Abstract
MicroRNAs (miRNAs) are endogenously short noncoding regulatory RNAs implicated in plant development and physiology. Nine small RNA (sRNA) libraries from three typical seed developmental stages (young, intermediate, and mature) were generated by deep sequencing to identify the miRNAs of J. curcas, a potential oilseed crop for the production of renewable oil. Strict criteria were adopted to identify 93 high confidence miRNAs including 48 conserved miRNAs and 45 novel miRNAs. Target genes of these miRNAs were involved in a broad range of physiological functions, including gene expression regulation, primary & secondary metabolism, growth & development, signal transduction, and stress response. About one third (29 out of 93) miRNAs showed significant changes in expression levels during the seed developmental process, indicating that the miRNAs might regulate its targets by their changes of transcription levels in seed development. However, most miRNAs were found differentially expressed in the late stage of seed development, suggesting that miRNAs play more important roles in the stage when seed accumulating organic matters and suffering dehydration stress. This study presents the first large scale identification of high confidence miRNAs in the developing seeds of J. curcas.
Similar content being viewed by others
Introduction
Huge consumption of fossil fuel leading to higher and higher levels of carbon dioxide released into the atmosphere is a key concern in the world. Biodiesel, as an alternative for fossil fuel, is an environmentally friendly biofuel. Jatropha curcas, a perennial, drought-resistant small tree belonging to the family of Euphorbiaceae, can be grown on non-agricultural land without sacrificing agricultural land. Its seed has a high content of oil that can be reformed as biodiesel for diesel engines1,2,3. However, improving the seed yield, the oil content, and the oil quality remains challenging, and these problems prevents fulfilling the promise of this energy crop4,5,6. Understanding the molecular processes of seed development and oil metabolism is crucial to solve these problems. Much efforts have therefore been made to analyze the gene expression in all kinds of tissues in Jatropha, especially in the developing seeds7,8,9,10. The genomic sequences and protein-encoding genes obtained through genome analysis have laid the foundation for further exploration of the genetics and genomics of J. curcas11,12,13.
MicroRNAs (miRNAs) are endogenous small non-coding RNAs that act as post-transcriptional regulators of gene expression in animals and plants by targeting mRNAs for cleavage or translational repression14,15,16. In plants, miRNAs are encoded in intergenic regions, where they are typically transcribed by RNA Polymerase II as long polyadenylated transcripts called pri-miRNAs. Then pri-miRNAs are recognized and processed by DICER-Like1 (DCL1) into miRNA precursors (pre-miRNAs), which are further processed to generate 18~24 nucleotide (nt) mature miRNAs16,17,18. The mature miRNAs bind to target mRNAs for cleavage or translational inhibition, depending on the degree of complementarity between the miRNAs and its target mRNAs15,18. In recent years, researchers found that miRNAs play critical roles in plant development and growth as well as a range of physiological processes, including abiotic and biotic stress responses and probably lipid metabolism19,20,21.
To our knowledge, only several recent studies focused on the identification of miRNAs from J. curcas. Wang et al. cloned 52 putative miRNAs from leaves and seeds without providing information on the miRNA precursor sequence22. Vishwakarma and Jadeja used known plant miRNAs to search for J. curcas miRNAs from EST and GSS sequences, and only 24 putative conserved miRNAs were identified23. Galli et al. investigated miRNAs through small RNAs (sRNA) deep sequencing from mature seeds, and revealed 180 conserved miRNAs and 41 precursors as well as 16 novel pre-miRNAs, but miRNAs involved in other developmental stages are yet to be found24. Furthermore, quite a few miRNAs identified in these reports could not be qualified as high confidence miRNAs according to the current strict criteria25,26.
In this study, high confidence miRNAs from J. curcas were identified through the deep sequencing of sRNA from three typical developing stages of seeds (young, intermediate and mature). The prediction of miRNA targets indicates that these miRNAs are involved in a wide range of physiological processes. In addition, differentially expressed miRNAs were found mainly in the late stages, providing important information on the possible function of miRNAs in seed development.
Results and Discussion
Deep Sequencing of sRNAs
To identify miRNAs and their expression patterns in the developing seeds of J. curcas, nine sRNA libraries from three typical developmental stages of seeds (young, intermediate and mature seeds; Supplementary Fig. S1) with three biological replicates were generated and sequenced by next-generation sequencing (NGS), resulting in 16.7~21.6 million raw reads for each sample and 162.7 million raw reads in total, which was a deep resource for extensive discovery of miRNAs. After trimming the adaptors, filtering out the low quality reads and noise, we obtained 13.8~18.9 million clean reads for each of the nine sRNA libraries, and no less than 95% of the reads had a high Detection Accuracy (>99.9%) (Table 1).
The clean reads were mapped to the SILVA rRNA, Rfam, tRNAdb, and Repbase databases to annotate the composition of the sRNA population. It showed that rRNA had the highest read frequency (19.65~41.56%) followed by tRNA (1.46~3.52%) in the total clean reads of every library. Repetitive elements, scRNA, snRNAs and snoRNA were less frequent in the sRNA library. Taken together, 55.24~77.1% of the clean reads were unannotated reads for the nine sRNA libraries, and these unannotated reads could be used for miRNA detection (Table 2).
Among the clean reads, the most abundant sequences were 21 nt (8.0%), 22 nt (6.1%), 23 nt (10.5%), and 24 nt (45.4%) (Supplementary Table S1). This distribution pattern of the sRNA size was similar to those seeds from other species, such as the developing seeds of canola27,28 and safflower29, and the mature seeds of J. curcas24. In these reports, the most abundant sRNA types are 21~24-nt long and 24-nt sRNA is also the most abundant one (>40%), indicating that 21~24-nt long sRNA might be the main sRNA products, and these plants might have similar sRNA biogenesis patterns in seeds.
Identification of High Confidence miRNAs
To analyze the unannotated reads for discovery of miRNAs, the miRDeep2 core algorithm with modifications for plant miRNAs was employed30,31. In brief, the unannotated reads were mapped onto the genome of J. curcas, and the flanking sequences of the mapped sites were extracted as candidate miRNA precursors. The mfe (minimal free energy) of the precursor structure, read counts of mature and star miRNA sequences, randfold p-values, and minimum 60% nucleotides paired in the candidate mature and star miRNAs, were considered and scored by the miRDeep2 core algorithms. In total, 108 potential miRNAs were identified in the developing seeds of J. curcas.
The stem-loop structures and other characters of miRNA precursors were displayed in miRDeep2 outputs (Supplementary Fig. S2 and Supplementary Table S2). These data were further manually examined for mismatches between the miRNA and star miRNA, the number of asymmetric bulges in the stem region, the size of asymmetric bulges, 3′ overhangs in duplex, dominance of the miRNA relative to other sRNAs in abundance, and precise miRNA/star miRNA excision, to annotate high confidence microRNAs according to the rules detailed by Meyers et al.25,32. Four rules must be followed if miRNAs could be accepted as high confidence ones: (1) The miRNA and star miRNA form a duplex with two nt 3′ overhangs (if star miRNA is absent, candidate miRNA must be found in multiple, independent libraries); (2) four or fewer mismatches between the miRNA and the other arm of the hairpin; (3) not more than one asymmetric bulge and less than two bases in the bulge, especially within the duplex; and (4) stem-loops that slightly violate one of these criteria could still be annotated as miRNAs, if precise miRNA/star miRNA excision is shown, i.e., more than 75% sRNA localized in one arm of miRNA/star miRNA duplex25. According to these rules, 93 high confidence miRNAs were obtained from the developing seeds (Supplementary Fig. S2 and Supplementary Table S2). That is to say, about 86% (93/108) miRNAs were qualified as high confidence miRNAs, confirming the efficiency and accuracy of the modified miRDeep2 in plant miRNA identification30,31. Most of the unqualified miRNAs were ruled out due to the imprecise miRNA/star miRNA excision or improper 3′ overhangs. If more stringent filtering strategies were adopted, three more rules should be added according to Kozomara and Griffiths-Jones26: (5) At least 10 reads must map with no mismatches to each of the two possible mature microRNAs derived from the hairpin precursor; (6) at least 50% of reads mapping to each arm of the hairpin precursor must have the same 5′ end; and (7) the predicted hairpin structure must have a folding free energy of <−0.2 kcal/mol/nt. Finally, we got 45 very high confidence miRNAs satisfying all the seven rules mention above (Supplementary Table S2). Almost all the 48 miRNAs failed to be qualified for very high confidence ones due to failure to satisfy the fifth rule: less than 10 reads of star miRNAs mapped with no mismatches. Considering the fact that star miRNAs are expressed at low level or even hardly detected for some miRNAs, some miRNAs could be approved as high confidence ones if they meet all the ancillary criteria25. For example, in the absence of star miRNA confirmation, a clear dominance of the miRNA sequences from one arm of a predicted stem-loop was found (Supplementary Fig. S2). Furthermore, the annotation was well supported by miRNA identification from multiple, independent libraries. Most of the 93 miRNAs were found in 9 independent libraries, except that Jcr4S02434_14511 and Jcr4S00022_291 could not be detected in young seeds. All the miRNAs identified in this work had high read counts. For instance, Jcr4S00096_1114 had the smallest (285) and Jcr4S01201_8767 had the highest read counts (741967). These features could be considered as ancillary criteria that strongly bolster miRNA annotations as suggested by Meyers et al.25 When we checked the structures of J. curcas miRNAs published in previous reports by Wang et al.22 and Galli et al.24, less than half of the miRNAs could satisfy Meyer’s rules (2) and (3) mentioned above, let alone other rules. Therefore, the miRNA screening criteria adopted by this work are stringent enough to qualify the 93 miRNAs as high confidence miRNAs.
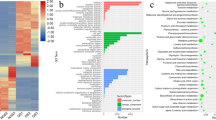
The lengths of the 93 mature miRNAs varied from 19 nt to 24 nt. More than half (50/93) of miRNAs were 21 nt, followed by 24-nt miRNAs (28) (Fig. 1a). Accordingly, 21-nt and 24-nt sRNAs were the most abundant ones (Fig. 1b). Plants have at least four Dicer-like proteins (DCL1-4), among which DCL1 mainly produces 18~21-nt long sRNA while DCL2, DCL3, and DCL4 produce 22-, 24-, and 21-nt long sRNA, respectively, and 21, 22, and 24 nt are typical sizes for plant Dicer-like (DCL)-derived products16. Although 24-nt sRNAs accounted for almost half (45.4%) of the total sRNAs and 21-nt sRNAs took up much less (8.0%) (Supplementary Table S1), it is interesting to note that more than half miRNAs (50/93) were 21 nt in length, which is quite accordant to the fact that DCL1 is responsible for most plant miRNA biogenesis and plant miRNAs are predominately 21 nt in length18. However, both the 24-nt miRNAs and 24-nt sRNAs were the second large population in total clean reads, indicating DCL3 might play a part in the sRNA production in the developing seeds. We also determined the frequency of the first base of mature miRNAs, and it showed that the 20-, 21-, and 22-nt miRNAs preferentially started with ‘U’ (7/7, 34/50, and 6/6, respectively), while 24-nt miRNAs preferred ‘C’ at the first base (9/24) (Fig. 1c). Similar base bias trend for 20-, 21-, and 22-nt miRNAs is also found in other plant species, such as Camellia33 and canola28. This is well in accordance to that plant miRNAs prefer to begin with a ‘U’ residue34. However, the first base of 24-nt miRNAs prefer ‘A’ in Camellia33 and canola28. The first base bias may determine their assembling into different AGO complexes and might result in a different regulatory outcome in the developing seeds of J. curcas16.
Identification of Conserved and Novel miRNAs
High confidence miRNAs were then aligned to known miRNAs from miRBase. A total of 48 miRNAs shared homology with known miRNAs were identified as conserved miRNAs, and 45 miRNAs without homology, novel miRNAs (Supplementary Table S2). The 48 conserved miRNAs belonging to 23 families (Jcu-MIR156, 159, 160, 162, 164, 166, 167, 168, 171, 172, 319, 390, 393, 394, 395, 396, 398, 403, 408, 477, 827, 1446, and 6445) could be found in at least one plant species except for J. curcas, and most (39/48) were identified in more than 10 species (Supplementary Table S2). Jcu-MIR156 and MIR166 were the biggest families with 6 members each, followed by Jcu-MIR319 (5 members). Four families (Jcu-MIR156, 159, 390, and 6445 (named nMIR001-5p in Galli’s report)) were also detected in the mature seeds of J. curcas in a previous report24. Jcu-MIR166 and Jcu-MIR167 families were both discovered in mature seeds based on deep sequencing24 and bioinformatic prediction based on expressed sequence tags23. Jcu-MIR167 (JcumiR027 in Wang’s work) was also identified in the leaf of J. curcas by cDNA cloning method, but not in the seed tissues22. The 45 novel miRNAs belonged to 34 families (named Jcu-nMIR001~Jcu-nMIR034). In previous reports, Jcu-nMIR020 (nMIR002-5p in Galli’s work) was only observed in the mature seeds24 and Jcu-nMIR005 (JcumiR019-3p in Wang’s work) was identified in the leaves instead of seed tissues of Jatropha22. However, Jcu-MIR167 and Jcu-nMIR005 were detected in all the three developing seed stages with extremely high read counts (>400,000) in this work, but neither were detected in the seeds of Jatropha by cDNA cloning method22, implying that even high abundance of miRNAs could not guarantee their detectability by the cloning method, because some miRNAs could be missed by chance when cloned into the pGEMT vector and sequenced22. To our knowledge, a total of 17 conserved (except Jcu-MIR156, 159, 166, 167, 390, and 6445) and 32 novel (except Jcu-nMIR020 and Jcu-nMIR005) miRNAs families were identified in J. curcas for the first time.
Prediction of miRNA Targets
In plants, as a post-transcriptional regulator, the mature miRNA binds to target mRNAs by their complementarity for cleavage or translational repression, and therefore miRNA target prediction is crucial for understanding the functions of miRNAs14,15,16,18. Based on the complementarity, target mRNAs of J. curcas miRNAs were predicted by using the web-based psRNATarget35 and Targetfinder software36.
A total of 331 potential target genes were found for 73 miRNAs with an average of 4.5 targets per miRNA (Supplementary Table S2). Some miRNAs had a larger number of targets, such as Jcu-MIR156 and Jcu-MIR396 (>12 targets for each family members), which were also observed to have the most targets in cassava37; The target genes of 22 conserved and 24 novel miRNA families were obtained. In other words, targets were spotted for almost all conserved miRNAs (46/48) and conserved miRNA families (22/23), whereas only for about half of novel miRNAs (27/45) and novel miRNA families (24/34), and a similar trend was observed in the targets determination in a previous report24. These results are in line with the idea that novel plant miRNAs, in contrast to deeply conserved miRNAs, are more divergent and tend to lack targets38.
A total of 188 unique target sequences were obtained after removing the redundancy in the 331 potential targets for the 73 miRNAs, and the functions of the 188 unique targets were annotated according to previous reports11,12 or by using BLASTX against NCBI nr and Swiss-Prot/Uniprot protein databases (Supplementary Table S3), followed by a GO analysis to evaluate their putative functions in regards to the cellular component, the molecular function and the biological process (Supplementary Fig. S3). The majority of miRNA targets are localized in nucleus or membrane, and some are in cytoplasm, ribosome, chloroplast, mitochondrion, cell wall, or even extracellular region; most miRNA targets participate in DNA binding, ATP binding, or had transcription factor activity; the miRNA targets mainly participate in DNA-templated regulation of transcription, oxidation reduction, developmental process, metabolic processes, protein phosphorylation or ubiquitination, electron transport, and signaling pathways (Supplementary Fig. S3). Most targets are involved in transcription regulation, and have a broad range of physiological functions. The function categories of miRNA targets in GO analysis are overlapped and complicated, and we therefore categorized the targets roughly into five aspects of plant physiological process according to the annotation in NCBI nr, Uniprot data, and previous reports: gene expression regulation, primary & secondary metabolism, growth & development, signal transduction, and stress response. To be concise, the miRNAs identified in this work are parenthesized behind their targets in the following text.
The first aspect comprises genes involved in all levels of gene expression regulation: DNA replication, transcription, post-transcriptional regulation, translational process and posttranslational modification. Non-structural maintenance of chromosomes element 4 (Jcu-MIR6445) is involved in DNA replication, recombination and repair. Most targets are involved in gene transcription process and some are well known families of transcription factors, such as DELLA protein, leucine zipper protein, AP2/ERF and B3 domain-containing protein, NAC domain-containing protein, WRKY, squamosa promoter-binding-like protein, MYB, GAMYB, etc (Supplementary Table S3). The targets are also involved in post-transcriptional regulation, such as endoribonuclease Dicer homolog 1 (Jcu-MIR162) and argonaute 2 (Jcu-MIR403), which contribute to the production of miRNA or siRNA. Pentatricopeptide repeat (PPR) proteins (Jcu-nMIR021) are a large family of modular RNA-binding proteins which mediate several aspects of gene expression primarily in organelles but also in nucleus. These proteins facilitate processing, splicing, editing, stability and translation of RNAs39, and thus it is classified as RNA processing and modification. Translation initiation factor eIF-2B subunit (Jcu-nMIR025) and eukaryotic peptide chain release factor GTP-binding subunit ERF3A (Jcu-MIR1446) are typical factors involved in translational process. Some target genes function at posttranslational modification level, in which protein turnover is regulated through ubiquitin-dependent pathway, such as RING-H2 finger protein ATL5 (Jcu-nMIR011), F-box only protein 6 (Jcu-MIR394), E3 ubiquitin-protein ligase (Jcu-MIR396), ubiquitin-conjugating enzyme (Jcu-nMIR021), and ubiquitin-like modifier-activating enzyme (Jcu-MIR172). The results indicate that miRNAs are involved indirectly in all levels of gene expression regulation, though miRNAs per se are post-transcriptional regulators.
The second aspect includes the genes involved in primary & secondary metabolism. Some target genes play roles in primary metabolism, i.e., protein, carbohydrate, and lipid metabolism. A sec. 1 family domain-containing protein MIP3 (Jcu-nMIR003) is required for proper maturation of seed storage proteins and it forms a complex with MAG2, ZW10/MIP1 and MIP2 on the endoplasmic reticulum which may be responsible for efficient transport of seed storage proteins. The 1, 4-alpha-glucan-branching enzyme 2–2 (Jcu-nMIR011) is involved in starch biosynthetic process. Eight miRNA targets are related to lipid metabolic pathways (Supplementary Table S3), among which three targets were involved in the lipid or fatty acid biosynthetic processes: cytochrome P450 (Jcu-MIR156f) catalyzes the omega-hydroxylation of various fatty acids, which is important for cutin biosynthesis, trichome differentiation, establishment of apical dominance and senescence40; GPI mannosyltransferase 1 (Jcu-MIR403) works in the pathway of cosylphosphatidylinositol-anchor biosynthesis, a part of glycolipid biosynthesis; and malonate-CoA ligase (Jcu-MIR403) catalyzes the formation of malonyl-CoA directly from malonate and CoA. Among the eight targets related to lipid metabolic pathways, five targets play a part in the lipid catabolic processes: monoacylglycerol lipase ABHD6 (Jcu-nMIR013), sn1-specific diacylglycerol lipase beta (Jcu-nMIR008), and phospholipase D zeta 1(Jcu-MIR827) are lipase; long-chain-alcohol oxidase FAO2 (Jcu-MIR1446) is involved in the omega-oxidation pathway of lipid degradation; acyl-coenzyme A thioesterase 9 (Jcu-nMIR011), catalyzes the hydrolysis of acyl-CoAs to the free fatty acids and CoAs, probably regulating intracellular levels of acyl-CoAs, free fatty acids and CoASH. Although increasing the oil content and improving the oil quality are still a challenging task5,41, the discovery of these miRNAs involved in lipid metabolism might provide an alternative way to solve the problems; for example, Jcu-nmiR011 might have a potential application in controlling the level of free fatty acids in the seed oil of J. curcas. Except for the primary metabolism, target genes involved in the secondary metabolism were also identified: deacetylvindoline O-acetyltransferase (Jcu-MIR156f) was shown to be involved in the biosynthesis of vindoline, a precursor of vinblastine and vincristine42; premnaspirodiene oxygenase (Jcu-MIR159) functions in the biosynthesis of solavetivone, a potent antifungal phytoalexinin43; chalcone synthase 2 (Jcu-nmiR014) participates in the biosynthesis of flavonoid, which is an important secondary metabolite in plants; and laccase-9 (Jcu-MIR398) is an important enzyme in lignin degradation and detoxification of lignin-derived products, which has become a focus recently for environmental protection. These results suggest that miRNAs play important roles in a wide range of plant primary and secondary metabolism and have promising application in agriculture, medicine and environment protection.
The third aspect is growth & development, including genes involved in cell division, cell differentiation and seed development. Growth-regulating factors (GRF1, 2, 4, 5, 7, 8 and 9) targeted by Jcu-MIR396 are transcription activators associated with the cell expansion of leaves and cotyledons. The evidences obtained from Arabidopsis, rice, and maize proved that miR396-GRF, a conserved regulatory network in plants, plays an important role in leaf growth and seed development, acting in the regulation of meristem function, at least partially through cell proliferation control44,45. Protein CUP-SHAPED COTYLEDON 2 (Jcu-MIR164) involved in meristem initiation and cell division patterns is also a target of Ath-MIR164 in Arabidopsis46. Coiled-coil domain-containing protein SCD2, AUGMIN subunit 5, and trafficking protein particle complex II-specific subunit targeted by Jcu-MIR156, 477, and nmiR001 respectively, take part in cell division and cell expansion47,48,49. Callose synthase 9 (Jcu-MIR319), cellulose synthase-like protein G3 (Jcu-MIR395), and leucine-rich repeat extensin-like protein 2 (Jcu-nMIR005) are involved in cell wall organization. Homeobox-leucine zipper protein ATHB-8, ATHB-14 (PHABULOSA), ATHB-15 (CORONA), and REVOLUTA belong to class III homeodomain-leucine zipper genes, which are transcription factors playing overlapping and different roles in the cell differentiation, vascular tissue formation, integument development, and embryogenesis50,51,52. Previous reports showed that ATHB15 is targeted by MIR166 in Arabidopsis53. Our results indicate that class III homeodomain-leucine zipper genes with related functions are all targeted by Jcu-MIR166, which might be an efficient strategy for the regulation of target genes coordinately. Transcription factor TCP family (2, 3 and 4), targeted by Jcu-MIR319, plays a pivotal role in cell division and leaf development and is required during early steps of embryogenesis54,55. As MIR319-TCP regulates leaf development in Arabidopsis55, MIR319-TCP pathway is probably conserved and might be related to seed development in J. curcas. Floral homoerotic protein APETALA 2 (Jcu-MIR172) activity is required in plant ovule development and seed development in addition to its known function in floral development56. It is worth to mention that miRNA-target network must be complicated in controlling plant development; for instance, MIR319 targets transcription factor TCP, negatively regulating the expression of CUP-SHAPED COTYLEDON through the induction of MIR16457.
The fourth aspect is signal transduction, including genes targeted by Jcu-MIR156, 159, 390, 393, 394, 396, 403, 827, nMIR010, nMIR014, nMIR021, nMIR024, and nMIR033. Some targets are under the control of plant hormones. Histidine kinase 3 (Jcu-nMIR033a) may participate in cytokinin-activated signaling pathway that regulates many developmental processes including seed germination, cell division, and seed size58. Auxin signaling F-box 2 (Jcu-MIR393) is involved in embryogenesis regulation by auxin, and it is shown to be targeted by MIR393 for embryogenesis in Arabidopsis59. Auxin response factor 16 (Jcu-MIR160), though classified as transcriptional factors, belongs to auxin-activated signaling pathway60. These genes may play a role in cell division and cell differentiation, and thus eventually in seed growth and formation under the control of hormones and miRNAs. Calmodulin is known to be involved in signal transduction by mediates the control of a large number of enzymes, ion channels and other proteins. Calmodulin-7 (Jcu-nMIR021) is transcription factor in Arabidopsis seedling development61. These results suggest that miRNAs should control a large range of plant activities since some signal transduction pathways are under the regulation of miRNAs.
Finally, it is no wonder that a lot of targets are involved in stress response, since seeds should be suffered desiccation and other stress during developmental process. Late embryogenesis abundant protein (Jcu-MIR394) is a group of well known proteins which may play an essential role in seed survival and in controlling water exchanges during seed desiccation. At least four targets of miR159 are related to defense and stress response, such as superoxide dismutase and peroxidase (Supplementary Table S3). AP2/ERF and B3 domain-containing transcription factor (Jcu-MIR156) may be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways according to Uniprot annotation. In Arabidopsis, MIR156, MIR159 and MIR394 are known as dehydration stress-responsive miRNAs. DEAD-box ATP-dependent RNA helicase 38 (Jcu-MIR393) is essential for mRNA export from the nucleus and plays an important role in the positive regulation of CBF/DREB transcription factors62,63, and J. curcas DREB gene expression is induced by cold, salt and drought stresses64.
Differentially Expressed miRNAs During Seed Development
Plant miRNAs play pivotal roles in the regulation of plant growth and development and therefore it is crucial to outline the spatiotemporal expression patterns of miRNAs. To gain insight into the possible developmental stage-dependent roles of miRNAs in Jatropha, the expression levels and patterns of miRNAs in seed were determined at three different developmental stages. The miRNA expression levels were shown as TPM values which were comparable between different samples36. About one third (29/93) were differentially expressed miRNAs (fold change >2, and p < 0.05) (Supplementary Table S4), which were presented as the base-2 logarithm of fold change of expression levels between young, intermediate, and mature stages (Fig. 2). In the 29 differentially expressed miRNAs, 11 miRNAs were down-regulated whereas 18 miRNAs were up-regulated, and the variation was continuous from young through mature stages without fluctuation (Fig. 2). Among these differentially expressed miRNAs, Jcu-MIR171a/b and Jcu-MIR390 were down-regulated most significantly. Jcu-MIR171b targets scarecrow-like protein 15 (XM_012223680.2), which is a transcription factor involved in plant development. MIR390 targets protein NRT1/ PTR FAMILY 4.6 (XM_012224813.2) which is involved in cellular abscisic acid (ABA) transport and uptake65. Jcu-nMIR001, nMIR002 and nMIR013 were very highly expressed in mature seeds, but hardly detected in young and intermediate stages (Supplementary Table S4). Chloroplastic photosynthetic NDH subunit of subcomplex B 3 (XM_012223240.2; target of Jcu-nMIR001) shuttles electrons in the photosynthetic electron transport in photosystem I to produce energy66. Another target of Jcu-nMIR001 is trafficking protein particle complex II-specific subunit 130 (XM_012215242.2), which is responsible for growing cell plate in mitotic active cells48. Jcu-nMIR002 targets auxin-responsive protein SAUR50-like (XM_020682576.1), which promotes cell expansion during plant growth67. Jcu-nMIR013 targets monoacylglycerol lipase ABHD6 (XM_012226208.2) which participates catabolic process of lipids, and the high level of Jcu-nMIR013 in mature seeds should repress the lipid catabolic process and so benefit the lipid accumulation. Jatropha 11 kD late embryogenesis abundant protein belongs to late embryogenesis abundant (LEA) group 1 (XM_012232372.2) involved in the acquisition of desiccation tolerance during late phase of embryogenesis68,69. It is targeted by Jcu-MIR394, which was down-regulated significantly during seed maturation, probably facilitating the accumulation of LEA protein at mature stage to improve desiccation tolerance. However, it is hard to find an unambiguous inverse corelation between the differentially expressed miRNAs and the target mRNAs levels (data not shown), because miRNAs might play a part role in the regulation of mRNA levels and some targets were regulated by miRNAs at translational level instead of cleavage. Degradome sequencing should be a better choice to solve this problem28.
The differentially expressed miRNAs during seed development of J. curcas. The miRNA expression levels were estimated for each sample by mapping sRNA back onto the miRNA precursor sequence, and read count for each miRNA was obtained from the mapping results. Differential expression analyses of miRNAs were performed based on Read counts using the DESeq. 2, in which p values were adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate (FDR). The variation of miRNA levels in three different developmental stages, i.e., young (Y), intermediate (I) and mature (M) seeds were shown as the base-2 logarithm of ratios (log2 fold change) between different developmental stages (I/Y, M/I and M/Y). The miRNAs whose expression levels between any two of the three different developmental stages changed significantly (|log2 fold change| > 1 and FDR adjusted p < 0.05) were assigned as differentially expressed miRNAs (indicated by *).
To our surprise, most differentially expressed miRNAs were observed during seed development from intermediate to mature stages, and only three miRNAs (Jcu-miR390 and Jcu-395a/b) were observed from young to intermediate. Reverse transcription (RT) quantitative real-time PCR (qPCR) was then performed for the validation of differentially expressed miRNAs. In addition to Jcu-miR390 and Jcu-395a/b, three more miRNAs (Jcu-miR477, Jcu-nMIR011, and Jcu-nMIR023) were observed to be differentially expressed from young to intermediate seeds, which was still much less than the number of differentially expressed miRNAs from young to mature (29 miRNAs). Nevertheless, the trend of expression variation of miRNAs from RT-qPCR agreed very well with the sequencing results in terms of up or down regulation (Supplementary Fig. S4). A heatmap was made to gain an overview of the differentially expressed miRNAs (Fig. 3). The global expressions of the 29 differentially expressed miRNAs had high correlations among three replicates, and samples from three different developmental stages of J. curcas seeds were grouped into three clades accordingly, suggesting that the data sets were reliable and ideal for statistical analysis for differentially expressed miRNAs (Fig. 3). The heatmap profiles were similar between young seeds and intermediate ones, which might be due to the young seeds were in the histodifferentiation stage and the intermediate seeds were in the early transition from histodifferentiation to seed filling stage70, and the two stages might have overlapped physiological process such as cell division and tissue differentiation. Nevertheless, a huge difference for the global profile of miRNA levels in mature seeds was observed when compared with the young and intermediate stages (Fig. 3). During the transition from intermediate to mature stage, seeds were developing much faster than before in regards to length, fresh weight and dry weight, and were suffering desiccation stress (Supplementary Fig. S1). These results from deep sequencing and RT-qPCR both suggested that the post-transcriptionally regulation of genes by miRNAs mainly happened at the late seed developmental stages when seeds experienced organic matter accumulation and desiccation stress.
The global expression of the 29 differentially expressed miRNAs in three developmental stages (young, intermediate and mature) of J. curcas seeds. Three independent biological replicates of developing seeds at each stage (young: S07, S08 and S09; intermediate: S04, S05, and S06; mature: S01, S02, and S03) were shown. The colors indicate relative expression levels of miRNAs, where high levels are indicated by red, and low levels are green.
Identification of IsomiRNAs
Some miRNA variants, known as isomiRNAs (isomiRs), have additional nucleotides in the 5′ or 3′ terminus when compared to canonical miRNAs, which might be a consequence of inaccuracies in Dicer pre-miRNA processing or post-transcriptional modification71,72,73,74. A total of 36 isomiRs belonging to 13 families (Jcu-MIR 159, 166, 167, 319, 393, 396, nMIR003, nMIR004, nMIR005, nMIR010, nMIR022, nMIR026, nMIR027; see Supplementary Table S6) were detected. Among these isomiRs, thirty 3′ isomiRs (3′ deletion, extension and nontemplated modifications), seven 5′ isomiRs (5′ deletion and extension) and four polymorphic isomiRs (only harbor distinct nucleotide compositions within the miRNA sequences) were identified, which agrees to the fact that 3′ isomiRs are the most frequently observed type of isomiR in animals and plants, in terms of both number and abundance71. Interestingly, no polymorphism within the seed sequence (2–9 nt) was observed, and it might suggest that the modification in these miRNA variants has little effect on target selection, because most of the isomiRs had similar targets with the canonical miRNAs (data not shown). The target mRNAs are recognized by multiple isomiRs plus the canonical miRNA, which might increase the specificity and efficacy of the target silencing75.
When inspecting the expression of these isoform variants among nine libraries, most isoforms displayed a similar trend of expression during seed development, whereas only a few isoforms had significant divergence. It is noteworthy that isomiR319-1 increases while isomiR319-3 decreases sharply during the seed developing process (Supplementary Table S5), suggesting that isomiR members in the same family might have different regulation mechanism.
Conclusions
The identification of 93 high confidence miRNAs and their targets provides useful insights into miRNA-mediated regulatory mechanism, substance accumulation, and the adaption to dehydration stress during seed development of J. curcas. A lot of miRNAs were predicted to regulate target genes involved in transcription regulation, lipid metabolism and other important biological processes. These miRNAs might be valuable in improving the yield and quality of J. curcas seed oil. Most differentially expressed miRNAs were observed in the late stage of seed development, indicating that miRNAs play important roles in the accumulation of organic matter and dehydration stress during seed maturation. However, further studies are necessary to validate the miRNA targets and the roles of miRNAs in the complex regulation mechanism.
Materials and Methods
Seed Collection and RNA Isolation
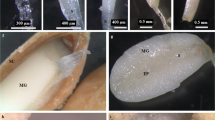
Fruits from J. curcas L. trees grown in Xishuangbanna Tropical Botanical Garden (21°54′ N, 101°46′ E, 580 m a.s.l.,) located in Mengla County, Yunnan Province, China, were collected randomly from 6 plants at 5–10 (young seeds, with seed length less than 10 mm), 12–20 (intermediate seeds, with seed length in the range of 13 to 17 mm) and 25–35 (mature seeds, with seed length more than 20 mm) days after flower opening (DAF) (Supplementary Fig. S1). Three independent biological replicates of developing seeds at each stage were collected and immediately frozen in liquid nitrogen and then stored at −80 °C. Total RNA was isolated from a pool of seeds from each stage with Trizol (Invitrogen, CA, USA), according to the manufacturer’s protocol.
sRNA Library Construction and Deep Sequencing
Total RNA (1.5 μg per sample) was sent to Biomarker Technologies (Beijing, China) for sRNA library construction and deep sequencing. The sRNA libraries were generated using NEBNext® UltraTMsmall RNA Sample Library Prep Kit for Illumina® (NEB, USA) following the manufacturer’s recommendations. In brief, RNA bands corresponding to a size range of 16–30 nt were separated and purified from the acrylamide gel. The sRNA molecules ligated with 5′ and 3′ adaptors were used for reverse transcription and subsequent PCR. Small RNA samples were sequenced using Illumina HiSeq. 2500 platform (San Diego, CA, USA) and single-end reads were generated. Three independent sRNA libraries were constructed from each of the three stages of developing seeds.
Sequence Data Analysis
The reads generated by sRNA-sequencing (named raw reads) were preprocessed with Fastx-toolkit (http://hannonlab.cshl.edu/fastx_toolkit/). First, the raw reads were processed to remove adaptors. Then reads with >20% bases <Q30, or with N base >10%, or with ployA/T/C/G, or read length < 18 nt or >30 nt, were filtered. The remaining reads named clean reads were used for further analysis. sRNAs derived from Viridiplantae rRNAs, scRNA, snRNAs, snoRNAs, tRNA, and repetitive elements (from the SILVA rRNA, Rfam, tRNAdb, and Repbase databases) were identified by mapping with Bowtie v 0.12.9 without mismatches76. The remaining reads named unannotated reads without rRNA, scRNA, snRNA, snoRNA, tRNA, or repetitive element were used for miRNA identification.
miRNA Identification
The unannotated reads containing miRNAs were processed for miRNA identification by miRDeep2 v 2.0.0.531 with modified parameters (sRNAs ≤ 15 hits; 250 nt flanking sequences) for plant miRNA characters30. In brief, the analysis procedure of miRDeep2 were: (1) The unannotated reads were aligned to the Jatropha genome sequences using Bowtie v 0.12.9 with perfect matches76, and only sRNAs with no more than 15 hits were kept and their flanking sequences (250 nt on each side) on the genome were extracted as candidate miRNA precursors, to satisfy the parameters of miRNAs in plant species as described25; (2) the precursors were folded in silico using RNAfold (v2.1.7; default parameters) to find the precursors with expected structures; and (3) the unannotated reads were aligned to the precursors, and statistical analyses of read distributions were generated for true miRNA evaluation. To select high confidence miRNAs, more strict criteria were adopted to check the miRDeep2 output figures manually25,26,32. Candidate miRNAs were then aligned to the known miRNAs from miRBase 21.0. The miRNAs shared homology with known miRNAs (allowed two mismatches) were identified as conserved miRNA, and those shared no homology to all known sequences in miRBase, novel miRNAs. Isoform miRNAs (isomiRNAs) were analyzed using isomiRID v 0.53 (using default parameters except for cutoff value 100; RangeSize: 18~26)77. The reads that were perfectly mapped in the annotated miRNA precursors but not representing the identified miRNA mature and star sequences, shifted not more than four positions from their original mature or star miRNA 5′ position (allowing one mismatch in the middle), and had a total number of reads not less than 50% of the total reads of their reference miRNA or had a total reads of more than 10,000, were considered as isoform miRNAs in Jatropha.
miRNA Target Prediction
The miRNA target prediction was performed by aligning the mature miRNA sequences against J. curcas RNA sequences, with Targetfinder v 1.6 using a strict prediction score cutoff value 3 (default = 4)36 and psRNAtarget (2017 release) using default parameters35. Target annotation was performed by using BLASTX (blast 2.2.25+; -evalue 1e-6, -word_size 6, and -max_target_seqs. 10) based on sequence similarity with genes from the NCBI nr, Swiss-Prot/Uniprot protein databases (http://www.expasy.ch/sprot), and Clusters of Orthologous Groups of proteins (COG) database (http://www.ncbi.nlm.nih.gov/COG/). Blast2GO was employed to obtain Gene Ontology (GO) annotation (http://www.geneontology.org) according to molecular function, biological process, and cellular component ontology with default parameters to evaluate their putative functions78. For those miRNA targets could not be mapped by Blast2GO, eggNOG (v4.5) was used to characterize their functional terms79.
In Silico Expression Analysis of miRNAs
The read count for each miRNA was obtained from each sample by mapping sRNA back onto the precursor sequence using the quantifier module of miRDeep2 v 2.0.0.5 with parameters allowing one mismatch and a small window of 2 nt upstream and 5 nt downstream as described by Friedländer et al.31. To compare the abundances of miRNA between different samples, a normalized read counts named TPM value for each sample was defined as ‘counts of reads mapped to miRNA × 1 000 000’/‘reads mapped to the reference genome’33,36. The DESeq2 software provides statistical routines for determining differential expression in digital miRNA expression data using a model based on the negative binomial distribution80, and therefore the read counts of miRNAs from different samples were further analyzed using the DESeq2R package (1.14.1) with default parameters. The resulting ‘log2 fold change’ represented the base-2 logarithm of fold change of expression levels between any two of young, intermediate, and mature stages; the resulting adjusted p values were p values adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate (FDR). The miRNAs whose expression levels between any two of the three different developmental stages varied significantly (|log2 fold change| >1; FDR adjusted p < 0.05) were assigned as differentially expressed ones.
RT-qPCR
RT-qPCR was performed following previously reported procedures81. In brief, small RNAs (<200 bp) were extracted with the BioTeKe miRNA extraction kit (BioTeKe Corporation, China). The extracted small RNAs were treated with DNase I and polyadenylated by poly(A) polymerase (New England Biolabs) following by reverse transcription using SuperScript™ III reverse transcriptase (Invitrogen) according to the manufacturer’s instructions. Real-time PCR was performed following a standard SYBR Premix Ex Taq II (TaKaRa) protocol. PCRs were performed in ABI StepOne (USA) as follows: 95 °C for 5 min; then followed by 40 cycles of the normal program (95 °C for 10 s, 60 °C for 30 s). The differences in gene expression were calculated using the 2−ΔΔCt analysis method82, and the transcription levels were determined by relative quantification using the 5.8 S RNA as internal reference gene and the young seeds as control. All primers use in RT-qPCR experiments were listed in Supplementary Table S6.
Data Availability
All sequencing data were deposited in the NCBI Sequence Read Archive under ID SRA616450. All data generated or analyzed during this study are included in this published article and its Supplementary Information files.
References
Yang, M. F. et al. Proteomic analysis of oil mobilization in seed germination and postgermination development of Jatropha curcas. J. Proteome Res. 8, 1441–1451 (2009).
Kandpal, J. B. & Madan, M. J. Jatropha curcus: a renewable source of energy for meeting future energy needs. Renew. Energ. 6, 159–160 (1995).
Abdulla, R., Chan, E. S. & Ravindra, P. Biodiesel production from Jatropha curcas: a critical review. Crit. Rev. Biotechnol. 31, 53–64 (2011).
Openshaw, K. A review of Jatropha curcas: an oil plant of unfulfilled promise. Biomass Bioenergy 19, 1–15 (2000).
Berchmans, H. J. & Hirata, S. Biodiesel production from crude Jatropha curcas L. seed oil with a high content of free fatty acids. Bioresour. Technol. 99, 1716–1721 (2008).
Xu, G., Huang, J., Yang, Y. & Yao, Y. A. Transcriptome analysis of flower sex differentiation in Jatropha curcas L. using RNA sequencing. PLoS ONE 11, e0145613 (2016).
Natarajan, P. et al. Gene discovery from Jatropha curcas by sequencing of ESTs from normalized and full-length enriched cDNA library from developing seeds. BMC Genomics 11, 606 (2010).
Costa, G. G. L. et al. Transcriptome analysis of the oil-rich seed of the bioenergy crop Jatropha curcas L. BMC Genomics 11, 462 (2010).
Xu, R. H., Wang, R. L. & Liu, A. Z. Expression profiles of genes involved in fatty acid and triacylglycerol synthesis in developing seeds of Jatropha (Jatropha curcas L.). Biomass Bioenergy 35, 1683–1692 (2011).
Jiang, H. W. et al. Global analysis of gene expression profiles in developing physic nut (Jatropha curcas L.) seeds. Plos One 7, e36522 (2012).
Zhang, L. et al. Global analysis of gene expression profiles in physic nut (Jatropha curcas L.) seedlings exposed to salt stress. Plos One 9, e97878 (2014).
Sato, S. et al. Sequence analysis of the genome of an oil-bearing tree, Jatropha curcas L. DNA Res. 18, 65–76 (2010).
Hirakawa, H. et al. Upgraded genomic information of Jatropha curcas L. Plant Biotechnol. 29, 123–130 (2012).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 136, 669–687 (2009).
Kurihara, Y. & Watanabe, Y. Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc. Natl. Acad. Sci. USA 101, 12753–12758 (2004).
Rogers, K. & Chen, X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 25, 2383–2399 (2013).
de Lima, J. C., Loss-Morais, G. & Margis, R. MicroRNAs play critical roles during plant development and in response to abiotic stresses. Genet. Mol. Biol. 35, 1069–1077 (2012).
Korbes, A. P. et al. Identifying conserved and novel microRNAs in developing seeds of Brassica napus using deep sequencing. Plos One 7, e50663 (2012).
Chuck, G., Candela, H. & Hake, S. Big impacts by small RNAs in plant development. Curr. Opin. Plant Biol. 12, 81–86 (2009).
Wang, C. M. et al. Isolation and Identification of miRNAs in Jatropha curcas. Int. J. Biol. Sci. 8, 418–429 (2012).
Vishwakarma, N. P. & Jadeja, V. J. Identification of miRNA encoded by Jatropha curcas from EST and GSS. Plant Signal. Behav. 8, e23152 (2013).
Galli, V. et al. Identifying microRNAs and transcript targets in Jatropha seeds. Plos One 9, e83727 (2014).
Meyers, B. C. et al. Criteria for annotation of plant MicroRNAs. Plant Cell 20, 3186–3190 (2008).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–73 (2014).
Huang, D., Koh, C., Feurtado, J., Tsang, E. & Cutler, A. MicroRNAs and their putative targets in Brassica napus seed maturation. BMC Genomics 14, 140 (2013).
Wang, Z., Qiao, Y., Zhang, J., Shi, W. & Zhang, J. Genome wide identification of microRNAs involved in fatty acid and lipid metabolism of Brassica napus by small RNA and degradome sequencing. Gene 619, 61–70 (2017).
Cao, S. et al. Comparative profiling of miRNA expression in developing seeds of high linoleic and high oleic safflower (Carthamus tinctorius L.) plants. Front. Plant Sci. 4, 489 (2013).
Yang, X. & Li, L. miRDeep-P: a computational tool for analyzing the microRNA transcriptome in plants. Bioinformatics 27, 2614–2615 (2011).
Friedlander, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52 (2012).
Taylor, R. S., Tarver, J. E., Hiscock, S. J. & Donoghue, P. C. Evolutionary history of plant microRNAs. Trends Plant Sci. 19, 175–182 (2014).
Yin, H. et al. Phylogenetic tree-informed microRNAome analysis uncovers conserved and lineage-specific miRNAs in Camellia during floral organ development. J. Exp. Bot. 67, 2641–2653 (2016).
Bartel, B. & Bartel, D. P. MicroRNAs: at the root of plant development? Plant Physiol. 132, 709–717 (2003).
Dai, X. & Zhao, P. X. psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res. 39, W155–159 (2011).
Fahlgren, N. et al. High-throughput sequencing of Arabidopsis microRNAs: evidence for frequent birth and death of MIRNA genes. Plos One 2, e219 (2007).
Rogans, S. J. & Rey, C. Unveiling the micronome of cassava (Manihot esculenta Crantz). Plos One 11, e0147251 (2016).
Cuperus, J. T., Fahlgren, N. & Carrington, J. C. Evolution and functional diversification of MIRNA genes. Plant Cell 23, 431–442 (2011).
Manna, S. An overview of pentatricopeptide repeat proteins and their applications. Biochimie 113, 93–99 (2015).
Pinot, F. & Beisson, F. Cytochrome P450 metabolizing fatty acids in plants: characterization and physiological roles. FEBS J. 278, 195–205 (2011).
Divakara, B. N., Upadhyaya, H. D., Wani, S. P. & Gowda, C. L. L. Biology and genetic improvement of Jatropha curcas L.: A review. Appl. Energy 87, 732–742 (2010).
Power, R., Kurz, W. G. & De Luca, V. Purification and characterization of acetylcoenzyme A: deacetylvindoline 4-O-acetyltransferase from Catharanthus roseus. Arch. Biochem. Biophys. 279, 370–376 (1990).
Takahashi, S. et al. Functional characterization of premnaspirodiene oxygenase, a cytochrome P450 catalyzing regio- and stereo-specific hydroxylations of diverse sesquiterpene substrates. J. Biol. Chem. 282, 31744–31754 (2007).
Zhang, K. et al. Investigation of miR396 and growth-regulating factor regulatory network in maize grain filling. Acta Physiol. Plant. 37, 1–12 (2015).
Samad, A. F. A. et al. MicroRNA and transcription factor: key players in plant regulatory network. Front. Plant Sci. 8, 565 (2017).
Laufs, P., Peaucelle, A., Morin, H. & Traas, J. MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 131, 4311–4322 (2004).
McMichael, C. M. et al. Mediation of clathrin-dependent trafficking during cytokinesis and cell expansion by Arabidopsis stomatal cytokinesis defective proteins. Plant Cell 25, 3910–3925 (2013).
Jaber, E. et al. A putative TRAPPII tethering factor is required for cell plate assembly during cytokinesis in Arabidopsis. New Phytol. 187, 751–763 (2010).
Hotta, T. et al. Characterization of the Arabidopsis augmin complex uncovers its critical function in the assembly of the acentrosomal spindle and phragmoplast microtubule arrays. Plant Cell 24, 1494–1509 (2012).
Prigge, M. J. et al. Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17, 61–76 (2005).
Baima, S. et al. The arabidopsis ATHB-8 HD-zip protein acts as a differentiation-promoting transcription factor of the vascular meristems. Plant Physiol. 126, 643–655 (2001).
Green, K. A., Prigge, M. J., Katzman, R. B. & Clark, S. E. CORONA, a member of the class III homeodomain leucine zipper gene family in Arabidopsis, regulates stem cell specification and organogenesis. Plant Cell 17, 691–704 (2005).
Ochando, I. et al. Mutations in the microRNA complementarity site of the INCURVATA4 gene perturb meristem function and adaxialize lateral organs in Arabidopsis. Plant Physiol. 141, 607–619 (2006).
Pagnussat, G. C. et al. Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 132, 603–614 (2005).
Schommer, C. et al. Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol. 6, e230 (2008).
Jofuku, K. D., den Boer, B. G., Van Montagu, M. & Okamuro, J. K. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 6, 1211–1225 (1994).
Koyama, T., Furutani, M., Tasaka, M. & Ohme-Takagi, M. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis. Plant Cell 19, 473–484 (2007).
Riefler, M., Novak, O., Strnad, M. & Schmulling, T. Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 18, 40–54 (2006).
Wojcik, A. M. & Gaj, M. D. miR393 contributes to the embryogenic transition induced in vitro in Arabidopsis via the modification of the tissue sensitivity to auxin treatment. Planta 244, 231–243 (2016).
Yang, J. H., Han, S. J., Yoon, E. K. & Lee, W. S. Evidence of an auxin signal pathway, microRNA167-ARF8-GH3, and its response to exogenous auxin in cultured rice cells. Nucleic Acids Res. 34, 1892–1899 (2006).
Kushwaha, R., Singh, A. & Chattopadhyay, S. Calmodulin7 plays an important role as transcriptional regulator in Arabidopsis seedling development. Plant Cell 20, 1747–1759 (2008).
Gong, Z. et al. RNA helicase-like protein as an early regulator of transcription factors for plant chilling and freezing tolerance. Proc. Natl. Acad. Sci. USA 99, 11507–11512 (2002).
Gong, Z. et al. A DEAD box RNA helicase is essential for mRNA export and important for development and stress responses in Arabidopsis. Plant Cell 17, 256–267 (2005).
Tang, M., Liu, X., Deng, H. & Shen, S. Over-expression of JcDREB, a putative AP2/EREBP domain-containing transcription factor gene in woody biodiesel plant Jatropha curcas, enhances salt and freezing tolerance in transgenic Arabidopsis thaliana. Plant Sci. 181, 623–631 (2011).
Kanno, Y. et al. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl. Acad. Sci. USA 109, 9653–9658 (2012).
Ifuku, K., Endo, T., Shikanai, T. & Aro, E. M. Structure of the chloroplast NADH dehydrogenase-like complex: nomenclature for nuclear-encoded subunits. Plant Cell Physiology 52, 1560–1568 (2011).
Ren, H. & Gray, W. M. SAUR proteins as effectors of hormonal and environmental signals in plant growth. Mol. Plant 8, 1153–1164 (2015).
Hundertmark, M. & Hincha, D. K. LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genomics 9, 118 (2008).
Candat, A. et al. The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26, 3148–3166 (2014).
Liu, H. et al. Proteomic analysis of the seed development in Jatropha curcas: From carbon flux to the lipid accumulation. J. Proteomics 91, 23–40 (2013).
Neilsen, C. T., Goodall, G. J. & Bracken, C. P. IsomiRs–the overlooked repertoire in the dynamic microRNAome. Trends Genet. 28, 544–549 (2012).
Guo, L. & Chen, F. A challenge for miRNA: multiple isomiRs in miRNAomics. Gene 544, 1–7 (2014).
Liang, T., Yu, J., Liu, C. & Guo, L. IsomiR expression patterns in canonical and Dicer-independent microRNAs. Mol. Med. Report. 15, 1071–1078 (2017).
Ebhardt, H. A., Fedynak, A. & Fahlman, R. P. Naturally occurring variations in sequence length creates microRNA isoforms that differ in argonaute effector complex specificity. Silence 1, 12 (2010).
Ahmed, F. et al. Comprehensive analysis of small RNA-seq data reveals that combination of miRNA with its isomiRs increase the accuracy of target prediction in Arabidopsis thaliana. RNA Biol. 11, 1414–1429 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
de Oliveira, L. F., Christoff, A. P. & Margis, R. isomiRID: a framework to identify microRNA isoforms. Bioinformatics 29, 2521–2523 (2013).
Gotz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 36, 3420–3435 (2008).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44, D286–293 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Zeng, C. Y. et al. Conservation and divergence of microRNAs and their functions in Euphorbiaceous plants. Nucleic Acids Res. 38, 981–995 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-delta delta Ct method. Methods 25, 402–408 (2001).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31370674; 21606020); Beijing Natural Science Foundation (2164059); Funding Project for Scientific Research Quality Improvement in Beijing University of Agriculture (GJB2013001). We thank Prof. Rogerio Margis for helping us in the isomiR analysis.
Author information
Authors and Affiliations
Contributions
M.F.Y. and L.Q.M. designed the experiments, analyzed the data and wrote the main manuscript text; F.Y.X. prepared Supplementary Figs S1–S4; H.S.L. analyzed the data and prepared all other figures and tables. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, M., Lu, H., Xue, F. et al. Identifying High Confidence microRNAs in the Developing Seeds of Jatropha curcas. Sci Rep 9, 4510 (2019). https://doi.org/10.1038/s41598-019-41189-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41189-y
This article is cited by
-
Comparative miRNome and transcriptome analyses reveal the expression of novel miRNAs in the panicle of rice implicated in sustained agronomic performance under terminal drought stress
Planta (2024)
-
Comparative analysis of the carrot miRNAome in response to salt stress
Scientific Reports (2023)
-
An insight into microRNA biogenesis and its regulatory role in plant secondary metabolism
Plant Cell Reports (2022)
-
MiRNA fine tuning for crop improvement: using advance computational models and biotechnological tools
Molecular Biology Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.