Abstract
To understand the mechanism underlying metastasis, identification of a mechanism-based and common biomarker for circulating tumour cells (CTCs) in heterogenous breast cancer is needed. SET, an endogenous inhibitor of protein phosphatase 2A, was overexpressed in all subtypes of invasive breast carcinoma tissues. Treatment with SET-targeted siRNAs reduced the motility of MCF-7 and MDA-MB-231 cells in transwell assay. SET knockdown reduced the number of mammospheres by 60–70% in MCF-7 and MDA-MB-231 cells, which was associated with the downregulation of OCT4 and SLUG. Hence, we analysed the presence of SET-expressing CTCs (SET-CTCs) in 24 breast cancer patients. CTCs were enriched using a size-based method and then immunocytochemically analysed using an anti-SET antibody. SET-CTCs were detected in 6/6 (100%) patients with recurrent breast cancer with a median value of 12 (12 cells/3 mL blood), and in 13/18 (72.2%) patients with stage I–III breast cancer with a median value of 2.5, while the median value of healthy controls was 0. Importantly, high numbers of SET-CTCs were correlated with lymph node metastasis in patients with stage I–III disease. Our results indicate that SET contributes to breast cancer progression and can act as a potential biomarker of CTCs for the detection of metastasis.
Similar content being viewed by others
Introduction
Breast cancer is a heterogeneous disease classified into four clinical subtypes based on the expression of hormone receptors (HR), including oestrogen and progesterone receptors (ER/PR) and human epidermal growth factor receptor 2 (HER2)1. Although adjuvant systemic therapy for breast cancer patients improves survival, some patients still eventually develop metastatic lesions. Early detection of metastasis, as well as the development of new mechanism-based agents, is important for overcoming metastatic disease. To achieve this goal, a comprehensive understanding of the characteristics of primary tumours and cancer cells in circulation, called circulating tumour cells (CTCs), is critical, as CTCs are believed to be “seeds” of metastasis2,3. Currently, the enumeration of epithelial cell adhesion molecule (EpCAM)-positive CTCs (EpCAM-CTCs) has allowed estimation of overall metastatic burden in patients with breast cancer4. However, EpCAM-CTCs are undetectable in one third of patients with metastatic disease5. As many cancer cells lose epithelial features due to epithelial-mesenchymal transition (EMT) during metastasis6,7, identification of a new molecule that plays a key role in metastasis and that is expressed in CTCs is necessary. The molecule is expected to help improve early detection of metastasis and evaluation of treatment response in patients.
SET is a 39-kDa oncoprotein encoded by the set gene, which was discovered as a component of a set-can fusion gene in acute myeloid leukemia (AML)8. Subsequently, Li et al. observed that SET protein is a potent and specific inhibitor of protein phosphatase 2A (PP2A) based on the results that a heat-stable PP2A inhibitor called I2PP2A was a truncated form of SET9,10. PP2A is a major serine/threonine phosphatase regulating multiple signalling pathways, including those involved in cell proliferation and metastasis, which antagonises kinase activities11. The inhibition of PP1 and PP2A is a common mechanism underlying tumour promotion in various organs of rodents: The okadaic acid class compounds, including okadaic acid, dinophysistoxin-1, calyculin A, microcystin-LR, and nodularin, are chemical inhibitors of PP1 and PP2A and induce tumour-promoting activities in mouse skin, rat glandular stomach, and rat liver12. Furthermore, the transforming simian vacuolating virus 40 (SV40) small T-antigen inhibits PP2A activity and transforms human cells13. Unusual inhibition of PP2A promotes the malignant transformation of normal cells. Studies have shown that PP2A is a unique tumour suppressor, which is rarely mutated or deleted, but its function is impaired by “inhibitor proteins”14. Abnormal overexpression of cellular PP2A inhibitors induces dysregulation of signalling pathways by increasing the levels of phosphoproteins, similar to abnormal activation of serine/threonine kinases. SET is a physiological endogenous PP2A inhibitor that is 30-fold weaker than the okadaic acid class of compounds12. Importantly, SET protein levels are high in various cancers, including breast, colon, pancreas, gastric, liver and lung cancers. Likewise, SET protein levels were also found to be high in leukemia, including chronic myeloid leukemia (CML) and acute myeloid leukemia (AML) and lymphoma15,16,17,18,19,20,21,22. Overexpression of SET perturbs various signalling pathways, and high SET levels correlate with drug resistance and a poor prognosis in various human cancers. In addition to SET, cancerous inhibitor of PP2A (CIP2A), another well-known cellular inhibitor of PP2A, is often overexpressed in various cancer tissues23,24. Among the known cellular inhibitors of PP2A (SET, CIP2A and protein phosphatase methylesterase 1 (PME-1)), SET is the most potent and regulates 30.5% of the phosphopeptides dephosphorylated by PP2A. In addition, the majority of the CIP2A- and PME-1-target peptides overlapped with the target peptides of SET25. Furthermore, targeting of SET with siRNA also down-regulates CIP2A26. Therefore, we hypothesised that SET might be the desired target molecule, if it is expressed in CTCs.
In this study, we found that SET was overexpressed in breast cancer tissues of all subtypes. Targeting SET using siRNA in two breast cancer cell lines, MCF-7 (HR-positive) and MDA-MB-231 (triple-negative), revealed that SET plays a significant role in controlling motility and stemness. Using a size-based enumeration method, we identified heterogenous CTCs in breast cancer patients, as reported previously27,28. We compared numbers of SET-expressing CTCs (SET-CTCs) with those of the epithelial-CTCs (expressing pan-cytokeratin and EpCAM) and mesenchymal-CTCs (expressing vimentin and gross cystic disease fluid protein-15 (GCDFP-15)) from the same breast cancer patients. The results presented here indicate that the oncoprotein SET is a useful biomarker for detecting CTCs, irrespective of EMT status, which may be beneficial for breast cancer treatment.
Results
Overexpression of SET in invasive breast carcinoma tissues
SET protein is a nuclear protein, and hence strong expression of SET was detected in the nuclei of invasive breast carcinoma tissues (Fig. 1a). Immunohistochemical staining of breast tissue array containing 24 cases each of normal tissue, adjacent normal tissue, and invasive ductal carcinoma revealed that the SET protein was significantly overexpressed in invasive carcinoma tissues compared to normal and adjacent normal tissues. Immunostaining scores of SET in normal, adjacent normal, and invasive carcinomas were 32.9 ± 15.9, 32.8 ± 18.7, and 148.1 ± 26.2, respectively (Fig. 1b). Next, we analysed the relationship between SET overexpression and breast cancer subtypes with varying expression of HR and HER2, using another breast cancer tissue array including 100 invasive carcinoma tissues from stage II–III breast cancers. As all invasive ductal carcinoma tissues were significantly stained with the anti-SET antibody, we classified them into three levels: level 1 (weak), level 2 (moderate), and level 3 (strong), corresponding to SET scores < 100, 100–130, and ≥ 130. More than 60% of all subtypes, including HR-positive/HER2-negative, HR/HER2-positive, HR-negative/HER2-positive, and triple-negative, showed strong level 3 SET expression (Table 1). In addition, we observed that higher SET expression was associated with higher Ki67 levels, indicating a correlation with high proliferation activity. Thus, the SET protein was overexpressed in invasive carcinoma irrespective of hormone receptor expression status.
Immunohistochemical analysis of breast cancer tissue array. (a) Strong expression of SET was detected in nuclei of invasive carcinoma, but not in those of normal and adjacent normal tissues. (b) Immunostaining scores were determined as described in “Methods” section. Invasive carcinoma tissues showed significantly higher score than those of normal and adjacent normal tissues. *P < 0.01.
Contribution of SET to cell motility and stemness
SET protein is frequently overexpressed in various breast cancer cell lines including MCF-7 and MDA-MB-23115. Using MCF-7 cells as the HR-positive and epithelial type cell line, and MDA-MB-231 cells as the triple-negative and EMT-induced cell line, we analysed the effects of SET knockdown with SET-targeted siRNAs (siSET-1 and siSET-2) in transwell and mammosphere formation assays. Two SET protein bands were detected in MCF-7 and MDA-MB-231 cell lysates. The less abundant 41 kDa protein was confirmed as SET-α. Both cell lines expressed large amounts of SET-β and small amounts of SET-α. Treatment with siSET-1 and siSET-2 reduced both SET isomers protein levels by > 50% in both MCF-7 and MDA-MB-231 cells, but not after treatment with siControl (Fig. 2a). Similar to the results observed with SET knockdown, the level of CIP2A was also reduced by > 50% in both cell lines, suggesting that SET inhibition restored PP2A activity in combination with CIP2A downregulation. This is in agreement with the results of a previous report showing that targeting SET downregulates the CIP2A mRNA and increases PP2A activity in triple-negative breast cancer cells26. We observed that SET knockdown reduced the motility of cells using a transwell assay. Treatment with siSET-1 and siSET-2 reduced the number of migrated cells from 27.3 ± 3.8 to 6.0 ± 2.0 and 5.7 ± 3.1 (79.1% inhibition) in MCF-7 cells, and from 182 ± 54.5 to 64.0 ± 19.2 and 66.3 ± 34.0 (63.5% inhibition) in MDA-MB-231 cells (Fig. 2b). SET knockdown suppressed motility by > 60%, suggesting a significant role of SET overexpression in metastasis in breast cancer.
Knocking down of SET with siSETs reduced motility and mammosphere formation in MCF-7 and MDA-MD-231 cells. (a) Treatment with siSET-1 and siSET-2 reduced SET protein level, which was associated with reduction of CIP2A. Original full images in the figure were shown in Supplementary information, Figure S1. Due to cut the membrane prior hybridization or due to clear backgrounds, some of blot images have absent membrane edges, but specificities of bands were confirmed by knocking down using siRNAs. (b) Motility of MCF-7 and MDA-MD-231 cells was reduced upon SET knockdown. (c) SET knockdown inhibited mammosphere formation in MCF-7 and MDA-MB-231 cells. (d) OCT4 and SLUG expression in mammospheres was suppressed by SET knockdown in MCF-7 and MDA-MB-231 cells. *P < 0.01.
Furthermore, SET knockdown significantly suppressed mammosphere formation in MCF-7 and MDA-MB-231 cells. MCF-7 cells produced approximately 300 spheres with an average major axis of 150 μm for 2 weeks. Upon treatment with siSET-1 and siSET-2, the number of mammospheres reduced to 35.3% and 47.3% (64.7% and 52.7% inhibition), respectively; in contrast, treatment with siControl showed a weak reduction (Fig. 2c). Similarly, MDA-MB-231 cells formed about 250 spheres with an average major axis of 220 μm for 3 weeks. Treatment with siSET-1 and siSET-2 reduced mammosphere formation to 25.9% and 34.1% (74.1% and 65.9% inhibition), respectively. Furthermore, treatment with siSET-1 and siSET-2 significantly reduced the expression of OCT4, a stemness-related gene, and SLUG, an EMT-related gene, in mammospheres in both MCF-7 and MDA-MB-231 cells (Fig. 2d); however, CD133, NANOG, or SOX2 expression was not altered (Supplementary Table S1). These results suggested that SET overexpression contributed to the maintenance of stemness and probably metastasis in both HR-positive and triple-negative breast cancers.
Inhibition of tumour formation by MCF-7 cells following SET knockdown
Next, we examined the suppressive effect of siSET on tumour formation in vivo using a xenograft model of SCID/Beige mice. Non-treated, siControl-treated, and siSET-1-treated MCF-7 cells (5 × 106 cells/site) were subcutaneously implanted at two sites in the flanks. At the end of the experiment on day 45, small tumours had developed at 75% (6/8) of the injected sites in the siSET-treated group, although tumours were found at 100% of the sites in the non-treated (8/8) and siControl-treated (6/6) groups (Fig. 3a,b). Tumour volume in the siSET-treated group (43.8 ± 11.9 mm3) was significantly smaller than that in the non-treated (149.9 ± 33.0 mm3) and siControl-treated (89.2 ± 37.9 mm3) groups. Tumour weight also decreased significantly from 115.1 mg ± 28.9 mg to 26.9 ± 23.5 mg (Fig. 3b). Immunohistochemistry confirmed the reduction in SET protein levels in tumours developed in the siSET-treated group: H-scores of tumours were 187.5 ± 52.0 for non-treated group, 198.3 ± 45.8 for siControl, and 66.7 ± 39.8 for siSET group, P < 0.05 (Fig. 3c). These results also indicated the contribution of SET to the progression of breast cancer.
Knocking down of SET inhibited tumour formation of MCF-7 cells in SCID/Beige mice. (a) siSET-treated MCF-7 cells (●) showed lower incidence (right) and smaller tumours (left) than non-treated (○) and siControl-treated MCF-7 (△) cells. (b) The images show all tumours formed in non-treated (8 sites), siControl-treated (6 sites), and siSET-1-treated (8 sites) mice at the end of the experiment; the graph indicates the weight of each tumour in the three groups. (c) Immunohistochemical analysis showed that SET protein level was significantly lower in tumours developed in siSET-treated group than in those in the non-treated and siControl-treated groups. *P < 0.05.
Significance of SET-expressing CTCs in patients with breast cancer
To examine whether SET can identify both epithelial and non-epithelial CTCs, we used a size-base microfluidic device, ClearCell FX, which successfully enriched heterogenous CTCs from blood samples of breast cancer patients27. Before assessing the presence of SET-expressing CTCs (SET-CTCs) in blood samples of patients with breast cancer, we confirmed that the size-based microfluidic device was more efficient in CTC enumeration than the CELLSEARCH system, which has been cleared by the Food and Drug Administration, USA4. The size-based device detected 3–12 epithelial-CTCs in 3.0 mL of blood in all five examined patients with advanced disease, whereas the CELLSEARCH system did not detect any CTCs in 7.5 mL of blood from the same patients. Furthermore, CTCs without epithelial features were also detected in two patients (Supplementary Table S2).
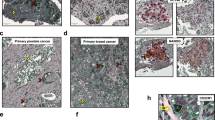
SET-CTCs were immunocytochemically analysed as nucleated cells (DAPI-positive) > 10 μm in diameter that stained with anti-SET (green) antibody, but not with anti-CD45 antibody (red) in patients with breast cancer (Fig. 4a). SET-CTCs were present in 13/18 (72.2%) stage I–III patients with a median value of 2.5 (2.5 cells/3 mL: range, 0–14), and 6/6 (100%) patients with recurrent disease with a median value of 12 (range, 3–38). In contrast, SET-CTCs were detected in 4/10 healthy controls with a median value of 0 (range, 0–3), indicating that the baseline of SET-CTCs may be 3 cells. When the cut-off value set to 4, the positivity of SET-CTCs was 0% for healthy controls, 44.4% for patients in stage I–III, and 83.3% for patients with recurrent disease (Table 2). The number of SET-CTCs and positivity of patients in stage I–III disease and recurrent disease were significantly higher than those in the healthy controls (P < 0.05) (Table 2, Fig. 4a). Furthermore, the number of CTCs in patients with recurrent disease was significantly higher than that in patients with stage I–III disease. To understand the relationship between SET-CTCs and other CTC subpopulations, including CTCs with epithelial and mesenchymal features, we examined the numbers of epithelial-CTCs, which are pan-cytokeratin-and EpCAM-positive and CD45-negative, and mesenchymal-CTCs, which are vimentin-positive and GCDFP-15-positive in the same patients, as described in “Methods” section. Epithelial-CTCs were detected in 3/18 patients with stage I–III disease (16.7%) with a median value of 0 (range, 0–5), and in 4/6 (66.7%) patients with recurrent breast cancer with a median value of 4 (range, 0–21) (Table 2, Fig. 4b). In contrast, mesenchymal-CTCs were detected in 13/18 (72.2%) patients with stage I–III disease with a median value of 1.5 (range, 0–26), and in 5/6 (83.3%) patients with recurrent disease with a median value of 6.5 (range, 0–12) (Table 2, Fig. 4c). As shown in Fig. 4d, all 7 patients with epithelial-CTCs also harboured SET-CTCs, and 6 patients harboured mesenchymal-CTCs. Among 18 patients with mesenchymal-CTCs, 14 were positive for SET-CTCs. These results suggested that SET is widely expressed in both epithelial- and mesenchymal-CTCs. In this study, most patients were HR-positive/HER2 negative. Three patients with recurrent disease and one stage I-III patients were triple-negative. SET-CTCs were found in all four triple-negative patients. However, epithelial-CTCs were not found in the triple-negative stage I-III patient.
Immunocytochemical analysis of SET-CTCs, epithelial-CTCs and mesenchymal-CTCs. (a) Staining of the nucleus with anti-SET antibody (green), along with DAPI staining, but not with anti-CD45 antibody (absence of red). The number of SET-CTCs differed significantly between healthy control and patients with stage I–III disease, healthy control versus patients with recurrent disease, and patients with stage I–III disease versus patients with recurrent disease. (b) Representative staining of epithelial-CTCs with anti-pan-cytokeratin and anti-EpCAM antibodies (green), along with DAPI staining (blue), but not anti-CD45 antibody (absence of red). The number of epithelial-CTCs differed significantly between healthy control and patients with recurrent disease, and patients with stage I–III disease versus patients with recurrent disease. (c) Representative staining of a mesenchymal-CTC with anti-vimentin antibody (green) and anti-GCDFP15 antibody (red), along with DAPI staining (blue). The number of mesenchymal-CTCs differed significantly between healthy control and patients with stage I–III disease, and healthy control versus patients with recurrent disease. (d) SET-CTCs were detected not only in patients with epithelial-CTCs but also in those with mesenchymal-CTCs. Collared bars show the number of SET-CTCs (orange), epithelial-CTCs (blue), and mesenchymal-CTCs (gray) in each patient. (e) Patients with lymph node metastasis showed higher numbers of SET-CTCs than those with stage I–III disease. (f) No significant difference was observed in the number of epithelial- or mesenchymal-CTCs between patients with or without lymph node metastasis. *P < 0.05.
Next, we analysed the relationship between the number of SET-CTCs and lymph node metastasis among patients with stage I–III disease, and observed that patients with lymph node metastasis showed a significantly higher number of SET-CTCs than patients without lymph node metastasis; the median numbers were 3 and 1, respectively (Fig. 4e). Unlike in SET-CTCs, no correlation was observed between lymph node metastasis and numbers of epithelial-CTCs or mesenchymal-CTCs (Fig. 4f). Thus, although the sample number size was small, our results clearly indicate that SET is a promising biomarker for detecting CTCs in patients with various subtypes of breast cancer, which can be used for early monitoring of metastasis.
Discussion
Here, we report for the first time that SET is a possible biomarker for CTCs, and it can be used for the detection of metastasis in patients with breast cancer irrespective of the clinical subtype. Immunohistochemical analysis of tissue microarrays, as well as in vitro and in vivo studies using siSETs indicated that overexpression of SET plays a significant role in progression, especially motility and stemness in breast cancer. Our results are consistent with previous reports showing that knockdown of SET with siRNA significantly reduces tumour growth of MDA-MB-231, MDA-MB-436, and MDA-MB-468 in xenograft experiments in non-obese diabetic/SCID/γ-chain null mice15. Furthermore, SET overexpression is associated with worse recurrence-free survival and it has been shown to reduce tamoxifen-induced anti-tumour effect in patients with primary breast cancer29. High SET protein levels are also associated with poor prognosis of metastatic colon cancer, non-small-cell lung cancer, pancreatic cancer, gastric cancer, liver cancer, CML, AML, and lymphoma16,17,18,19,20,21,22. SET enhances cell migration, colony-forming activity, and EMT transition in colorectal cancer30. High expression of SET is shown to contribute to the progression of various types of cancer, suggesting the usefulness of SET as a biomarker of CTCs in other cancers in addition to breast cancer. Therefore, we suggest that SET is a promising non-specific diagnostic marker for monitoring disease progression and treatment response for various cancers, rather than a specific biomarker for breast cancer.
In this study, we used the size-based microfluidic device as a label-free method, which is based on differences in the mechanical features, such as the size of tumour cells and white blood cells (tumour cells are larger than white blood cells)27. Various methods have been developed for isolation of CTCs based on their physical properties, biological characteristics, and tumour-specific surface proteins31. Among all of them, the method used in this study is one that can enumerate heterogenous CTCs28. In the analysis of epithelial-CTCs, large cells were found to be negative for epithelial marker and CD45. We think that some of the large cells were SET-CTCs and mesenchymal-CTCs. In addition, some SET-CTCs were positive for epithelial or mesenchymal markers. Therefore, we believe that CTCs can be detected in patients with undetectable epithelial-CTCs and mesenchymal-CTCs using SET as a biomarker. The number of SET-CTCs in patients with recurrent disease was significantly higher than that in patients with stage I–III disease, and a high number of SET-CTCs were associated with the presence of lymph node metastasis in patients with stage I–III breast cancer. One limitation of the study is that the number of examined patients was small. And, few SET-positive large cells were detected in the healthy controls. As SET is a ubiquitously expressed protein, we need to improve the specificity of the method used for SET-CTC analysis. We think that further follow-up of patients with early-stage breast cancer is required to determine a cut-off value for detection of early metastasis. In our study, we observed 14 SET-CTCs/3 mL of the peripheral blood of a 37-year-old patient with stage II breast cancer, who had axillary lymph node metastases and multiple tumours in the ipsilateral breast. This patient showed the highest number of SET-CTCs among stage I-III patients and the number higher than even the median value of patients with recurrent disease. We believe that combination analysis with SET, epithelial and mesenchymal markers will improve the clinical significance of CTC analysis by increasing sensitivity and specificity. Ito et al. reported that determining the numbers of both mesenchymal- and epithelial-CTCs might predict survival for breast cancer patients receiving eribulin32. Furthermore, analysis of SET-CTCs will enable real-time monitoring of treatment response to therapeutic drugs. We performed a preliminary CTC analysis to compare CTCs numbers before and after treatment in three patients with breast cancer. Two patients treated with eribulin showed a clear reduction in the number of SET-CTCs (from 8 to 0 and from 10 to 1); while the patient treated with conventional chemotherapy did not (from 14 to 22). Although their clinical response has to be confirmed, this pilot study shows the possibility of monitoring therapy response using SET-CTCs.
There are two major isoforms of SET, SET-α and SET-β, and SET-β is reported to be more widely expressed compared with SET-α. In MCF-7 and MDA-MB-231 cells, we detected a major band with 39 kDa and a minor band with 41 kDa using a pan-SET antibody. A monoclonal antibody for specific SET-a reacted with the protein corresponding to the minor band. Therefore, we suggest that low amounts of SET-α and high amounts of SET-β are expressed in both cell lines. Recently, distinctions in transcriptional regulation and function between two isoforms have been reported. Since we used a pan-SET antibody in this CTC study, further investigations using specific antibodies for SET-α and SET-β will provide more insights into the significance of SET isomers in cancer progression.
SET directly binds to the catalytic subunit of PP2A and inhibits its phosphatase activity10. Therefore, SET is a potent inhibitor of protein dephosphorylation among PP2A inhibitors and inhibition of SET with siRNA strongly impacts targeted peptides25. As PP2A is a critical negative regulator of various pathways related to MYC, Wnt, extracellular signal-regulated kinase (ERK), and Akt, SET overexpression induces stabilisation and activation of c-Myc transcriptional activity and related signalling pathways15,33. Furthermore, SET maintains cancer cell stemness by stabilizing phosphorylated E2F1 via inhibition of PP2A in gastric cancer34, and stable SET knockdown inhibits migration and invasion of breast cancer cells via reduction of matrix metalloproteinase 9 (MMP-9)35. Although we did not investigate whether c-Myc, E2F1, or MMP-9 were inhibited by SET knockdown in breast cancer cells in this study, we observed that treatment with siSET-1 and siSET-2 significantly reduced phosphorylation of Akt in MCF-7 cells (data not shown), suggesting the involvement of the Akt signalling pathway. Therefore, SET plays a key role in malignant progression in breast cancer of all subtypes. These results suggest that SET is a proper target to inhibit multiple pathways involving proliferation, motility, invasion, and stemness, which result in tumour promotion, progression, and metastasis.
Recently, reactivation of PP2A activity using small molecules that either antagonise its inhibitors (SET and CIP2A), or directly activate PP2A, has attracted attention as a new strategy for cancer therapy14,33. These are named PP2A activating drugs. Several SET-targeting agents have been developed. For example, COG112, anapolipoprotein E-mimetic peptide interacting with SET, inhibited the motility of various cancer cells33; FTY720, a synthetic sphingosine analogue, binds to SET, leading to PP2A activation and anti-tumour activity in lung cancer and colorectal cancer24,36; and OP449 (COG449), a synthetic peptide, directly binds to SET and/or interferes with its PP2A-inhibition function37. OP449 showed anti-leukemic activity toward tyrosine kinase inhibitor (TKI)-resistant CML and AML in primary patient samples. Notably, combination of TKI and OP449 synergistically reduced the viability and clonogenic potential of leukemic cell lines and primary CD34+ CML progenitors38. The combined effects of an anticancer agent, such as trastuzumab, and SET-targeting drugs on breast cancer patients should be investigated in future.
Based on our results, we concluded that analysis of SET-CTCs will provide new insights regarding the mechanism of metastasis, and useful information for treatment design using SET-targeting drugs in near future.
Methods
Cell culture and reagents
Human breast cancer cell lines, MCF-7 and MDA-MB-231, were purchased from the American Type Culture Collection (Manassas, VA, USA). The cells were grown in Roswell Park Memorial Institute (RPMI) 1640 medium containing 10% foetal bovine serum (FBS; Sigma-Aldrich, MO, USA) at 37 °C in the presence of 5% CO2. The cells, which were confirmed to be mycoplasma-free, were used at less than 20 passages and were authenticated using short-tandem repeat profiling by ATCC. Anti-SET, anti-SET-α (ab1183), and anti-gross cystic disease fluid protein-15 (GCDFP-15) antibodies were purchased from Abcam Plc, (Cambridge, UK), and anti-CIP2A and anti-vimentin antibodies were from Santa Cruz Biotechnology, Inc. (Danvers, MA, USA). Anti-YAP, anti-pan-keratin (CK4, 5, 6, 8, 10, 13, 18), anti-EpCAM, and anti-CD45 antibodies were obtained from Cell Signaling Technology (MA, USA). AF647-anti-cytokeratin (pan-reactive, BioLegend, CA, USA), PE-anti-CD45 (Abnova Co., Taipei, Taiwan), anti-cytokeratin C-36H5 (CK 7, 8, 18, 19, MiltenyiBiotec GmbH, Gladbach, Germany), anti-glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (Trevigen, MD, USA), AF594 anti-rabbit IgG, and AF488 anti-mouse IgG (Life Technologies, CA, USA) were used for the experiments.
Immunohistochemistry
Human paraffin embedded tissue arrays were purchased from US Biomax, Inc. (Rockville, MD, USA). Breast cancer and adjacent normal tissue array (BR724), and breast carcinoma with breast tissue microarray (BC08116d) were used. The tissue arrays were incubated with anti-SET antibody (1:800) for 40 min at 37 °C after blocking with Protein Block (DAKO, Tokyo, Japan), and the secondary antibody (N-Histofine simple stain MAX-PO (Multi), Nichirei Bioscience Inc. Tokyo, Japan) was applied as described previously39. The stained tissues were observed and photographed at equivalent to 40 × using NanoZoomer-XR and NDP.view2 (Hamamatsu Photonics K.K., Shizuoka, Japan). The relative staining intensity of SET in breast tissues was reviewed by two independent individuals. The score was calculated based on the staining intensity as percentage of stained cells × intensity score.
Knockdown of SET with siRNA
SET-targeted siRNAs (siSET-1 and siSET-2) and a control siRNA were purchased from OriGene Technologies, Inc. (Rockville, MD, USA). The sequences of siSET-1 and siSET-2 were 5′-GGTCTCTGTTTAGTAAATACATCAC-3′ and 5′-GCTATGTGGATTGAGTAATGGGATT-3′, respectively. siRNA (10 nM) was transfected into MCF-7 cells using Lipofectamine 3000 (Invitrogen, CA, USA), and in MDA-MB-231 cells using Lipofectamine RNAiMAX (Invitrogen)40. Western blot analysis, mammosphere formation, and transwell assay were conducted after 2 days.
Western blotting
Whole cell lysates were obtained using a lysis buffer, as described previously41. The cell lysates (10 μg) were subjected to sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS–PAGE) and transferred onto a nitrocellulose membrane. The membranes were blocked with 3% skim milk in 20 mM Tris-HCl (pH 7.6) containing 150 mM NaCl with 0.1% Tween 20, and then incubated with the primary antibody overnight at 8 °C. After incubation with horseradish peroxidase-conjugated secondary antibody against rabbit IgG (GE Healthcare, UK) or goat IgG (Santa Cruz, CA, USA), the bound antibody was detected using ImmunoStar LD (Wako Pure Chemical Industries, Ltd., Tokyo, Japan) and a C-DiGit Blot Scanner (LI-COR Biosciences Inc., NE, USA), as described previously41. At least three independent experiments were conducted.
Transwell assay
Cell motility was determined using a Transwell cell culture chamber (Becton–Dickinson, NJ, USA). Cells (1 × 104) in serum-free RPMI 1640 medium were added to the upper chamber and incubated for 6 h with 1% FBS in the lower chamber. The cells that migrated to the other side of the filter with pore size of 8 μm were stained with 0.4% trypan blue and counted, as described previously40. The data represent an average of three independent experiments that were conducted in duplicate.
Mammosphere formation assay
MCF-7 (100 cells/well) or MDA-MB-231 cells (500 cells/well) were seeded into ultra-low-attachment 96-well plates (Corning Inc., Corning, NY, USA) and grown in Dulbecco’s modified Eagle’s medium/F12 medium (Gibco, Rockville, MD) containing 0.45% methylcellulose, 50 ng/mL epidermal growth factor, 50 ng/mL fibroblast growth factor (PeproTech, USA), and B27 supplement (Invitrogen) for 2 or 3 weeks. Each treatment was conducted in six wells. All wells were imaged using an All-in-One fluorescence microscope (BZ-X700, Keyence, Tokyo, Japan), and the number of spheres measuring > 100 µm in a minor axis were counted41. Three independent experiments were conducted.
Quantitative reverse transcription RT-PCR (qRT-PCR)
The mammospheres were collected using a filter with pore size of 77 μm (Spheroid Catch, Watson Co. Ltd., Tokyo, Japan) and total RNA was isolated using Isogen (Nippon Gene Co. Ltd., Toyama, Japan). Total RNA was reverse transcribed into cDNA using oligo(dT)16 and MuLV reverse transcriptase (Thermo Fisher Scientific, Tokyo, Japan), and real-time PCR was conducted using SYBR Green I (LightCycler 480, Roche Life Science, Basel, Switzerland), as described previously41.
Tumour xenograft experiments
All animal experimental protocols were approved by the Animal Care and Use Committee of Saitama University (H30-A-1-12). All methods were carried out in accordance with the guidelines for the Care and Use of Experimental Animals of Saitama University. This in vivo study was carried out in compliance with the ARRIVE guidelines. Five-week-old female SCID/Beige mice were obtained from Charles River Laboratories Japan, Inc., Kanagawa, Japan, and randomly divided into non-treated, siControl-treated, and siSET-treated groups (3 or 4 mice/group). After treatment with siSET-1 or siControl for 2 days, 5 × 106 cells/100 μL saline of MCF-7 cells were injected at two sites of the subcutaneous flanks of mice. Body weight were monitored weekly, and tumour size was measured using a caliper gauge every 3–4 days for 46 days. Tumour volume was calculated using the formula width2 × length/240. Tumour weight were determined after dissection of tumour tissues from euthanized mice with pentobarbital sodium solution (64.8 mg/mL, Kyoritsu Seiyaku Co., Tokyo) at the study end-point.
Enumeration of CTCs from peripheral blood and immunocytochemistry
Blood samples were obtained from 29 patients with breast cancer and 10 healthy volunteers who were enrolled between December 2016 and March 2020. Informed signed consent was obtained from patients and healthy volunteers before blood collection, and the study was approved by the ethics committees of the Saitama Cancer Center (H26-423, H29-669) and Saitama University (H26-21, R1-E-2). All methods were performed in accordance with relevant guidelines and regulations of the Saitama University and the Saitama Cancer Center. Blood samples were collected in sterile EDTA-coated vacutainer tubes.
CTCs in blood were enumerated using ClearCell FX (Clearbridge BioMedics, Pte Ltd., Singapore) according to the manufacturer’s instruction27. After treatment with a red blood cell lysis buffer, the nucleated cell fraction from 7.5 mL blood was loaded on the CTChip. The enriched CTCs were collected via centrifugation at 500 × g for 15 min at room temperature. This enumeration was conducted twice per individual, and the total CTCs-enriched fraction was divided into four samples to analyse SET-, epithelial-, and mesenchymal-CTCs using immunocytochemical staining.
The CTCs-enriched fractions were fixed with 4% paraformaldehyde in phosphate buffered saline (PBS) containing 0.1% Triton X-100 for 10 min. Immunocytochemical staining was conducted after blocking with 5% FBS and 5% human FcR blocking reagent (MiltenyiBiotec GmbH) in PBS for 30 min. For SET-CTCs analysis, the CTCs were incubated with anti-SET and anti-CD45 antibodies for 30 min, and then stained with appropriate fluorescently labelled secondary antibodies, along with 4′6-diamidino-2-phenylindole (DAPI). SET-positive (green), CD45-negative, and DAPI-positive cells more than 10 μm in diameter were designated as SET-CTCs. For epithelial-CTCs, an anti-cytokeratin and EpCAM antibody cocktail including anti-pan cytokeratin, anti-cytokeratin (C-36H5), anti-EpCAM, and anti-CD45 antibodies were used. For mesenchymal-CTCs, anti-vimentin and anti-GCDFP 15 antibodies were used, and vimentin-positive (green) and GCDFP-15-positive (red) cells were designated as mesenchymal-CTCs. The size and staining intensity of CTCs were analysed using Hybrid Cell Count (BZ-H3A) under an All-in-One fluorescence microscope BZ-X700 (Keyence Co., Osaka, Japan).
Statistical analysis
Statistical analyses for in vitro and in vivo studies were performed using one-way analysis of variance (ANOVA), followed by Dunnett’s test. Statistical tests in CTCs analysis were performed using SPSS ver. 26 (SPSS Inc. USA). Each experiment was conducted independently at least thrice, and values are expressed as mean ± standard deviation (SD). The association between CTC numbers and clinical parameters was assessed using Student’s t-test. P < 0.05 was considered statistically significant.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Nagini, S. Breast cancer: Current molecular therapeutic targets and new players. Anticancer Agents Med. Chem. 17, 152–163 (2017).
Gao, Y. et al. Clinical significance of pancreatic circulating tumor cells using combined negative enrichment and immunostaining-fluorescence in situ hybridization. J. Exp. Clin. Cancer Res. 35, 66. https://doi.org/10.1186/s13046-016-0340-0 (2016).
Boral, D. et al. Molecular characterization of breast cancer CTCs associated with brain metastasis. Nat. Commun. 8, 196. https://doi.org/10.1038/s41467-017-00196-1 (2017).
Riethdorf, S. et al. Clinical applications of the CellSearch platform in cancer patients. Adv. Drug Deliv. Rev. 125, 102–121. https://doi.org/10.1016/j.addr.2018.01.011 (2018).
Mego, M. et al. Characterization of metastatic breast cancer patients with nondetectable circulating tumor cells. Int. J. Cancer. 129, 417–423 (2011).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Heerboth, S. et al. EMT and tumor metastasis. Clin. Transl. Med. 4, 6. https://doi.org/10.1186/s40169-015-0048-3 (2015).
von Lindern, M. et al. Can, a putative oncogene associated with myeloid leukemogenesis, may be activated by fusion of its 3′ half to different genes: Characterization of the set gene. Mol. Cell. Biol. 12, 3346–3355 (1992).
Li, M., Guo, H. & Damuni, Z. Purification and characterization of two potent heat-stable protein inhibitors of protein phosphatase 2A from bovine kidney. Biochemistry 34, 1988–1996 (1995).
Li, M., Makkinje, A. & Damuni, Z. The myeloid leukemia-associated protein SET is a potent inhibitor of protein phosphatase 2A. J. Biol. Chem. 271, 11059–11062 (1996).
Sangodkar, J. et al. All roads lead to PP2A: Exploiting the therapeutic potential of this phosphatase. FEBS J. 283, 1004–1024 (2016).
Fujiki, H. et al. The concept of the okadaic acid class of tumor promoters is revived in endogenous protein inhibitors of protein phosphatase 2A, SET and CIP2A, in human cancers. J. Cancer Res. Clin. Oncol. 144, 2339–2349 (2018).
Weinberg, R. A. The Biology of Cancer 2nd edn, 476 (Garland Science, Taylor & Francis Group, 2014).
Westermarck, J. & Neel, B. G. Piecing together a broken tumor suppressor phosphatase for cancer therapy. Cell 181, 514–517 (2020).
Janghorban, M. et al. Targeting c-MYC by antagonizing PP2A inhibitors in breast cancer. Proc. Natl. Acad. Sci. U. S. A. 111, 9157–9162 (2014).
Cristobal, I. et al. Deregulation of the PP2A inhibitor SET shows promising therapeutic implications and determines poor clinical outcome in patients with metastatic colorectal cancer. Clin. Cancer Res. 21, 347–356 (2015).
Farrell, A. S. et al. Targeting inhibitors of the tumor suppressor PP2A for the treatment of pancreatic cancer. Mol. Cancer Res. 12, 924–939 (2014).
Christensen, D. J. et al. SET oncoprotein overexpression in B-cell chronic lymphocytic leukemia and non-Hodgkin lymphoma: A predictor of aggressive disease and a new treatment target. Blood 118, 4150–4158 (2011).
Liu, H. et al. Overexpression of PP2A inhibitor SET oncoprotein is associated with tumor progression and poor prognosis in human non-small cell lung cancer. Oncotarget 6, 14913–14925. https://doi.org/10.18632/oncotarget.3818 (2015).
Yuan, X. et al. Overexpression of SET oncoprotein is associated with tumor progression and poor prognosis in human gastric cancer. Oncol. Rep. 38, 1733–1741 (2017).
Cristobal, I. et al. Overexpression of SET is a recurrent event associated with poor outcome and contributes to protein phosphatase 2A inhibition in acute myeloid leukemia. Haematologica 97, 543–550. https://doi.org/10.3324/haematol.2911.050542 (2012).
Bayarkhangai, B. et al. A comprehensive and perspective view of oncoprotein SET in cancer. Cancer Med. 7, 3084–3094 (2018).
Soo Hoo, L., Zhang, J. Y. & Chan, E. K. Cloning and characterization of a novel 90 kDa ‘companion’ auto-antigen of p62 overexpressed in cancer. Oncogene 21, 5006–5015 (2002).
Oaks, J. & Ogretmen, B. Regulation of PP2A by sphingolipid metabolism and signaling. Front. Oncol. 4, 388. https://doi.org/10.3389/fonc.2014.00388 (2015).
Kauko, O. et al. Phosphoproteome and drug-response effects mediated by the three protein phosphatase 2A inhibitor proteins CIP2A, SET, and PME-1. J. Biol. Chem. 295, 4194–4211 (2020).
Liu, C. Y. et al. Targeting SET to restore PP2A activity disrupts an oncogenic CIP2A-feedforward loop and impairs triple negative breast cancer progression. EBioMedicine 40, 263–275. https://doi.org/10.1016/j.ebiom.2018.12.032 (2019).
Lee, Y., Guan, G. & Bhagat, A. A. ClearCell® FX, a label-free microfluidics technology for enrichment of viable circulating tumor cells. Cytometry A 93, 1251–1254. https://doi.org/10.1002/cyto.a.23507 (2018).
Ramalingam, N. et al. Label-free enrichment and integrated full-length mRNA transcriptome analysis of single live circulating tumor cells from breast cancer patients. Cancer Res. 77(13 Supplement), 2923 (2019).
Huang, Y. H. et al. SET overexpression is associated with worse recurrence-free survival in patients with primary breast cancer receiving adjuvant tamoxifen treatment. J. Clin. Med. 7, 245. https://doi.org/10.3390/jcm7120515 (2018).
Cristobal, I. et al. Deregulation of SET is associated with tumor progression and predicts adverse outcome in patients with early-stage colorectal cancer. J. Clin. Med. 8, 346. https://doi.org/10.3390/jcm8101517 (2019).
Sharma, S. et al. Circulating tumor cell isolation, culture, and downstream molecular analysis. Biotechnol. Adv. 36, 1063–1078. https://doi.org/10.1016/j.biotechadv.2018.03.007 (2018).
Ito, M. et al. Impact of circulating tumour cells on survival of eribulin-treated patients with metastatic breast cancer. Med. Oncol. 36, 89. https://doi.org/10.1007/s12032-019-1314-9 (2019).
Switzer, C. H. et al. Targeting SET/I2PP2A oncoprotein functions as a multi-pathway strategy for cancer therapy. Oncogene 30, 2504–2513 (2011).
Enjoji, S. et al. Stemness is enhanced in gastric cancer by a SET/PP2A/E2F1 axis. Mol. Cancer Res. 16, 554–563 (2018).
Li, J. et al. Stable SET knockdown in breast cell carcinoma inhibits cell migration and invasion. Biochem. Biophys. Res. Commun. 453, 7–12 (2014).
Cristobal, I. et al. PP2A inhibition is a common event in colorectal cancer and its restoration using FTY720 shows promising therapeutic potential. Mol. Cancer Ther. 13, 938–947. https://doi.org/10.1158/1535-7163.MCT-13-0150 (2014).
Neviani, P. & Perrotti, D. SETting OP449 into the PP2A-activating drug family. Clin. Cancer Res. 20, 2026–2028 (2014).
Agarwal, A. et al. Antagonism of SET using OP449 enhances the efficacy of tyrosine kinase inhibitors and overcomes drug resistance in myeloid leukemia. Clin. Cancer Res. 20, 2092–2103 (2014).
Devanand, P. et al. Inhibition of TNFα-interacting protein α (Tipα)-associated gastric carcinogenesis by BTG2/TIS21 via downregulating cytoplasmic nucleolin expression. Exp. Mol. Med. 50, e449. https://doi.org/10.1038/emm.2017.281 (2018).
Iida, K. et al. Cell softening in malignant progression of human lung cancer cells by activation of receptor tyrosine kinase AXL. Sci. Rep. 7, 17770. https://doi.org/10.1038/s41598-017-18120-4 (2017).
Namiki, K. et al. (−)-Epigallocatechin gallate inhibits stemness and tumourigenicity stimulated by AXL receptor tyrosine kinase in human lung cancer cells. Sci. Rep. 10, 2444. https://doi.org/10.1038/s41598-020-59281-z (2020).
Acknowledgements
We thank all patients and healthy volunteers for providing blood samples; Prof. Hirota Fujiki, the Faculty of Medicine, Saga University for contribution to our concept and for a stimulating discussion; Dr. Tatsuro Watanabe, the Faculty of Medicine, Saga University for fruitful discussion; Dr. Yuri Yamaguchi, Research Institute of Clinical Oncology, Saitama Cancer Center for providing breast cancer cell lines, and Mrs. Kaori Suzuki and Mr. Nobuyoshi Matsuzawa of Saitama University for their technical assistance. We would like to thank Editage for English language editing. This work was supported by a Grant from the Japan Society for the Promotion of Science (JSPS) KAKENHI (116k10487 to S.E.N.), Eisai Co. Ltd (to M.S.) and Takeda Science Foundation (to M.S.).
Author information
Authors and Affiliations
Contributions
K.T., S.E.N., and M.S. conceived the experiments, K.T., P.W., Y.K., M.K., and M.S. conducted the experiments, and analysed the results, K.T., S.E.N. and M.S. wrote the manuscript. K.K., K.T., K.I., Y.S., and H.M. reviewed and edited the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tozuka, K., Wongsirisin, P., Nagai, S.E. et al. Deregulation of protein phosphatase 2A inhibitor SET is associated with malignant progression in breast cancer. Sci Rep 11, 14238 (2021). https://doi.org/10.1038/s41598-021-93620-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93620-y
This article is cited by
-
Cancer progression by the okadaic acid class of tumor promoters and endogenous protein inhibitors of PP2A, SET and CIP2A
Journal of Cancer Research and Clinical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.