. Biol. 287, 121–136 (2004).
Fouquerel, E., Barnes, R. P., Wang, H. & Opresko, P. L. Measuring UV photoproduct repair in isolated telomeres and bulk genomic DNA. Methods Mol. Biol. 1999, 295–306 (2019).
Mazzucco, G. et al. Telomere damage induces internal loops that generate telomeric circles. Nat. Commun. 11, 5297 (2020).
Lopes, M. Electron microscopy methods for studying in vivo DNA replication intermediates. Methods Mol. Biol. 521, 605–631 (2009).
Kimura, M. et al. Measurement of telomere length by the Southern blot analysis of terminal restriction fragment lengths. Nat. Protoc. 5, 1596–1607 (2010).
Conomos, D. et al. Variant repeats are interspersed throughout the telomeres and recruit nuclear receptors in ALT cells. J. Cell Biol. 199, 893–906 (2012).
Simpson, J. T. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat. Methods 14, 407–410 (2017).
Rand, A. C. et al. Mapping DNA methylation with high-throughput nanopore sequencing. Nat. Methods 14, 411–413 (2017).
An, N., Fleming, A. M., White, H. S. & Burrows, C. J. Nanopore detection of 8-oxoguanine in the human telomere repeat sequence. ACS Nano 9, 4296–4307 (2015).
Müller, C. A. et al. Capturing the dynamics of genome replication on individual ultra-long nanopore sequence reads. Nat. Methods 16, 429–436 (2019).
Hennion, M. et al. FORK-seq: replication landscape of the Saccharomyces cerevisiae genome by nanopore sequencing. Genome Biol. 21, 125 (2020).
Sinden, R. R., Bat, O. & Kramer, P. R. Psoralen cross-linking as probe of torsional tension and topological domain size in vivo. Methods 17, 112–124 (1999).
Doksani, Y., Wu, J. Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Wu, P., Takai, H. & de Lange, T. Telomeric 3′ overhangs derive from resection by Exo1 and Apollo and fill-in by POT1b-associated CST. Cell 150, 39–52 (2012).
Schwartz, D. C. & Cantor, C. R. Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell 37, 67–75 (1984).
Acknowledgements
We are grateful to D. Piccini, who assisted us in setting up the separation in sucrose gradient. We are grateful to the members of the IFOM Cell Biology Unit for their invaluable assistance with growing large cell cultures. We are grateful to the IFOM Imaging facility for assisting with the acquisition of single-molecule images. We thank C. Villa at IFOM for her assistance and expertise in the preparation of the Supplementary Video files. Y.D. lab is supported by the Associazione Italiana per la Ricerca sul Cancro, AIRC, IG 19901.
Author information
Authors and Affiliations
Contributions
A.H. contributed to the development of the procedure. M.G. performed the experiments of the small-scale enrichment procedure. E.Z. performed the large-scale enrichment procedure on the HT1080 cell line. G.M. performed the other experiments with the large-scale enrichment procedure and prepared the rest of the figures of the manuscript. Y.D. conceived the procedure, oversaw the development of the protocol and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Jack Griffith and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Mazzucco, G. et al. Nat. Commun. 11, 5297 (2020): https://doi.org/10.1038/s41467-020-19139-4
Extended data
Extended Data Fig. 1 Single-molecule analysis of telomeric DNA enrichment.
Single-molecule analysis of telomere enrichment. SV40 MEFs DNA from a non-enriched sample, a sample enriched with the large-scale procedure and a sample enriched with the small-scale procedure were combed on a silanized glass coverslip with the procedure described in Box 2. IF: immunofluorescence, the DNA was denatured in situ and incubated first with an antibody against single-stranded DNA, which labels all DNA molecules (green) and subsequently with a fluorescent (TTAGGG)3 peptide nucleic acid (PNA) probe that labels the telomeric molecules (magenta).
Extended Data Fig. 2 Quantification of telomeric DNA enrichment from HT1080 cells.
Quantification of telomeric DNA enrichment from the human HT1080 cell line in a dot blot. After the large-scale procedure, the indicated amounts of DNA for each enrichment round were loaded on a dot blot and hybridized with a telomeric probe. Fold-enrichment numbers (from one experiment) indicate the telomeric signal per ng of DNA, expressed relative to the value of bulk DNA.
Extended Data Fig. 3 No DNA damage induced during fractionation in sucrose gradient and DNA concentration.
Control experiment showing that separation in sucrose gradient and concentration in Amicon ultra centrifugal filters does not induce detectable DNA damage. Equal amounts of a plasmid prep (Addgene no. 71116) were either stored at 4 °C (input) or separated in a sucrose gradient, collected and concentrated in Amicon ultra centrifugal filters. As a positive control, the same plasmid DNA was incubated with 0.3 µg ml−1 ethidium bromide in TE 1× and irradiated for 10 min with 254 nm UV in a stratalinker. Equal amounts of the three samples were loaded in the agarose gel shown. Note that, in the positive control, treated with UV, all supercoiled form of the plasmid is lost, while no loss of the supercoiled form is detected after separation in sucrose gradient and sample concentration.
Supplementary information
Supplementary Information
Supplementary Methods
Supplementary Video 1
Video showing the psoralen cross-linking procedure
Supplementary Video 2
Video showing the manual preparation of sucrose gradients
Supplementary Video 3
Video showing sample loading on the sucrose gradient
Supplementary Video 4
Video showing the collection of sucrose fractions
Source data
Source Data Fig. 1
Whole gels and membranes used in Fig. 1
Source Data Fig. 3
Whole gels and membranes used in Fig. 3
Source Data Fig. 4
Whole gels and membranes used in Fig. 4
Source Data Fig. 5
Whole gels and membranes used in Fig. 5
Source Data Fig. 6
Whole gel used in Fig. 6
Source Data Fig. 7
Whole membrane used in Fig. 7a
Source Data Fig. 7
Data used for the graph in Fig. 7b
Source Data Fig. 8
Whole gel used in Fig. 8
Source Data Extended Data Fig. 1
Images used in Extended Data Fig. 1
Source Data Extended Data Fig. 2
Whole membrane used in Extended Data Fig. 2
Source Data Extended Data Fig. 3
Whole gel used in Extended Data Fig. 3
Rights and permissions
About this article
Cite this article
Mazzucco, G., Huda, A., Galli, M. et al. Purification of mammalian telomeric DNA for single-molecule analysis. Nat Protoc 17, 1444–1467 (2022). https://doi.org/10.1038/s41596-022-00684-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00684-9
This article is cited by
-
Short-term molecular consequences of chromosome mis-segregation for genome stability
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.