Abstract
RAD52 motif containing 1 (RDM1) is involved in DNA damage repair pathway and RDM1−/− cells increase sensitivity to cisplatin, a common chemotherapy drug. Lung cancer is the leading cause of cancer death worldwide. However, the role of RDM1 in lung cancer is unknown. Here, we find that the mRNA and protein expression levels of RDM1 are significantly increased in human lung tumors, especially in lung adenocarcinoma. The lung adenocarcinoma patients with higher mRNA expression of RDM1 show the worse clinical outcomes. Knockdown of RDM1 in lung adenocarcinoma cells reduces cell proliferation and promotes apoptosis, consistent with the role RDM1 in the overexpression experiments. Xenograft mouse model shows stable knockdown of RDM1 significantly inhibits lung adenocarcinoma tumor growth. These in vitro and in vivo results conclude that RDM1 plays an oncogenic role in human lung adenocarcinoma. Interestingly, P53/RAD51/RAD52 can be regulated by RDM1, and the negative regulation of P53 by RDM1 may be one of major mechanisms for RDM1 to accomplish its oncogenic functions in lung adenocarcinoma. Therefore, RDM1 may be a new target for the treatment of lung adenocarcinoma.
Similar content being viewed by others
Introduction
The risk of cancer can be significantly increased by disruption of genomic integrity resulted from dysfunctional DNA damage response signaling and/or aberrant activity of the key components in the DNA repair pathways. The DNA repair machineries work constantly to remove numerous DNA lesions caused by chemotherapeutic agents such as cisplatin, which contributes to drug resistance in many cancers. As resistance to standard cisplatin-based chemotherapy becomes a frequent phenomenon, cancer treatment targeting important components in the DNA repair pathways emerges to be an imminent and compelling task. RDM1 (RAD52 motif 1, or RD motif) is involved in cellular response to cisplatin, and shows similarities to RAD52, a key regulator in DNA recombination and repair, where the RD motif of RDM1 functionally resembles the N-terminal region of RAD521,2,3. Importantly, RDM1−/− cells exhibited the increased sensitivity to cisplatin4. More interestingly, our initial comprehensive bioinformatics exploration in multiple Oncomine expression datasets has identified RDM1 as one of the significantly up-regulated genes in human lung adenocarcinoma. Despite these discoveries, however, to date, little is known about the role of RDM1 in human cancer. Given the potential role of RDM1 in the DNA repair pathways that constitute an important aspect of cancer initiation and progression, we proposed that RDM1 might display oncogenic properties in lung cancer.
Lung cancer is a leading cause of cancer deaths, and remains one of the refractory cancer types. Lung cancer is divided into two major categories: small cell lung cancer and non-small cell lung cancer (NSCLC)5. Lung adenocarcinoma, one of major subtype of NSCLC, accounts for 40% of all lung cancers. The five-year survival rate of lung cancer is the lowest among the major cancers, including colon, breast, and prostate cancers6. Even with major clinical interventions, such as surgery, radiation therapy, chemotherapy, targeted cancer therapy, and immunotherapy, the survival rate has not been improved significantly, and lingers at only 15% within five years of treatment7. The clinical staging of lung cancers follows the TNM classification system, where the determining factors include: the size of the primary tumor (T), the effects on the regional lymph nodes (N), and the distant metastatic status (M). Recent years have witnessed some successes in targeted therapies for particular mutations in lung adenocarcinoma, such as those in EGFR and ALK, and these strategies have been approved for use as first-line treatment in adenocarcinoma8,9,10. Furthermore, investigation of the mutational landscape in lung adenocarcinoma can add new targets to the growing biomarker panel that may assist with the diagnosis of this cancer. As a result, it is imperative to uncover more novel molecules, which will be beneficial to the treatment and diagnosis of lung adenocarcinoma.
In this study, we found that the mRNA and protein expressions of RDM1 were up-regulated in human lung adenocarcinoma samples. Significantly, up-regulation of RDM1 mRNA level was correlated with poor clinical characteristics and risk factors, including staging, survival, recurrence, and smoking, as demonstrated by multiple Oncomine expression analyses. We knocked down and overexpressed RDM1 in two lung adenocarcinoma cell lines, PC9 and A549, and then evaluated cancer-related phenotypes, including cell proliferation and apoptosis. We further evaluated the in vivo growth of the RDM1-knockdown cells in a mouse xenograft model. The current data support an oncogenic function of RDM1 in human lung adenocarcinoma, supporting by the observation that RDM1 negatively affected the mRNA and protein expression of P53. Our study reveals the oncogenic function of RDM1 in human lung adenocarcinoma.
Results
RDM1 is up-regulated in human lung adenocarcinoma tumors and correlated with poor clinical outcomes
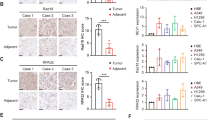
Recent work has revealed high levels of RDM1 in papillary thyroid carcinoma11. But expression of RDM1 in lung cancer remains to be explored. We therefore performed multiple Oncomine analyses in published datasets to examine the RDM1 levels in human lung cancer with various clinical characteristics (Fig. 1)12,13,14,15. Interestingly, RDM1 is significantly over-expressed in lung adenocarcinoma and large cell carcinoma compared with the normal tissues (Fig. 1A). Consistent with the Oncomine results, our immunohistochemistry (IHC) and Western Blot analyses showed the protein level of RDM1 in human lung adenocarcinoma in increased compared to that of adjacent normal lungs (Fig. 1B–D).
RDM1 is up-regulated in human lung adenocarcinoma tumors and correlated with poor clinical outcomes.(A) Oncomine box plots of RDM1 levels in human lung adenocarcinoma (left panel) or large cell lung carcinoma (right panel) and normal tissues. The data resources are listed under each plot. (B) Immunohistochemistry (IHC) analysis of RDM1 in clinical lung adenocarcinoma samples. (C) Statistical analysis of the expression of RDM1 in B (D) Western Blot analysis of analysis of RDM1 in clinical lung adenocarcinoma samples. (E) Oncomine box plots of RDM1 levels in advanced N stage of lung adenocarcinoma. (F) Oncomine box plots of RDM1 levels in lung adenocarcinoma patients deceased at 1 year or 5 years of diagnosis. (G) Oncomine box plots of RDM1 levels in lung adenocarcinoma patients who had recurrent cancer at 3 years or 5 years. (H) Oncomine box plots of RDM1 levels in smokers of lung adenocarcinoma.
Based on these findings, we further examined whether higher expression of RDM1 levels were positively associated with various clinical outcomes in human lung adenocarcinoma. Indeed, higher expression of RDM1 mRNA levels was found in the advanced N stage tumors (1.5 fold or higher in tumors, p < 0.05, Fig. 1E)14. Furthermore, at 1 year or 5 years of diagnosis, RDM1 levels were significantly higher in those who had deceased compared with those who were still alive (1.5 fold or higher, p < 0.01, Fig. 1F). Similar results were also true for patients with recurrence at 3 or 5 years (1.6 fold, p < 10−5, Fig. 1G). Therefore, in general, high RDM1 levels were correlated with poor clinical outcomes in human lung adenocarcinoma. The findings, that RDM1 was over-expressed in lung adenocarcinoma and that high RDM1 was strongly associated with poor clinical outcomes, suggesting the oncogenic role of RDM1 in human lung adenocarcinomas. Finally, we further investigated the expression patterns of RDM1 in smokers since smoking is the leading risk factor of lung cancer. Noticeably, we observed that RDM1 was also highly expressed in lung tumors of smokers compared with nonsmokers, suggesting a positive link between smoking and the RDM1 level (2 folds, p < 0.001, Fig. 1H)16.
RDM1 positively regulates human lung adenocarcinoma cell growth
Given that RDM1 was up-regulated in human lung adenocarcinoma, we first investigated whether RDM1 positively regulated cell proliferation. We used two lung adenocarcinoma cell lines, PC9 and A549, as our in vitro models. The knockdown efficiency was confirmed by qRT-PCR showing that the mRNA levels of RDM1 were significantly inhibited after the transfection of siRNA-RDM1 (siRDM1) compared with that of siRNA-Control (siN) (Fig. 2A). It was evident that, in both cell lines, siRDM1 cells grew significantly slower than siN at later time points (i.e. 72 or 96 h, p < 0.05) (Fig. 2B). These results were further confirmed by clonogenic assay at 72 h under RDM1 knockdown condition (Fig. 2C,D) or overexpression condition (Fig. 2E,F). These results concluded that RDM1 positively regulated the cell proliferation of lung adenocarcinoma.
RDM1 positively regulates lung adenocarcinoma cell growth. (A) Knockdown efficiency of RDM1 was evaluated by qRT-PCR in A549 and PC9 cells. (B) Cell growth for RDM1-knockdown and control cells was recorded as OD450 at 0, 24, 48, 72, and 96 h. (C,D) Clonogenic assay for A549 (C) and PC9 (D) cells at Day 5 after plating the cells with transfection of siRNAs. (E) Overexpression of RDM1 was evaluated by qRT-PCR in A549 and PC9 cells. (F) Clonogenic assay for A549 cell at Day 3 after plating the cells with overexpression of RDM1.
Knockdown of RDM1 inhibits human lung adenocarcinoma cell growth in vivo
To further investigate the oncogenic role of RDM1 in lung adenocarcinoma in vivo, RDM1 was silenced with specific shRNA in A549 cells (A549 shRDM1) (Fig. 3A). Next, we injected A549 shRDM1 cells into 6-week old immunocompromised nude mice and recorded the tumor sizes. At the time of sacrifice, the shRDM1 tumors were significantly smaller compared with the tumors of the control group, confirming the oncogenic in vivo role of RDM1 in lung adenocarcinoma (Fig. 3B,C). IHC staining of RDM1 protein expression was performed to confirm the knockdown efficiency of RDM1 in the shRDM1 tumors (Fig. 3D). Therefore, knockdown of RDM1 inhibited cell growth of lung adenocarcinoma in vivo.
Knockdown of RDM1 inhibits human lung adenocarcinoma cell growth in vivo. (A) Total protein was isolated from stable RDM1- knockdown (shRDM1) cell line and analyzed by immunoblotting with the antibodies against RDM1. (B) A549 cells with the stable RDM1- knockdown (shRDM1) were injected subcutaneously into 6-week old immunocompromised mice. At endpoint, tumors were removed, photographed and measured. (C) The weights of tumors are presented as Mean ± S.D. (n = 5) *p < 0.05. (D) IHC staining of RDM1 was performed for RDM1-knockdown (shRDM1) or control (shN) tumors.
Knockdown of RDM1 induces cell apoptosis in human lung adenocarcinoma cells
Defective growth is usually associated with dysregulated apoptosis, we therefore measured the apoptosis in the RDM1-kockdown cells. The flow cytometry analysis on apoptosis by double-staining of Annexin V-FITC and propidium iodide (PI) showed that siRDM1 cells had more apoptotic populations than siN. For siRDM1 A549 cells (Fig. 4A), more apoptotic cells (Annexin V+/PI− and Annexin V+/PI+ combined) were detected (3.7% in knockdown cells vs. 1.9% in the control cells) (Fig. 4C). Notably, for PC9, the populations of both early stage (Annexin V+/PI−) (3.0%) and late stage apoptotic (Annexin V+/PI+) cells (0.5%) in siRDM1 cells were higher than the control cells (Fig. 4B,D). Consistent with the results of RDM1 knockdown, overexpression of RDM1 reduced the cell apoptosis in A549 and PC cells (Fig. 4E–H). Collectively, RDM1 in lung adenocarcinoma cells negatively regulated apoptosis, further supporting the notion that RDM1 might play an oncogenic role in lung adenocarcinoma cells.
RDM1 negatively regulates cell apoptosis in human lung adenocarcinoma cells. (A) Flow cytometry analysis of Annexin V-FITC and Propidium Iodide (PI) double-stained populations for apoptosis in siRDM1A549 cells. (B) Flow cytometry analysis of Annexin V-FITC and PI double-stained populations for apoptosis in siRDM1 PC9 cells. (C) The ratios of apoptotic cells to all cells were quantified based on three independent experiments of A. Data are expressed as mean ± SD (n = 3). **p < 0.01, siControl versus siRDM1. (D) The ratios of apoptotic cells to all cells were quantified based on three independent experiments of B. Data are expressed as mean ± SD (n = 3). **p < 0.01, siControl versus siRDM1. (E,F) Flow cytometry analysis of A549 (E–G) and PC9 (F-H) cell apoptosis after overexpression of RDM1. Similar methods were conducted as these descriptions in (A–D).
RDM1 regulates P53-RAD52-RAD51 in human lung adenocarcinoma cells
TP53 is the top mutated gene in human lung cancer, including lung adenocarcinoma16,17,18. And TP53 is a central tumor suppressor and has diverse roles in DNA-repair system19. Given that the critical role of RDM1 in DNA damage pathway, we hypothesize that RDM1 might regulate P53 expression in A549 and PC9 cells, which are human lung adenocarcinoma cells expressing wildtype TP5320. Western blot assay showed that knockdown of RDM1 in A549 and PC9 cells significantly increased the protein level of P53 (Fig. 5A). This results suggest the oncogenic role of RDM1 in human lung adenocarcinoma cells is partially by the negative regulation of P53. Furthermore, the analyses in String datasets indicated that RDM1 could interact with the typical DNA repair factors, such as RAD52 (Fig. 5B). It has been reported that Rad52 physically interacts with the Rad51 recombinase and serves as a mediator in the Rad51-catalyzed DNA strand exchange reaction21. To investigate the RDM1’s effect on these typical DNA repair factors, we knocked down or overexpressed RDM1 in A549 cells using siRNA. We found that knockdown of RDM1 decreased the protein expression of RAD51 and RAD52 with the increased protein expression of P53 (Fig. 5C–E). In sum, RDM1 regulated P53-RAD52-RAD51 in human lung adenocarcinoma cells.
RDM1 regulates P53-RAD51-RAD52 in human lung adenocarcinoma cells. (A) Western blots of P53 and RDM1 in A549 siRDM1 cells and PC9 siRDM1 cells. (B) The protein interaction analysis in String datasets. (C) Western blots of RAD51, RAD52, P53 and RDM1 in A549 siRDM1 cells. (D) Western blots of RAD51, RAD52, P53 and RDM1 in A549 cells with the overexpression of RDM1.
P53 negatively regulates the expression of RAD52 and RAD51 in human lung adenocarcinoma cells
It was reported that P53 inhibit the expression of Rad51 at transcriptional level by its specific binding to DNA22. Consistent with previous reports, our analyses of TP53 ChiP-Seq databases (GSM2746540, GSM1294879 and GSM2501568) show that TP53 has binding sites on RAD51 or RAD52 promoter region (Sup. Figs 1–2). Therefore, we next examined the effect of P53 on RAD51 and RAD52. The mRNA and protein expression of RAD51 and RAD52 were decreased after overexpression of P53 (Fig. 6). Taken together, these results suggested that RDM1 potentiates RAD52-RAD51 signaling via P53-mediated transcriptional suppression.
RDM1 regulates the mRNA expression and protein stability of P53 in human lung adenocarcinoma cells
Given that the dysregulation of P53 may be partially responsible for the oncogenic mechanism of RDM1 in lung adenocarcinoma cells, we next examined the potential regulation P53 by RDM1. First, we employed shRNA system to confirmed the regulation of RDM1 on P53-RAD52-RAD51 to exclude the potential off-target of siRNA. Consistent with the siRNA results, knockdown of RDM1 increased the mRNA and protein expression of P53-RAD52-RAD51 (Fig. 7A,B). Meanwhile, the expression of P53 protein was more stable after knockdown of RDM1 (Fig. 7C). These results suggested P53 could be regulated by RDM1 at the transcriptional level and through protein modification.
RDM1 regulates mRNA expression and protein stability of P53 in human lung adenocarcinoma cells. (A,B) Western blots (A) and qRT-PCR (B) analyses of RAD51, RAD52, P53 and RDM1 in A549 cells after knockdown of RDM1. (C) Western blots analysis of RDM1 and P53 in A549 cells after knockdown of RDM1 with Cycloheximide (CHX) treatment.
Discussion
In this study, we found that RDM1 played an oncogenic role in human lung adenocarcinoma cells. We observed that the mRNA and protein expression levels of RDM1 was up-regulated in human lung adenocarcinoma and correlated with poor clinical outcomes. RDM1 played an oncogenic role in human lung adenocarcinoma cells, which may be partially by inhibition of P53. Therefore, our study provides a new potential target for the treatment of lung cancer.
It has been proposed that RDM1 and RAD52 share similar functions in DNA double strand break repair and homologous recombination. Yet its function in cancer-related pathways is seldom explored. A recent work was shown that high RDM1 level was found in papillary thyroid carcinoma11. But the role of RDM1 in human lung adenocarcinoma remains unknown. In this study, we identified RDM1 as an oncogenic target in lung adenocarcinoma. First of all, analyses of human lung cancer samples had shown that RDM1 was over-expressed in lung adenocarcinoma tumors, initializing the hypothesis that RDM1 may impose an oncogenic function in lung cancer. Second, knockdown of RDM1 could reduce cell proliferation of lung adenocarcinoma cells, which might be ascribed to the increased apoptosis. Similar role of RDM1 was confirmed in the overexpression experiments. These results served as direct evidences that RDM1 might benefit for cancer cell survival. Consistent with the in vitro results, the in vivo growth of the RDM1-defective cells was significantly inhibited. These findings provided an important link between the chemotherapeutic resistance and the oncogenic functionality in lung adenocarcinoma. As a previous study indicated that RDM1 might play a secondary role in double-strand DNA break (DSB) repair and homologous recombination, despite that the RDM1−/− cells exhibited elevated sensitivity to cisplatin4, its function as an oncogenic protein in lung adenocarcinoma became more prominent. Our discovery also restated the notion that dysregulation of the DNA damage response (DDR) led to a predisposition to cancer23. Since it is conceivable that disruption of the DDR signaling may affect the response to DNA-damaging anticancer therapy (such as cisplatin-based ones), future work should be focused on testing whether and how DDR pathways will be altered upon treatment of cisplatin in the context of manipulating RDM1 functionality in lung adenocarcinoma cells.
Another interesting finding of our study was that RDM1 might regulate the expression of P53, as RDM1-knockdown cells showed the significant upregulation of P53 protein. TP53 not only is top mutated gene in human lung adenocarcinoma24, but also plays a central role in regulating cell growth and apoptosis25. P53 regulates various downstream targets, and triggers growth arrest or apoptosis19. Of the DNA damage that initiates a P53 response, the molecular mechanisms by which P53 is activated following DNA double strand breaks are the most comprehensively understood19. In our study, the finding that RDM1 regulates P53 expression can partially explain these phenotypes observed in RDM1 silencing cells. However, whether regulation of P53 by RDM1 is the only major mechanism contributing to the decreased cell growth in RDM1 knockdown cells needs further study. Meanwhile, how P53 regulated by RDM1 remains a question. TP53 can be regulated at different levels, including the transcriptional, epigenetic or post-translational levels18,26. Our data (Fig. 7) suggest that the inside mechanism related to transcriptional and post-translational levels. Therefore, following studies can examine the potential mechanisms from these two aspects.
Finally, as our Oncomine analyses have shown that up-regulation of RDM1 is generally correlated with poor clinical outcomes, we propose that RDM1 can be a potential prognostic marker for lung adenocarcinoma. Specifically, high RDM1 levels are significantly associated with advanced lung adenocarcinoma tumors, tumors from smokers, and patients of poor outcomes (i.e. short survivals and increased recurrence). As it has been proposed that combinatorial marker panel will out-perform any single marker for prognostic predictions27,28, we propose that RDM1 can serve as an additional marker to the marker panel for the diagnosis and/or stratification for clinical management of lung adenocarcinoma.
In summary, our study reveals the oncogenic function of RDM1 in lung adenocarcinoma. In light of the importance of chemotherapeutic resistance and DDR pathways, future work should address the following key questions: (1) whether and how does RDM1 in DNA damage repair is related to its oncogenic functions in lung adenocarcinoma; (2) how does RDM1 regulate P53; and (3) can RDM1 be targeted for alleviating cisplatin-resistance in the therapy of lung adenocarcinoma? Answers to these questions may offer new opportunities for diagnosing, treating, and managing this major subtype of NSCLC.
Materials
Animals
Six-week old mice were purchased from Shanghai SLAC Laboratory Animal CO. LTD. Mice were bred in the Animal Core Facility by following procedures ap-proved by the East China Normal University of Institutional Animal Care and Use Committee. Rules of animal welfare were applied throughout the experiment.
Cell culture
Human non-small cell lung adenocarcinoma cell lines PC9 and A549 were obtained from ATCC. Cells were cultured in RPMI 1640 medium supplemented with 2 mM L-glutamine, 10% fetal bovine serum (FBS), 100 U/ml penicillin, and 100 mg/ml streptomycin. Cell cultures were maintained in 37 °C and humidified atmosphere consisting 5% CO2.
Clinical tumor samples
Lung adenocarcinoma tumors were collected from Shanghai Cancer Center from October 2016 to July 2017. The patients were selected according to the following criteria: (1) all patients were diagnosed and confirmed by pathology; (2) patients with NSCLC were at early stages (Stage I and II) according the clinical staging method and had no other cancers; and (3) No preoperative chemotherapy or radiotherapy was administered to the cancer patients included in this study.
All samples were collected in accordance with ethical guidelines, and written informed consent was received. All patients were approached based on approved ethical guidelines, and patients who agreed to participate in this study were required to sign consent forms before being included in the study. All experimental protocols and methods were approved by Medical ethical committee of Shanghai Cancer Center (No. 2017-01-025). We also confirmed that all methods were performed in accordance with the relevant guidelines and regulations.
RNA interference of RDM1
RDM1 siRNA sequences is 5′-GCACCAGACAUAAGGCAGUTT-3′ and RDM1 shRNA sequence is 5′-GCGAAUUACUACUUUGGUUTT-3′. The specificity of RDM1 siRNA or shRNA sequences was confirmed by BLAST search against the human genome database. siControl-RNA was purchased from the GenePharma company.
Overexpression of RDM1
pcDNA3.1 vector was used to express human RDM1 cDNA. Lipofectamine 2000 (Thermo Fisher Scientific) was employed to transfect 1ug pcDNA3.1-RDM1 or pcDNA3.1-vector into A549 and PC9 cells.
Quantitative real-time PCR
Quantitative Real-time PCR using SYBR green reagents was performed as previously described29. The primer sequence for these detected genes: RDM1 forward: 5′-GCCCATCCTGGTTTCTATGCCC-3′; RDM1 reverse: 5′-AGACGAACCTTGACTGGAGAT-3′; RAD51forward: 5′- CAGTGATGTCCTGGATAATGTAGC-3′; RAD51 reverse: 5′-TTACCACTGCTACACCAAACTCAT-3′; RAD52 forward: 5′- TCAAGTACCGCGTGAAACCA-3′; RAD52 reverse: 5′- CGATCTTTGTTGCGGAACGG-3′; P53 forward: 5′- GGCAGACTTTTCGCCACAG-3′; P53 reverse: 5′- CAGGCACAAACACGAACCTC-3′; β-actin forward: 5′- CGTCATACTCCTGCTTGCTG-3′; β-actin reverse: 5′- GTACGCCAACACAGTGCTG-3′.
Proliferation assay
Cell proliferation assay was performed based on a colorimetric assay system (Cell Counting Kit-8, Dojindo Molecular Technologies, Inc., USA). Both targeted-knockdown (siRDM1) and control cells were seeded at a density of 1 × 105 cells/well before assay. At each time point, the absorbance at 450 nm was measured according to manufacturer’s instructions.
Analysis of apoptosis
Double-staining of Annexin V-FITC-Propidium iodide (PI) was used for analyzing apoptosis. Briefly, following collection and wash, cells were resuspended by cold PBS at a density of 1 × 106 cells/mL. PI (final concentration 100 µg/mL) and Annexin V-Alexa Fluor488 conjugate were mixed and added into the cell suspensions, followed by incubation at room temperature for 15 min. Flow cytometry was used to analyze apoptosis with the parameters of 494/518 nm set for Annexin V channel and 535/617 nm for PI.
Mouse xenograft tumor model
ShRDM1 and control cells were suspended in cold PBS (5 × 105 cells/100 µL) and injected into the mice subcutaneously on both flanks. Mice were continuously monitored for 6 weeks so the tumor size and weight were recorded. End-point mice were euthanized by CO2 asphyxiation, followed by removal of tumors. IHC was performed after the samples were embedded in paraffin, sectioned, and stained with hematoxylin and eosin (H&E) or IHC antibodies.
Western blot
For Western blot, proteins were separated on 12% SDS-polyacrylamide gels and transferred to polyvinylidene difluoride (PVDF) membranes. Membranes were blocked in 5% non-fat milk in Tris-buffered saline (plus 0.1% Tween-20). RDM1 (proteintech, 20156-1-AP), p53 (santa cruz, sc-6243), RAD51 (abcam, ab133534), RAD52 (abcam, ab124971) and β-actin (MBL, M177) antibodies were purchased. Blots were detected by chemiluminescence and exposed on X-ray films.
Immunohistochemistry (IHC)
IHC was performed for continuous sections from paraffin-embedded blocks. Antigen retrieval was performed by microwaving for 3 min in citrate-buffered solution (pH 6.0). Blocking was done by incubation with 10% goat serum at room temperature for 30 min. Sections were incubated with RDM1 (proteintech, 20156-1-AP) antibody that are indicated in this study for overnight at 4 °C. Staining with secondary antibody (conjugated with horseradish peroxidase) was performed for 1 hr at room temperature. Sections were finally stained with 3, 3- diaminobenzidine tetrahydrochloride (DAB) and counterstained with hematoxylin. Normal rabbit serum was used as negative control in place of the primary antibody. The IHC results are quantified using ImageJ (fiji-win64) as previously described29.
Statistical analysis
Student’s t tests were performed to obtain the statistical significance. A P value < 0.05 was considered significant.
References
van den Bosch, M., Lohman, P. H. & Pastink, A. DNA double-strand break repair by homologous recombination. Biol Chem 383, 873–892, https://doi.org/10.1515/BC.2002.095 (2002).
West, S. C. Molecular views of recombination proteins and their control. Nat Rev Mol Cell Biol 4, 435–445, https://doi.org/10.1038/nrm1127nrm1127 (2003).
Messaoudi, L. et al. Subcellular distribution of human RDM1 protein isoforms and their nucleolar accumulation in response to heat shock and proteotoxic stress. Nucleic Acids Res 35, 6571-6587, gkm753 [pii] https://doi.org/10.1093/nar/gkm753 (2007).
Hamimes, S. et al. RDM1, a novel RNA recognition motif (RRM)-containing protein involved in the cell response to cisplatin in vertebrates. J Biol Chem 280, 9225–9235, https://doi.org/10.1074/jbc.M412874200 (2005).
Liu, J. et al. ErbB2 Pathway Activation upon Smad4 Loss Promotes Lung Tumor Growth and Metastasis. Cell Rep, https://doi.org/10.1016/j.celrep.2015.02.014 (2015).
Jemal, A. et al. Cancer statistics, 2006. CA Cancer J Clin 56, 106–130, 56/2/106 (2006).
Spira, A. & Ettinger, D. S. Multidisciplinary management of lung cancer. N Engl J Med 350, 379–392, https://doi.org/10.1056/NEJMra035536350/4/379 (2004).
Bronte, G. et al. Driver mutations and differential sensitivity to targeted therapies: a new approach to the treatment of lung adenocarcinoma. Cancer Treat Rev 36(Suppl 3), S21–29, https://doi.org/10.1016/S0305-7372(10)70016-5S0305-7372(10)70016-5 (2010).
Shaw, A. T. & Engelman, J. A. Ceritinib in ALK-rearranged non-small-cell lung cancer. N Engl J Med 370, 2537–2539, https://doi.org/10.1056/NEJMc1404894 (2014).
Vansteenkiste, J. F. Ceritinib for treatment of ALK-rearranged advanced non-small-cell lung cancer. Future Oncol 10, 1925–1939, https://doi.org/10.2217/fon.14.94 (2014).
Li, W., Huang, Q., Sun, D., Zhang, G. & Tan, J. RDM1 gene overexpression represents a therapeutic target in papillary thyroid carcinoma. Endocr Connect 6, 700–707, https://doi.org/10.1530/EC-17-0209EC-17-0209 (2017).
Okayama, H. et al. Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res 72, 100–111, https://doi.org/10.1158/0008-5472.CAN-11-14030008-5472.CAN-11-1403 (2012).
Hou, J. et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS One 5, e10312, https://doi.org/10.1371/journal.pone.0010312 (2010).
Ding, L. et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455, 1069–1075, https://doi.org/10.1038/nature07423nature07423 (2008).
Bild, A. H. et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439, 353–357, https://doi.org/10.1038/nature04296 (2006).
Cancer Genome Atlas Research, N. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550, https://doi.org/10.1038/nature13385 (2014).
Cancer Genome Atlas Research, N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525, https://doi.org/10.1038/nature11404 (2012).
Liu, J. et al. REGgamma modulates p53 activity by regulating its cellular localization. J Cell Sci 123, 4076–4084, https://doi.org/10.1242/jcs.067405 (2010).
Meek, D. W. The p53 response to DNA damage. DNA Repair (Amst) 3, 1049–1056, https://doi.org/10.1016/j.dnarep.2004.03.027S1568786404000862 (2004).
Ummanni, R. et al. Evaluation of reverse phase protein array (RPPA)-based pathway-activation profiling in 84 non-small cell lung cancer (NSCLC) cell lines as platform for cancer proteomics and biomarker discovery. Biochim Biophys Acta 1844, 950–959, https://doi.org/10.1016/j.bbapap.2013.11.017 (2014).
Krejci, L. et al. Interaction with Rad51 is indispensable for recombination mediator function of Rad52. J Biol Chem 277, 40132–40141, https://doi.org/10.1074/jbc.M206511200M206511200 (2002).
Lazaro-Trueba, I., Arias, C. & Silva, A. Double bolt regulation of Rad51 byp53: a role for transcriptional repression. Cell Cycle 5, 1062–1065, https://doi.org/10.4161/cc.5.10.2764 (2006).
Curtin, N. J. DNA repair dysregulation from cancer driver to therapeutic target. Nat Rev Cancer 12, 801–817, https://doi.org/10.1038/nrc3399nrc3399 (2012).
Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550, https://doi.org/10.1038/nature13385nature13385 (2014).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310, https://doi.org/10.1038/35042675 (2000).
Saldana-Meyer, R. & Recillas-Targa, F. Transcriptional and epigenetic regulation of the p53 tumor suppressor gene. Epigenetics 6, 1068–1077, https://doi.org/10.4161/epi.6.9.16683 (2011).
Groeger, A. M. et al. Expression of p21 in non small cell lung cancer relationship with PCNA. Anticancer Res 20, 3301–3305 (2000).
Coate, L. E., John, T., Tsao, M. S. & Shepherd, F. A. Molecular predictive and prognostic markers in non-small-cell lung cancer. Lancet Oncol 10, 1001–1010, https://doi.org/10.1016/S1470-2045(09)70155-XS1470-2045(09)70155-X (2009).
Liu, J. et al. Mig-6 deficiency cooperates with oncogenic Kras to promote mouse lung tumorigenesis. Lung Cancer 112, 47–56, https://doi.org/10.1016/j.lungcan.2017.08.001 (2017).
Acknowledgements
This work was supported by the National Basic Research Program of China (2016YFC0902102, 2015CB910402). This study was also funded by the National Basic Research Program (2011CB504200, 2015CB910403). We thank Dr. Xiaotao Li for help in this study. This work was also supported in part by grants from National Natural Science Foundation of China (81401837, 81471066, 81261120555, 31200878, 31071875, 81271742, 31401012, 31730017), the Science and Technology Commission of Shanghai Municipality (14430712100), Shanghai Rising-Star Program (16QA1401500) and Shanghai natural science foundation (17ZR1407900, 12ZR1409300, 14ZR1411400).
Author information
Authors and Affiliations
Contributions
L.T., W.j.C., S.h.S. and K.L. conducted the experiments and provided experimental methods and data. J.L. provided Oncomine and Chip-seq analyses. G.p.N., W.j.Y. J.L. and L.L. designed the study, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tong, L., Liu, J., Yan, W. et al. RDM1 plays an oncogenic role in human lung adenocarcinoma cells. Sci Rep 8, 11525 (2018). https://doi.org/10.1038/s41598-018-30071-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30071-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.