Abstract
Neck contrast-enhanced CT (CECT) is a routine tool used to evaluate patients with cervical lymphadenopathy. This study aimed to evaluate the ability of convolutional neural networks (CNNs) to classify Kikuchi-Fujimoto’s disease (KD) and cervical tuberculous lymphadenitis (CTL) on neck CECT in patients with benign cervical lymphadenopathy. A retrospective analysis of consecutive patients with biopsy-confirmed KD and CTL in a single center, from January 2012 to June 2020 was performed. This study included 198 patients of whom 125 patients (mean age, 25.1 years ± 8.7, 31 men) had KD and 73 patients (mean age, 41.0 years ± 16.8, 34 men) had CTL. A neuroradiologist manually labelled the enlarged lymph nodes on the CECT images. Using these labels as the reference standard, a CNNs was developed to classify the findings as KD or CTL. The CT images were divided into training (70%), validation (10%), and test (20%) subsets. As a supervised augmentation method, the Cut&Remain method was applied to improve performance. The best area under the receiver operating characteristic curve for classifying KD from CTL for the test set was 0.91. This study shows that the differentiation of KD from CTL on neck CECT using a CNNs is feasible with high diagnostic performance.
Similar content being viewed by others
Introduction
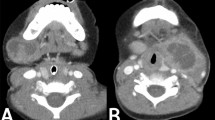
In benign cervical lymphadenopathy, both Kikuchi-Fujimoto disease (KD) and cervical tuberculous lymphadenitis (CTL) can show enlarged cervical lymph nodes with or without necrosis, with similar imaging features on contrast-enhanced CT (CECT)1,2,3,4,5. Although both are benign, there are differences in their treatment and course6,7,8,9,10,11. Therefore, many studies have been performed to differentiate KD from CTL using CECT1,2,3,12. KD can show perinodal infiltration, indistinct margins of necrosis, stronger cortical enhancement of lymph nodes, and unilaterality1,2,12,13. CTLs can demonstrate a lower density of necrosis, unilocular necrosis, calcifications, and skin fistula on CECT4,9. However, the differential diagnosis using imaging alone is not easy. The condition needs to be confirmed by histopathologic diagnosis using fine needle aspiration, core needle biopsy, or excision13,14,15,16. However, because these histopathologic confirmations are invasive, additional information from convolutional neural networks (CNNs) could be useful to enhance conspicuity for deciding the diagnosis.
Deep learning methods with CNNs utilize multiple layered neural networks to develop robust predictive models without feature selection by human image evaluation experts17,18,19,20. In radiology, many studies using CNNs have focused on the detection or classification of lesions and the validation of the deep learning technique performance20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35. The performance of CNNs has been improved and found to be comparable to that of radiologists in many studies32,33,34,35,36. However, the deep learning application to neck CT could have many challenges in the detection and classification of cervical lymphadenopathy, because neck CT has many anatomic structures from skull base to thoracic inlet and CT has inferior soft tissue contrast to MR37,38,39,40,41. Accordingly, for the enhancement of performance, this study used the Cut&Remain method40, which is one of supervised augmentation methods and crops the image to focus on the lesion.
Recently, in head and neck imaging, deep learning methods have been used to differentiate metastatic lymph nodes in thyroid cancer, discriminate benign and malignant thyroid nodules, detect extracapsular extension of metastatic lymph nodes in head and neck cancers, and automatic lymph nodes segmentation20,24,34,35,36,41. However, no studies have investigated the feasibility of deep learning applications to classify of benign cervical lymph nodes. In this study, a deep learning method was developed to discriminate KD from CTL on CECT. The purpose of this study was to evaluate the ability of CNNs to differentiate benign cervical lymphadenopathy and classify KD and CTL in patients with benign cervical lymphadenopathies.
Results
Patient characteristics
Among the study cohort, KD occurred more frequently in women (75.2%) and the patients’ age of KD was significantly younger (mean age, 25.1 ± 8.7 years) than CTL (mean age, 41.0 ± 16.8 years) (p = 0.002 and < 0.001, respectively). The most common symptom was a palpable cervical mass, which was observed in 99.2% of patients with KD (124 of 125 patients) and 97.3% of patients with CTL (71 of 73 patients). Fever was more frequently observed in patients with KD (60.0%, 75 of 125 patients) than in those with CTL (5.5%, 4 of 73 patients). There was no significant difference in cervical lymph node enlargement unilaterality (p = 0.070) [95.2% for KD (119 of 125 patients) and 74.0% for CTL (54 of 73 patients)]. The abbreviations of the study are listed in Table 1. The detailed data of the study population are summarized in Table 2.
Diagnostic performance for classification
The diagnostic performance of the deep learning models is shown in Table 3. In the preprocessing, the CT images were divided into three groups, as follows: the original image with aspect ratios of 1.0, 1.5, and 2.0, the original image with an aspect ratio of 3.0, and the original image with an aspect ratio of 4.0. The test results of each group showed accuracies of 69.15%, 94.67%, and 86.05%, respectively. The aspect ratio of 3.0 setting showed an accuracy of 94.67%, a sensitivity of 99.52%, a specificity of 73.90%, a positive predictive value of 94.22%, and a negative predictive value of 97.32%, representing the best diagnostic performance. For the test set (1266 slices), the area under the receiver operating characteristic curve (AUC) of CNNs was 0.91 with an aspect ratio of 3.0, followed by 0.87 with an aspect ratio of 4.0 (Fig. 1).
Qualitative evaluation by Grad-CAM
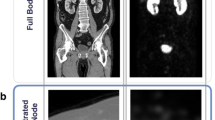
An augmentation technique was developed, where labeled lesions were mainly considered as cues for classification, with attention-guided networks39. To verify that the technique was indeed learning to recognize the lesions in target images, the activation maps for test images were visually shown. The vanilla ResNet-50 model was used to obtain Grad-CAM to observe the effect of the augmentation method clearly. Figure 2 shows the test examples and the corresponding Grad-CAM according to the aspect ratio. For aspect ratios of 3.0 and 4.0, Grad-CAM indicated enlarged lymph nodes in the right level IV and supraclavicular fossa. Grad-CAM analysis indicated that Cut&Remain force a model to focus on lesions, irrespective of the background.
Representative attention guide with CAM images in each Kikuchi-Fujimoto disease and cervical tuberculous lymphadenitis group. This figure shows test examples as well as the corresponding Grad-CAM according to the aspect ratio. Ground-truth annotation are shown with a red box. In case of aspect ratios (b) 3.0 and (c) 4.0, the Grad-CAM results indicate that the trained model identify the enlarged lymph nodes in right level IV and supraclavicular fossa.
Discussion
This study developed a deep learning algorithm for classifying KD and CTL in patients with benign cervical lymphadenopathy using neck CECT images. With the application of a bounding box, the deep learning algorithm exhibited remarkable performance in classifying KD and CTL. The results indicated that a CNNs could accurately distinguish KD from CTL with an AUC of 0.91 in the test dataset. Therefore, we expect the application of CNNs to be feasible for classifying cervical lymphadenopathy.
Recently, deep learning techniques have been validated for head and neck oncology imaging26,34,35,36. A prior study applied a deep learning method to diagnose metastatic lymph nodes and identify extracapsular extension in head and neck cancer, with an AUC of 0.91. Another study showed the high performance of deep learning CNNs, with an AUC of 0.95 for diagnosing metastatic lymph nodes in patients with thyroid cancer on CECT. This study result also showed a high performance, with an AUC of 0.91 for classifying KD and CTL on CECT. This study result was enhanced when the bounding boxes were applied using the Cut&Remain method. We hypothesize that dedicated comparisons between KD and CTL will be possible when expert experience and supervision are added.
Among benign cervical lymphadenopathies, KD and CTL are the major differential diagnoses in patients with acute cervical lymphadenopathy3. In prior studies with CT, the presence of indistinct margins of necrotic foci was found to be an independent predictor of KD, with 80% accuracy1. Another previous study demonstrated that bilateral involvement, ≥ five levels of nodal involvement, absence or minimal nodal necrosis, marked perinodal infiltration, absence of upper lung lesion, and mediastinal lymphadenopathy were independent findings that suggest KD rather than tuberculosis on CT. The investigators reported an AUC of 0.761 for these five CT findings, which was considerably lower than this result2. We suggest that combined CT findings and simultaneous deep learning could enhance diagnostic performance for benign cervical lymphadenopathy.
There have been many studies on cervical lymph node analysis using deep learning, particularly in oncology34,36,39. However, no previous investigations have evaluated the application of a deep-learning model to discriminate benign cervical lymphadenopathy. Notably, Adele discriminated between normal lymph nodes and lymphadenopathy; however, detailed lymph node disease classification was not covered41. In this study, the CNNs algorithm has several advantages. First, it shows higher performance than previous qualitative analysis studies. The performance could be related to the application of the Cut&Remain method, which is a simple and efficient supervised augmentation method. This method drives a model to focus on relevant subtle and small regions; therefore, it is possible to differentiate such small lesions from medical images and enhance performance. This method has been used to classify clavicle and femur fractures on radiography and 14 lesion classifications on chest radiographs. Previous studies have shown improved performance with the application of Cut&Remain40; this study demonstrated the best performance with the Cut&Remain method as well. We believe that as there are many anatomic structures on neck CECT, this method could be effective in future studies using deep learning applications for neck CT imaging.
This study had several limitations. First, the subject number of CTL was smaller than that of KD. Accordingly, to overcome this problem, data augmentation was performed on CTL. Second, it is important to differentiate malignant from benign lymph nodes, and this study aimed to differentiate and classify benign cervical lymph nodes only. Further studies are needed to distinguish metastatic from benign lymph nodes. Third, an external validation was not performed, and the deep learning algorithm may exhibit overfitting results. Finally, we did not evaluate radiologist performance and compared radiologist performance with that of the deep learning method. A study that investigates this will be further conducted in the future.
In conclusion, the deep learning method is helpful in the differentiation of benign cervical lymphadenopathy between KD and CTL. The performance of CNNs method can be enhanced with the Cut&Remain method. In the future, deep learning diagnostic algorithm could be developed for differentiation of cervical lymphadenopathy including benign and malignant lymph nodes with large number of patients.
Methods
This study was approved by the institutional review board (IRB) of Hanyang University Seoul Hospital (2020-07-048-004). The requirement of informed consent was waived due to its retrospective nature by the institutional review board (IRB) of Hanyang University Seoul Hospital. All experiments were performed in accordance with relevant guidelines and regulations.
Patients and datasets
We retrospectively investigated the medical records of 350 patients who were clinically diagnosed with KD and CTL from January 2012 to June 2020 at a tertiary hospital (193 KD, 157 CTL). Patients were excluded if they did not undergo pathologic confirmation (n = 56), lacked a neck CECT (n = 27), had incomplete clinical data (n = 32), or had a poor quality CECT due to metal dental artifacts (n = 3). Finally, 125 patients with KD and 73 with CTL were included. A flowchart of patient selection is shown in Fig. 3. All scans of neck CECT were downloaded from PACS in the DICOM format. A neuroradiologist and an otolaryngology resident reviewed the images and CT scans demonstrating only enlarged cervical lymph nodes were included. Finally, patients with KD consisted of 6306 slices, and those for CTL consisted of 1477 slices. The training datasets comprised training (4414 slices for KD and 1034 slices for CTL) and validation (631 slices for KD and 148 slices for CTL) slices. The test datasets were composed of 1261 slices for the KD group and 295 for the CTL group.
CT imaging protocol
Contrast-enhanced CT imaging was performed after the administration of intravenous iodine contrast agent (1.2 mL/kg, 2 mL/s, 30 s delay) using 120 kVp, 200 mAs, and 2 mm slice thickness reconstruction (Brilliance 64, Philips Healthcare, Best, The Netherlands; SOMATOM, Definition Flash, Siemens Healthcare, Erlangen, Germany).
Labeling
Figure 4 shows a schematic representation of the pipeline for the differentiation of KD from CTL. A radiologist (J.Y.L, a neuroradiologist with nine years of experience) manually identified cervical lymph node lesions on the CT images and then drew a rectangular bounding box on the cervical lymph nodes. The image was cropped based on the bounding box-labeled image. The labeled image was used as an augmentation technique to generate a new training sample.
Data augmentation technique to identify local key features
A novel data augmentation technique, called Cut&Remain, was applied to eliminate unimportant portions of the image while preserving the spatial location of the important areas40.
The CNNs assumed that \(x\in {\mathbb{R}}^{W\times H\times C}\) and \(y\) denote the training image and label, respectively. The goal of Cut&Remain technique is to generate a new training sample \((\widetilde{x},\widetilde{y})\), which can be described by Eq. (1) and used to train the model based on its original loss function.
where \(\mathrm{M}\in {\{\mathrm{0,1}\}}^{W\times H}\) denotes a binary mask indicating lesion and \(\odot\) is element-wise multiplication. To generate mask, \(M\), a bounding box annotation \(\mathrm{B}=({c}_{x},{c}_{x},\mathrm{ w},\mathrm{ h})\), was used to indicate the cropped region on image \(x\). For the bounding boxes, nine aspect ratios of {2.0, 2.5, 3.0} were considered. Within the cropped region, the value of binary mask \(\mathrm{M}\in {\{\mathrm{0,1}\}}^{W\times H}\) was 1 within B or 0 outside B. In each training step, an augmented sample \((\widetilde{x},\widetilde{y})\) was generated based on each training sample according to Eq. (1) and included in the mini-batch, as shown in Fig. 5.
Data augmentation was performed with rotation (− 10° to 10°), flip (horizontal 50%), zoom (95–105%), and translation shift (0–10% of the image size in the horizontal and vertical axes), which was applied before Cut&Remain augmentation with corresponding bounding boxes.
Deep learning training strategy
ResNet-50 was adapted as a backbone network and we have initialized the weights randomly. We used binary cross-entropy loss for classification and Adam with a momentum of 0.9. The initial learning rate was set to 0.0005. The model was trained for 2000 epochs in total, and the learning rate was reduced by a factor of 10 at 1000 epochs.
Statistical analysis
The sensitivity, specificity, and AUC were evaluated to assess the diagnostic performance of the deep learning algorithm. We report the AUC and average F1-score of five random runs with different initializations for the classification performance.
Data availability
The datasets generated for the current study are available from the corresponding author on reasonable request.
Code availability
The codes generated for the current study are available from the corresponding author on reasonable request.
References
Lee, S., Yoo, J. H. & Lee, S. W. Kikuchi disease: Differentiation from tuberculous lymphadenitis based on patterns of nodal necrosis on CT. AJNR Am. J. Neuroradiol. 33, 135–140. https://doi.org/10.3174/ajnr.A2724 (2012).
Baek, H. J., Lee, J. H., Lim, H. K., Lee, H. Y. & Baek, J. H. Diagnostic accuracy of the clinical and CT findings for differentiating Kikuchi’s disease and tuberculous lymphadenitis presenting with cervical lymphadenopathy. Jpn. J. Radiol. 32, 637–643. https://doi.org/10.1007/s11604-014-0357-2 (2014).
You, S. H., Kim, B., Yang, K. S. & Kim, B. K. Cervical necrotic lymphadenopathy: A diagnostic tree analysis model based on CT and clinical findings. Eur. Radiol. 29, 5635–5645. https://doi.org/10.1007/s00330-019-06155-2 (2019).
Reede, D. L. & Bergeron, R. Cervical tuberculous adenitis: CT manifestations. Radiology 154, 701–704 (1985).
Na, D. G. et al. Kikuchi disease: CT and MR findings. Am. J. Neuroradiol. 18, 1729–1732 (1997).
Golden, M. P. & Vikram, H. R. Extrapulmonary tuberculosis: An overview. Am. Fam. Physician 72, 1761–1768 (2005).
Polesky, A., Grove, W. & Bhatia, G. Peripheral tuberculous lymphadenitis: Epidemiology, diagnosis, treatment, and outcome. Medicine (Baltimore) 84, 350–362. https://doi.org/10.1097/01.md.0000189090.52626.7a (2005).
Bosch, X. & Guilabert, A. Kikuchi-Fujimoto disease. Orphanet. J. Rare Dis. 1, 18. https://doi.org/10.1186/1750-1172-1-18 (2006).
Fontanilla, J. M., Barnes, A. & von Reyn, C. F. Current diagnosis and management of peripheral tuberculous lymphadenitis. Clin. Infect. Dis. 53, 555–562. https://doi.org/10.1093/cid/cir454 (2011).
Dumas, G. et al. Kikuchi-Fujimoto disease: Retrospective study of 91 cases and review of the literature. Medicine (Baltimore) 93, 372–382. https://doi.org/10.1097/MD.0000000000000220 (2014).
WHO. Global Tuberculosis Report 2020 (WHO, 2020).
Shim, E. J., Lee, K. M., Kim, E. J., Kim, H. G. & Jang, J. H. CT pattern analysis of necrotizing and nonnecrotizing lymph nodes in Kikuchi disease. PLoS ONE 12, e0181169. https://doi.org/10.1371/journal.pone.0181169 (2017).
Park, S. G. et al. Efficacy of ultrasound-guided needle biopsy in the diagnosis of Kikuchi-Fujimoto disease. Laryngoscope 131, E1519–E1523. https://doi.org/10.1002/lary.29160 (2021).
Baek, C. H., Kim, S. I., Ko, Y. H. & Chu, K. C. Polymerase chain reaction detection of Mycobacterium tuberculosis from fine-needle aspirate for the diagnosis of cervical tuberculous lymphadenitis. Laryngoscope 110, 30–34. https://doi.org/10.1097/00005537-200001000-00006 (2000).
Ryoo, I., Suh, S., Lee, Y. H., Seo, H. S. & Seol, H. Y. Comparison of ultrasonographic findings of biopsy-proven tuberculous lymphadenitis and Kikuchi disease. Korean J. Radiol. 16, 767–775. https://doi.org/10.3348/kjr.2015.16.4.767 (2015).
Han, F. et al. Efficacy of ultrasound-guided core needle biopsy in cervical lymphadenopathy: A retrospective study of 6,695 cases. Eur. Radiol. 28, 1809–1817. https://doi.org/10.1007/s00330-017-5116-1 (2018).
Soffer, S. et al. Convolutional neural networks for radiologic images: A radiologist’s guide. Radiology 290, 590–606. https://doi.org/10.1148/radiol.2018180547 (2019).
Zhou, S. K. et al. A Review of deep learning in medical imaging: Imaging traits, technology trends, case studies with progress highlights, and future promises. Proc. IEEE 109, 820–838. https://doi.org/10.1109/jproc.2021.3054390 (2021).
Zaharchuk, G., Gong, E., Wintermark, M., Rubin, D. & Langlotz, C. P. Deep learning in neuroradiology. AJNR Am. J. Neuroradiol. 39, 1776–1784. https://doi.org/10.3174/ajnr.A5543 (2018).
Kann, B. H. et al. Pretreatment identification of head and neck cancer nodal metastasis and extranodal extension using deep learning neural networks. Sci. Rep. 8, 1–11. https://doi.org/10.1038/s41598-018-32441-y (2018).
Aggarwal, R. et al. Diagnostic accuracy of deep learning in medical imaging: A systematic review and meta-analysis. NPJ Dig. Med. 4, 1–23. https://doi.org/10.1038/s41746-021-00438-z (2021).
Laith, A. et al. Robust application of new deep learning tools: An experimental study in medical imaging. Multimed. Tools Appl. 81, 13289–13317. https://doi.org/10.1007/s11042-021-10942-9 (2022).
Sibille, L. et al. 18F-FDG PET/CT uptake classification in lymphoma and lung cancer by using deep convolutional neural networks. Radiology 294, 445–452. https://doi.org/10.1148/radiol.2019191114 (2020).
Lee, J. H., Ha, E. J. & Kim, J. H. Application of deep learning to the diagnosis of cervical lymph node metastasis from thyroid cancer with CT. Eur. Radiol. 29, 5452–5457. https://doi.org/10.1007/s00330-019-06098-8 (2019).
Yang, S. et al. Deep learning segmentation of major vessels in X-ray coronary angiography. Sci. Rep. 9, 16897. https://doi.org/10.1038/s41598-019-53254-7 (2019).
Crowson, M. G. et al. A contemporary review of machine learning in otolaryngology-head and neck surgery. Laryngoscope 130, 45–51. https://doi.org/10.1002/lary.27850 (2020).
Han, C. et al. In Neural Approaches to Dynamics of Signal Exchanges (eds Esposito, A. et al.) 291–303 (Springer, 2020).
Lee, C. et al. Classification of femur fracture in pelvic X-ray images using meta-learned deep neural network. Sci. Rep. 10, 13694. https://doi.org/10.1038/s41598-020-70660-4 (2020).
Lee, J. Y., Kim, J. S., Kim, T. Y. & Kim, Y. S. Detection and classification of intracranial haemorrhage on CT images using a novel deep-learning algorithm. Sci. Rep. 10, 20546. https://doi.org/10.1038/s41598-020-77441-z (2020).
Rahman, T. et al. Reliable tuberculosis detection using chest X-ray with deep learning, segmentation and visualization. IEEE Access 8, 191586–191601. https://doi.org/10.1109/access.2020.3031384 (2020).
Kim, M., Kim, J. S., Lee, C. & Kang, B. K. Detection of pneumoperitoneum in the abdominal radiograph images using artificial neural networks. Eur. J. Radiol. Open 8, 100316. https://doi.org/10.1016/j.ejro.2020.100316 (2021).
Kim, Y. et al. Deep learning in diagnosis of maxillary sinusitis using conventional radiography. Invest. Radiol. 54, 7–15. https://doi.org/10.1097/RLI.0000000000000503 (2019).
Ardakani, A. A., Kanafi, A. R., Acharya, U. R., Khadem, N. & Mohammadi, A. Application of deep learning technique to manage COVID-19 in routine clinical practice using CT images: Results of 10 convolutional neural networks. Comput. Biol. Med. 121, 103795. https://doi.org/10.1016/j.compbiomed.2020.103795 (2020).
Onoue, K., Fujima, N., Andreu-Arasa, V. C., Setty, B. N. & Sakai, O. Cystic cervical lymph nodes of papillary thyroid carcinoma, tuberculosis and human papillomavirus positive oropharyngeal squamous cell carcinoma: Utility of deep learning in their differentiation on CT. Am. J. Otolaryngol. 42, 103026. https://doi.org/10.1016/j.amjoto.2021.103026 (2021).
Zhou, H. et al. Differential diagnosis of benign and malignant thyroid nodules using deep learning radiomics of thyroid ultrasound images. Eur. J. Radiol. 127, 108992. https://doi.org/10.1016/j.ejrad.2020.108992 (2020).
Tomita, H. et al. Deep learning for the preoperative diagnosis of metastatic cervical lymph nodes on contrast-enhanced computed tomography in patients with oral squamous cell carcinoma. Cancers 13, 600. https://doi.org/10.3390/cancers13040600 (2021).
Huff, D. T., Weisman, A. J. & Jeraj, R. Interpretation and visualization techniques for deep learning models in medical imaging. Phys. Med. Biol. 66, 04TR01. https://doi.org/10.1088/1361-6560/abcd17 (2021).
Madhavan, M. V. et al. Res-CovNet: An internet of medical health things driven COVID-19 framework using transfer learning. Neural Comput. Appl. https://doi.org/10.1007/s00521-021-06171-8 (2021).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vis. 128, 336–359. https://doi.org/10.1007/s11263-019-01228-7 (2019).
Lee, C., Kim, Y., Lee, B. G., Kim, D. & Jang, J. Look at here: Utilizing supervision to attend subtle key regions. arXiv. https://doi.org/10.48550/arXiv.2111.13233 (2021).
Courot, A. et al. Automatic cervical lymphadenopathy segmentation from CT data using deep learning. Diagn. Interv. Imaging https://doi.org/10.1016/j.diii.2021.04.009 (2021).
Author information
Authors and Affiliations
Contributions
Study concept and design: J.Y.L. and K.T.; data acquisition: B.H.K., C.L., and J.Y.L.; computer software programming, image processing, and convolutional neural networks execution: C.L.; data analysis and interpretation: B.H.K., C.L., and J.Y.L.; manuscript drafting: B.H.K., C.L., and J.Y.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, B.H., Lee, C., Lee, J.Y. et al. Initial experience of a deep learning application for the differentiation of Kikuchi-Fujimoto’s disease from tuberculous lymphadenitis on neck CECT. Sci Rep 12, 14184 (2022). https://doi.org/10.1038/s41598-022-18535-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18535-8
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.