Abstract
Transgene insertion plays an important role in gene therapy and in biological studies. Transposon-based systems that integrate transgenes by transposase-catalyzed “cut-and-paste” mechanism have emerged as an attractive system for transgenesis. Hyperactive piggyBac transposon is particularly promising due to its ability to integrate large transgenes with high efficiency. However, prolonged expression of transposase can become a potential source of genotoxic effects due to uncontrolled transposition of the integrated transgene from one chromosomal locus to another. In this study we propose a vector design to decrease post-transposition expression of transposase and to eliminate the cells that have residual transposase expression. We design a single plasmid construct that combines the transposase and the transpositioning transgene element to share a single polyA sequence for termination. Consequently, the separation of the transposase element from the polyA sequence after transposition leads to its deactivation. We also co-express Herpes Simplex Virus thymidine kinase (HSV-tk) with the transposase. Therefore, cells having residual transposase expression can be eliminated by the administration of ganciclovir. We demonstrate the utility of this combination transposon system by integrating and expressing a model therapeutic gene, human coagulation Factor IX, in HEK293T cells.
Similar content being viewed by others
Introduction
Transgenesis plays a crucial role in unraveling the function of various genes in developmental processes and disease states1,2, recombinant protein generation3,4, gene therapy5,6 and reprogramming of somatic cells7,8. Viral vectors such as adenovirus, adeno-associated virus (AAV), retrovirus and lentivirus have been widely used for delivering transgenes. The non-integrative nature of adenoviral transgenesis may not be ideal for long-term gene therapy9. Immune response and limited cargo space also precludes the extensive use of adenoviral vectors. On the other hand, integrative lentiviruses and retroviruses ensure the permanency of transgene expression. However, immune response and limited cargo space is still a major drawback10,11. The possibility of the viral elements reconstituting the active, self-replicating viral form is also a concern12,13. Despite these drawbacks viral systems remain the vector of choice if efficient transgenesis is required14. On the other end of the transgene vector spectrum is plasmid-mediated transgenesis that is characterized by low efficiency15. However, nonviral vectors are generally considered to be safer than the viral options. Therefore, to fill the niche of a vector that is characterized by highly efficient transgenesis, the ability to integrate large transgenes and a superior biosafety profile, transposon-mediated delivery systems have been developed. Transposable elements are genetic elements that can move from one place in the genome to another. These naturally occurring elements have been copied to enable the movement of a transgene flanked by inverted terminal repeat sequences from a vector to the genome. Prominent among these “cut-and-paste” transposition systems are Sleeping Beauty, Tol2 and piggyBac16,17,18,19. Among them piggyBac system has generated great interest by virtue of its ability to achieve robust, highly efficient transposition even with large transgenes20,21,22,23. The successful utilization of piggyBac system to deliver reprogramming constructs for generating induced pluripotent stem cells (iPSCs) from somatic cells has also contributed to the enthusiasm24,25.
Transposition has been achieved by delivering the transposase enzyme as protein, mRNA or expression plasmid26,27,28. When delivered as plasmids, the transposase and the inverted terminal repeat-flanked transgene are usually loaded onto two separate plasmids vectors including the helper plasmid (transposase-bearing) and the donor plasmid (transgene-bearing). In a two-plasmid system, ensuring the co-delivery of both the elements is a challenge; this problem is exacerbated when delivery is less efficient, such as in vivo applications. Therefore, attempts have been made to combine these two elements together in a single plasmid for efficient co-delivery29,30. Excitement of using transposons for gene delivery has been tempered by the apprehension of heightened levels of genotoxicity and mutagenesis due to prolonged expression of transposase. This concern is based on the prospect of the transposon element continually hopping from one place in the genome to another under the influence of the continued presence of transposase31. There is also an apprehension that the lingering transposase would remove the already integrated transgenes, as not all transposons reenter the genome32. Moreover, prolonged expression can provoke an immune response to the foreign transposase that may prevent any subsequent re-administration. Therefore, it is prudent to inactivate the transposase gene once it has completed its function of transgene integration. Most studies using piggyBac have depended on the transposase-bearing plasmid to be lost through dilution following cell division. However, the cell division rates can vary widely in different tissues and hence this process of transposase dilution is not reliable. Delivering transposase in the form of plasmid also risks random integration of the transposase element into the genome33. This can happen from nonspecific, nuclease-mediated linearization of the plasmid, or from the formation of sheared linear forms during the preparation of the plasmid. These linear forms can integrate randomly into the genome, thereby perpetuating transposase expression in some cells even when most of the circular forms get diluted out due to cell division. The co-delivery of both transposase and transposon from a single plasmid has added another level of complexity to the problem of transposase integration; there might be heightened prospect of transposase integration due to the formation of transient linear forms after transposition (Figure 1A).
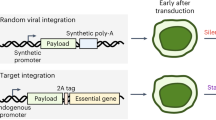
Schematic representation of potential fates of the transposase in a conventional donor-helper combination plasmid and an illustration depicting the mechanism of action of our self-inactivating combination plasmid variant.
(A) The possible fates of the transposase with the donor-helper combination system before transposition (BT) and post transposition (PT). Outcomes are shown in terms of insertional mutagenesis and duration of transposase expression. (B) Illustration depicting the self-inactivating plasmid construct Tsase-Tson(Prom-GOI-pA) and its mechanism of action. The lack of a dedicated polyA for the transposase leads to its inactivation after transposition. (hyPBase: hyperactive piggyBac transposase; Prom: promoter; pA: polyA; TRE: terminal repeat elements; GOI: Gene of interest).
A promising solution to this problem was suggested by Urschitz et al.29, in which the promoter for the transposase is included in the transposon, within the inverted terminal repeats. Therefore expression of the transposase is inactivated following excision of the transposon from the plasmid vector. However, this post-transposition self-inactivation mechanism is inefficacious when non-specific integration of the combined plasmid takes place prior to transposition34. Post transposition expression of a truncated, inactive transposase fragment, driven by the CAG promoter, is also a result of the self-inactivation process. The effect of continuous expression of the truncated transposase fragment on cellular processes has not been studied. Moreover, the CAG promoter, being in close proximity to the promoter driving the transgene, can lead to unpredictable effects on promoter function35. Compounding these problems, there is always the prospect of the CAG promoter aberrantly triggering genes in the vicinity of the integration site36. Moreover, inducible promoters or tissue-specific promoters may not be effective for driving the transgene as CAG may aberrantly trigger them due to the close proximity37. Therefore, to address these problems we have suggested an alternative solution that will make the combined plasmid system a safer and viable option for the delivery of piggyBac transposon-based systems. We propose a single plasmid construct that combines the transposase and the transpositioning transgene element to share a single polyA sequence for efficient termination. This self-inactivates the transposase element after transposition because polyA support for proper termination of the transposase transcript is lost as it is carried away along with the transpositioning transgene (Figure 1B). Additionally, we also propose to implement a HSV-tk/ganciclovir-based negative selection system in conjunction with the self-inactivation system to eliminate the rare transposase-expressing cells38.
Results
Donor-helper combination plasmid design
We first assessed the potential pitfalls of existing donor-helper (transposon-transposase combination) plasmid configuration by analyzing all the possible fates of the transposase in such a system (Figure 1A). Before transposition (BT), plasmids can either dilute out or integrate as a whole. Post transposition (PT), remaining plasmid elements (transposase) can either integrate, undergo cleavage, or recircularize and dilute out.
Our plasmid construct design suggests an approach to self-inactivate expression of the transposase after transposition (Figure 1B). This strategy is based on both the transposase and transposon (transgene) sharing a single polyA sequence for termination. A polyA sequence signal for the termination of transcription is essential for the proper expression of a gene. The shared polyA was placed distal to the transgene. Both the polyA and the transgene were flanked by the transposon terminal repeat elements (TREs). Therefore, the transposase should no longer be expressed after transposition as the polyA terminator sequence was carried away along with the transpositioning transgene.
Transposition efficiency of the combination plasmid system
We first verified the activity of the piggyBac transposon system (Supplementary Figure S1). We implemented the more efficient mutant form of piggyBac transposase called hyperactive piggyBac transposase (hyPBase) for our experiments39. HEK293T cells were transfected with two plasmids: hyPBase (Tsase-mO-pA) and the transposon (Tson(GFP-pA)). Control group was transfected with just the transposon. To distinguish transient plasmid-mediated transgene expression from permanently-integrated transposition-mediated expression, the transfected cells were passaged multiple times to dilute out the transient forms. At each passage, the percentage of cells expressing the transgene (GFP), relative to initial percentage upon transfection (passage 0), was measured using flow cytometry. Our experiments indicate that hyPBase system is functional with approximately 15% transgene integration after three passages, whereas transgene expression from the transposon-only control is rapidly lost by the second passage (~0.9%) at a split ratio of 1:10 (p<0.001). Therefore, for all the subsequent experiments we decided to limit the number of passages to three at the same split ratio of 1:10.
After verifying the efficacy of the transposase in a conventional two plasmid format we determined the efficacy of the transposase in our combination plasmid system. It was assessed using three different combination plasmid models (Figure 2A). Tsase-mO-Tson(GFP-pA) is the proposed combination plasmid design where the transposase lacks its own polyA and depends on that of the transgene (i.e. GFP) for expression. In this form, PT recircularization and integration usually cannot perpetuate transposase expression due to the absence of a dedicated polyA. Tsase-mO-Tson(GFP-pA) was compared with Tsase-mO-Tson(GFP-pA)-pA, which has a second polyA outside the 3′ terminal repeat element and Tsase-mO-pA-Tson(GFP-pA), which is the traditional combination plasmid design where both the transposase and transgene have their own dedicated polyA. In Tsase-mO-Tson(GFP-pA)-pA, the transposase expression post transposition can be driven by recircularization. In the case of Tsase-mO-pA-Tson(GFP-pA), both recircularization and integration drives PT transposase expression. BT integration in all the three forms can potentially lead to prolonged transposase expression. Plasmid dilution and PT plasmid cleavage in all the three combination plasmids leads to a decrease in transposase levels (Supplementary Figure S2).
Transposition efficiency of the combination plasmid system.
(A) Schematic design of three different types of combination plasmid constructs. Tsase-mO-Tson(GFP-pA) is the proposed self-inactivating plasmid design with a single polyA shared between the transposase and the transgene. Tsase-mO-Tson(GFP-pA)-pA has a second polyA near the transgene polyA, but outside the transposable element. Tsase-mO-pA-Tson(GFP-pA) is the traditional combination plasmid design where the transposase and transgene have their own dedicated polyA. (B) Percentage of transfected HEK293T cells with transgene (GFP) expression as measured by flow cytometry. All measurements were calculated relative to the expression at passage 0 (12 hours after transfection). All three plasmid constructs performed equally well (Two-way ANOVA p = 0.2). Two way ANOVA with Bonferroni post hoc at the third passage indicate no significant difference among the three plasmid types (p>0.05). The average transgene-positive cell fraction after three passages ranged from 15% to 19%. (C) Significant differences in GFP median fluorescence intensity (MFI) level were observed among three plasmid types (p<0.0001, two-way ANOVA). After three passages, the average MFI of Tsase-mO-Tson(GFP-pA) was found to be ~2.2 fold higher than that of Tsase-mO-Tson(GFP-pA)-pA and ~3.3 fold higher than that of Tsase-mO-pA-Tson(GFP-pA), with statistical significance of p<0.001 (two-way ANOVA with Bonferroni post hoc). There was no significant difference between Tsase-mO-Tson(GFP-pA)-pA and Tsase-mO-pA-Tson(GFP-pA) (p>0.05; two-way ANOVA with Bonferroni post hoc). (n = 4; error bars denote standard error of the mean (SEM)).
A two-way ANOVA analysis on the flow cytometry measurements showed that the nature of combination plasmids had no statistically significant effect (p = 0.2) on the percentage of transfected cells expressing GFP. All three plasmid constructs showed similar degree of transgene integration after three passages (p>0.05, P3 average: 15% to 19% (normalized to P0)) (Figure 2B and Supplementary Figure S3 A). Significant differences in GFP median fluorescence intensity (MFI) levels were observed among three plasmid types on a two-way ANOVA analysis (p<0.0001). After three passages, the average normalized MFI of Tsase-mO-Tson(GFP-pA) was 2.2 and 3.3 fold higher than that of Tsase-mO-Tson(GFP-pA)-pA and Tsase-mO-pA-Tson(GFP-pA), respectively which was found to be statistically significant on a Bonferroni post hoc test (p<0.001). There was no significant MFI difference between Tsase-mO-Tson(GFP-pA)-pA and Tsase-mO-pA-Tson(GFP-pA) (p>0.05) (Figure 2C and Supplementary Figure S3B). This implies that Tsase-mO-Tson(GFP-pA) is the most effective form for enabling high levels of transgene expression.
Comparison of the self-inactivating combination plasmid with the two plasmid system
The combination plasmid system Tsase-mO-Tson(GFP-pA) performed better than the two plasmid system in terms of percentage of cells where transposition took place and also the number of transposition events per cell as reflected by the MFI. In the two plasmid system, two separate plasmids Tsase-mO-pA and Tson(GFP-pA) were transfected together (Figure 3A). After three passages, Tsase-mO-Tson(GFP-pA) showed an average integration efficiency of ~22.3%, while Tsase-mO-pA + Tson(GFP-pA) showed only ~15% (p<0.001, normalized P0) (Figure 3B and Supplementary Figure S4). Furthermore, the normalized MFI level of Tsase-mO-Tson(GFP-pA) was significantly higher than that of Tsase-mO-pA + Tson(GFP-pA) group (p<0.0001) (Figure 3B and Supplementary Figure S4). Our results are consistent with previous studies that demonstrate more effective co-delivery of both the elements using the combined form29,30.
Comparison of the self-inactivating combination plasmid with the two plasmid system.
(A) Illustration depicting the plasmid constructs Tsase-mO-pA, Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA). The two plasmid system involves transfection of two separate plasmids, Tsase-mO-pA and Tson(GFP-pA), whereas the combination system involves transfection of a single plasmid, Tsase-mO-Tson(GFP-pA). (B) Percentage of transfected HEK293T cells with transgene (GFP) expression as measured by flow cytometry. All measurements were calculated relative to the transgene expression 12 hours after transfection (passage 0). Results show a significant difference between the transposition efficiency of combined system and that of two plasmid system (p<0.0001, two-way ANOVA). After three passages, Tsase-mO-Tson(GFP-pA) had an average transgene integration rate of ~22.3%, while the Tsase-mO-pA + Tson(GFP-pA) group had an average of ~15% (p<0.001; two-way ANOVA with Bonferroni post hoc). (C) Average GFP expression level (relative to P0) measured by calculating the median fluorescence intensity (MFI). The GFP median fluorescence intensity (MFI) level in the Tsase-mO-Tson(GFP-pA) group was significantly higher than Tsase-mO-pA + Tson(GFP-pA) (p<0.0001, two-way ANOVA). (n = 4; error bars denote SEM).
Quantification of residual transposase expression in the combination plasmid system
To determine if residual transposase expression can be reduced in our combined plasmid system, we first quantified residual transposase expression (Figure 4). The fluorescence level of mOrange, which was co-expressed with transposase from the same promoter, was measured using flow cytometry. All measurements were calculated relative to initial GFP expression after transfection (passage 0). After three passages, residual transposase (mOrange) expression in cells transfected with each of the three plasmids was less than 1%. No statistical significant difference was detected among the three samples (Figure 4A and Supplementary Figure S5A). On the other hand, the mOrange MFI (normalized to P0 mOrange MFI) of Tsase-mO-pA-Tson(GFP-pA) was significantly higher than that of Tsase-mO-Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA)-pA as early as P0 + 5 days. Similar trend was seen over the subsequent passages (P1 p value< 0.001, Bonferroni post hoc. No post hoc significance analysis was done on P2 and P3 cells due to the detection of very few mOrange+ cells). This result is consistent with increased expression of the transposase when it has its own dedicated polyA. However, the MFI between Tsase-mO-Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA)-pA was not significantly different (Figure 4B and Supplementary Figure S5B).
Quantification of residual transposase expression in the combination plasmid system.
(A) Percentage of HEK293T cells with residual transposase expression after transfection with each type of combination plasmid. mOrange fluorescence which is co-expressed with transposase, was measured by flow cytometry as a proxy to quantify transposase expression. All measurements were calculated relative to passage 0 transgene (GFP) expression. After three passages residual transposase (mOrange) expression in all three plasmids was less than 1% and not significantly different from each other (p>0.05, two-way ANOVA with Bonferroni post hoc test). (B) Median mOrange fluorescence intensity (MFI) of HEK293T cells transfected with the combination plasmids. There is a significant difference in the mOrange MFI amongst three plasmid types (p<0.0001, two-way ANOVA). From passage P0 + 5 days the MFI of Tsase-mO-pA-Tson(GFP-pA) was significantly higher than that of Tsase-mO-Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA)-pA (p<0.001 for both; two-way ANOVA with Bonferroni post hoc). However, the MFI between Tsase-mO-Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA)-pA was not significantly different (p>0.05, two-way ANOVA with Bonferroni post hoc). (n = 4; error bars denote SEM).
Strategy to eliminate residual transposase expression by employing HSV-tk/ganciclovir negative selection
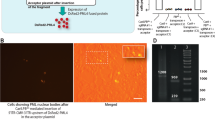
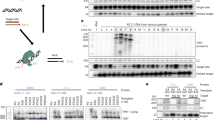
To address the problem of residual transposase expression, we designed a combination plasmid that also expresses Herpes Simplex Virus thymidine kinase (HSV-tk) in a bi-cistronic fashion along with the transposase (Figure 5A). This arrangement ensures that any cell expressing residual transposase is killed by the administration of ganciclovir (Figure 5B). There was no qualitative difference between the GFP immunofluorescence of the ganciclovir-treated cells and the no treatment group (Figure 5C). This finding was further verified quantitatively by flow cytometry (p = 0.23; student's t-test) (Figure 5D). Therefore, we can conclude that ganciclovir-mediated killing of the transposase-positive cells does not lead to a significant reduction of transgene (GFP)-positive cells. This corroborates the previous finding that residual transposase (mOrange) expression in all the three combination plasmids was very low (less than 1%) (Figure 4A). Furthermore, we placed the puromycin resistance gene after the same promoter as a fusion protein with HSV-tk. This enabled the cells that expressed residual transposase to be resistant to puromycin and sensitive to ganciclovir. Upon addition of puromycin, resistant colonies evolved, which indicated that there were cells with residual transposase expression (Figure 5C) thereby corroborating the previous finding of a residual transposase expression (Figure 4A). We also did not observe any puromycin resistant colony that was not GFP positive (Figure 5C). Upon addition of puromycin and ganciclovir not a single resistant colony emerged, thereby indicating that ganciclovir treatment was able to kill all the residual transposase expressing cells (Figure 5C). The absence of transposase expression upon ganciclovir treatment as detected by the sensitive RT-PCR assay is a strong evidence of complete removal of residual transposase by ganciclovir treatment (Figure 5E). The transfection of the combination plasmid along with puromycin and HSV-tk/ganciclovir treatment significantly altered the viability of cells as measured by MTT assay (p<0.001). Further analysis of the MTT assay data showed a significant difference between the transfected cells treated with only puromycin and those treated with both puromycin and ganciclovir (p<0.001) thereby reconfirming that ganciclovir treatment is functional (Figure 5F). Thus, our results demonstrate that HSV-tk/ganciclovir selection is an efficient way to get rid of residual transposase expressing cells in case the self-inactivation mechanism of the combination plasmid fails.
Strategy to eliminate residual transposase expression by employing HSV-tk/ganciclovir negative selection.
(A) Illustration depicting the plasmid construct Tsase-puro dTK-Tson(GFP-pA). (B) Cartoon illustrating the usage of HSV-tk/ganciclovir- mediated negative selection to kill cells expressing residual transposase. (C) GFP fluorescence images and phase contrast images of HEK293T cells transfected with Tsase-puro dTK-Tson(GFP-pA) under no treatment (HEK:T(-)), ganciclovir treatment (HEK:T(G)), puromycin treatment (HEK:T(P)) and both ganciclovir and puromycin treatment (HEK:T(P+G)) for 10 days after three passages (Scale Bar = 100 µm). (D) The percentage of transfected HEK293T cells with transgene (GFP) integration as measured by flow cytometry. Measurements were conducted after 3 passages. Ganciclovir was added for 7 days after the third passage in the treatment group (HEK:T(G)). Results show no significant difference between the ganciclovir treated (HEK:T(G)) and untreated cells (HEK:T(-)) (p = 0.23; student's t-test). (E) RT-PCR of transfected and untransfected HEK293T cells with different treatments. The samples (n = 3) were probed for transgene (GFP) and transposase expression. GAPDH was used as the PCR control. The gels were cropped and the full-length gels are presented in the supplementary information (Supplementary Figure S7). (F) Percentage cell viability of transfected and untransfected HEK293T cells with different treatments, as measured by the MTT assay. Results indicate that transfection of the combination plasmid along with puromycin and HSV-tk/ganciclovir treatment significantly alters the viability of cells as measured by MTT assay (p<0.001, one-way ANOVA). Transfected cells treated with puromycin formed puromycin-resistant colonies with both transgene and transposase expression. However, when transfected cells are treated with both puromycin and ganciclovir, no colonies emerged. MTT assay data also shows significant difference between HEK:T(P) and HEK:T(P+G) groups (p<0.001, one-way ANOVA with Bonferroni post hoc test). In addition, RT-PCR results show the absence of transposase expression in transfected cells with ganciclovir treatment. (purodTK: puromycin-delta thymidine kinase; HEK:NT: untransfected HEK293T cells; HEK:T: transfected HEK293T cells).
Transposition of the human FIX gene using the self-inactivating, HSV-tk/ganciclovir combination plasmid
We have demonstrated that a model therapeutic gene, human FIX, could be integrated and expressed in human cell lines by our self-inactivating hyPBase transposition system (Figure 6). Furthermore, we verified that HSV-tk/ganciclovir treatment for elimination of residual transposase expression was also applicable for human FIX transposition. ICC staining and RT-PCR showed expression of FIX (Figure 6B–C). RT-PCR also showed a decrease in transposase expression levels after ganciclovir treatment (Figure 6C). We also demonstrated stable human FIX expression over a prolonged period of 25 days and 5 passages (Figure 6D). Tsase-puro dTK-Tson(hFIX-pA) showed more sustained and higher hFIX protein production than the control Tson(hFIX-pA) group.
Transposition of the human FIX gene using the self-inactivating, HSV-tk/ganciclovir combination plasmid.
(A) Illustration depicting the plasmid constructs Tsase-puro dTK-Tson(hFIX-pA) and Tson(hFIX-pA). (B) Immunocytochemistry (ICC) staining of HEK293T cells transfected with the hFIX-bearing combination plasmid (Scale Bar = 100 µm). (C) RT-PCR of untransfected and transfected (with and without ganciclovir treatment) HEK293T cells using human FIX, transposase and GAPDH primers. The gels were cropped and the full-length gels are presented in the supplementary information (Supplementary Figure S8). (D) Chromogenic assay quantifying the release of functional human FIX protein over 5 passages (25 days) from HEK293T cells transfected with Tsase-puro dTK-Tson(hFIX-pA) and Tson(hFIX-pA). Tsase-puro dTK-Tson(hFIX-pA) shows sustained hFIX protein production over a prolonged period which is significantly higher than Tson(hFIX-pA) group (p = 0.0011, two-way ANOVA). After 25 days post transfection, a significant difference in the hFIX level exists between the Tsase-puro dTK-Tson(hFIX-pA) and Tson(hFIX-pA) (p<0.001; two-way ANOVA with Bonferroni post hoc). (HEK:NT: untransfected HEK293T cells; HEK:T: transfected HEK 293T cells; G: Ganciclovir treatment; (-): cells without any treatment).
Discussion
In this study we review all the potential fates of the transposase (integration, cleavage, dilution and recircularization) in a combination plasmid format and implement an approach to self-inactivate the transposase expression cassette after transposition. Undesirable integration of the transposase into the genome can cause insertional mutagenesis and/or continued expression of transposase. Insertional mutagenesis, which results from disruption, suppression or activation of an endogenous gene, can lead to aberrant behavior of the cell. Continued expression of transposase due to integration or slow rate of plasmid dilution can be a potential cause of genotoxicity and cytotoxicity40. Plasmid dilution, which occurs due to cell division, is a better outcome than integration as there is no insertional mutagenesis and the transposase expression is transient. However, this transiency can be extended if there is a delay in transgene dilution due to lack of cell division, an inherent property of certain cell types like neurons and skeletal myocytes. The most desirable outcome is the one in which the transposase expression cassette is cleaved after transposition and there is no integration or prolonged expression of transposase (Figure 1A). The measures used in this study have the potential to stop expression of transposase due to integration and recircularization. This approach also has the potential to bypass the limitations associated with the combined donor-helper plasmid solution described by Urschitz et al.29. Although there are instances in the literature of a single termination signal being shared by two open reading frames (ORFs)41, yet uncertainty of the effect of such a combination on transposition efficiency prompted us to compare the single polyA combination plasmid with the double polyA forms. We observed similar degree of transgene (GFP) integration with our combination plasmid (single polyA form) when compared with the traditional double polyA combination form thereby indicating that the lack of a dedicated polyA does not have any negative effect on transposition efficiency. All the combination plasmids showed an increase in MFI after the first passage (Figure 2C). It can be speculated that this upward trend is probably due to dilution of the plasmids from cells that received fewer copies of the plasmid. These cells have low transposase levels; consequently little or no transposition occurs. In contrast, the cells receiving multiple copies of the plasmid have higher transposase levels enabling multiple integration events. This shifts the cell population having a range of GFP copy numbers after transfection to a cell population having only high GFP copy number resulting in the GFP MFI increase at passages 1 and 2. Therefore, there appears to be a minimum level of transposase needed for efficient transposition. This also implies that for optimum transposition a transfection method has to be chosen which can deliver high copies of the plasmid into the nucleus. Therefore, a thorough optimization of the transfection process is needed for individual cell types. After three passages the MFI was found to be significantly higher in the case of single polyA combination plasmid compared to both the double polyA combination plasmid forms. It can be speculated that following transposition there is less transgene excision in the case of single polyA combination plasmid as the transposase gets inactivated faster. This in turn results in a higher MFI.
The combined system performed better than the two plasmid system with respect to both the percentage of transposition positive cells and the average expression levels. This effect can be accounted for by the reliable co-delivery of both elements to the same cell. Moreover, more plasmids can be packed in the transfection agent in the case of a combination plasmid. The two-plasmid system also lacks a self-inactivation mechanism for the transposase, which may lead to toxicity and transgene excision and consequently lower transposition efficiency.
After proving the efficacy of our self-inactivating combination plasmid we assessed the problem of residual transposase expression. Theoretically, residual transposase expression can be found even in the self-inactivating combination plasmid group if the whole plasmid integrates before transposition. This sort of integration happens due to random linearization of the plasmid during plasmid production or due to the action of cellular nucleases. The residual expression can also stem from post-transposition integration of the transposase. In most of such cases the transposase will be silent as it lacks a polyA. However, polyA-less transposase may express if integration occurs in an active area of the genome with an endogenous polyA support in a phenomenon akin to the polyA trap experiments42. To quantify the level of residual transposase expression we used the residual mOrange expression (co-expressed with transposase from the same promoter) as a proxy. In all three conditions very few cells expressed mOrange after three passages (< 1%, relative to P0 GFP+ cells). This suggests that integration of the transposase is not a frequently occurring phenomenon in the combination plasmid format. Median mOrange fluorescence intensity measurements showed that the Tsase-mO-pA-Tson(GFP-pA) group had a higher normalized MFI than the other two groups as early as passage P0 + 5 days. This difference in MFI can only be accounted for by the robust expression of the integrated residual transposase in the case of Tsase-mO-pA-Tson(GFP-pA) due to the presence of a dedicated polyA for the transposase. The other two plasmids are self-inactivated and have to depend on endogenous polyA for expression (Supplementary Figure S2). This may lead to considerably weaker transposase expression. Moreover, the MFI of Tsase-mO-Tson(GFP-pA) is similar to Tsase-mO-Tson(GFP-pA)-pA even in the early passages when the dilution effect on plasmids is not pronounced. Among the two plasmids only Tsase-mO-Tson(GFP-pA)-pA can theoretically support recircularization-mediated transposase expression (Supplementary Figure S2). This indicates that post-transposition plasmid recircularization is not a major mechanism determining the fate of the plasmid backbone after transposition. Therefore, when compared to integration and recircularization, cleavage seems to be the most prominent mechanism for the disposal of plasmid backbone.
After demonstrating the value of the self-inactivating combination plasmid-based transposition system, experiments were designed to solve the problem of residual transposase expression. We used HSV-tk/ganciclovir system for efficiently clearing residual transposase expression. The negative selection of transposase-free cells is based on the conversion of the prodrug ganciclovir to a toxic phosphorylated form by the enzyme HSV-tk. The FDA-approved status of ganciclovir enhances the applicability of our system in translational studies. Successful utilization of the HSV-tk/ganciclovir system also led us to ask if HSV-tk/ganciclovir treatment had a significant effect on the cells where transposition had already taken place. Studies revealed that very few transposition-positive cells (GFP+) expressed the transposase stably as addition of ganciclovir had no significant effect on the percentage of GFP-positive cells.
This study also highlights the potential of this vehicle to act as an efficacious gene delivery vector by using a model therapeutic gene (human FIX). However, a potential deterrent in using the vehicle may be the size of the construct which can decrease transfection efficiency. In future studies, the size of the transgene can be reduced by removing the puromycin cassette (which is not essential for gene therapy purposes) and fusing the HSV-tk cassette directly to the transposase by a self-cleaving 2a construct. These measures will decrease the size of the construct by more than 1 kb43. Moreover, the construct can be loaded onto a minicircle vector to make it smaller. The minicircle form will also remove the bacterial backbone leading to a higher and more reliable expression of the transposase44.
In conclusion, we expect that this donor-helper combination plasmid design along with the HSV-tk/ganciclovir fail-safe system will help realize the translational potential of hyperactive piggyBac transposon-based systems. The significance of this combination plasmid design can be extended to other transposition systems as well as targeted gene editing system like CRISPR/Cas9. This will ultimately impact cellular reprogramming strategies, cell-based therapeutics and gene therapy research.
Methods
Plasmid construction
In silico plasmid design was performed using PlasmaDNA software (University of Helsinki). Plasmid constructs pPB-Ubc and pCMV-hyPBase were acquired from Wellcome Trust Sanger Institute PiggyBac Transposase Resources (Hinxton, UK). Tson(GFP) was created by inserting GFP cassette distal to the Ubiquitin promoter of the transposon vector pPB-Ubc. The IRES-mOrange fragment was amplified by PCR from pCAG2LMKOSimO (Addgene Plasmid 20866) and cloned into pCMV-hyPBase to create Tsase-mO. Tsase-mO and Tson(GFP) constructs were combined to create Tsase-mO-pA-Tson(GFP-pA) and Tsase-mO-Tson(GFP-pA). The polyA component from pIRES-puro (Addgene Plasmid 16616) was inserted into Tsase-mO-Tson(GFP-pA) to create Tsase-mO-Tson(GFP-pA)-pA. The puro-dTK component of pLCA.66/2272 (Addgene Plasmid 22733) was used to construct Tsase-puro dTK-Tson(GFP). Tson(hFIX) and Tsase-puro dTK-Tson(hFIX) was created by replacing GFP with human FIX gene45.
Cell culture and transfection
HEK-293T cells were seeded on 0.1% gelatin coated 12-well plates. DMEM-HG (GIBCO-11960) supplemented with L-glutamine, pyruvate, MEM-NEAA (GIBCO) and 10% fetal bovine serum (FBS) (Atlanta Biologicals) was used to culture the cells. The calcium phosphate particle-based method was used for transfecting HEK-293T cells with the plasmid constructs. Once HEK-293T cells were more than 70% confluent, 1 ml of fresh medium was added to each well 2 hours before the transfection. The transfection mixture for individual wells was composed of 1.2 µg of plasmid DNA, 18.39 µl of TE 0.1X buffer, 3.14 µl of CaCl2 and water up to the total volume of 31.76 µl. Subsequently 31.76 µl of HBS 2X buffer was slowly added to the transfection mixture under continuous vortexing. The transfection mixture was added to each well after 10 minute incubation at room temperature.
Flow cytometry
The gene expression levels of GFP and mOrange in the transfected HEK-293T cells were measured by using FACSCanto II flow cytometer and FACSDiva software (BD Biosciences). Data analysis was done using FlowJo software (Treestar, Inc).
The transfection for each sample (n = 4 for each group) was done in duplicates; one replicate to passage and the other to measure gene expression level at passage 0 (12 hours after transfection). First passage (split ratio of 1:10) was performed after transfected cells were incubated for 5 days (labeled as Passage 0 + 5 days) to prevent rapid dilution of the transposon system even before transposition has taken place. The subsequent passages (split ratio of 1:10) were performed when cells became more than 90% confluent in approximately 2–3 days.
HSV-tk/ganciclovir selection, cell viability test and RT-PCR
In order to eliminate residual transposase-expressing cells, HEK-293T cells were transfected with Tsase-puro dTK-Tson(GFP) and passaged multiple times. Cells were treated with either 3 µg/ml puromycin (Sigma), 4 µM ganciclovir (Invivogen), or both puromycin and ganciclovir. Treatment was maintained by changing the medium every day for 10 days. Transfected cells that received no treatment and untransfected cells were used as controls. On the 10th day, fluorescence (GFP) and phase contrast images were taken and cell viability (MTT assay, Sigma M5655) was measured according to manufacturer's protocol.
To validate the expression of our plasmid construct, RT-PCR using primers for transposase (5′-TGATGACCTGCAGCAGAAAG-3′, 5′-GCTGATGTTGTCCCTCAGGT-3′) and GFP (5′-GAGCAAGGGCGAGGAGCT-3′, 5′-GTCCTCCTTGAAGTCGATGCC-3′) was performed. After transfected cells were treated for 10 days with either puromycin or ganciclovir, total RNA was extracted using Qiagen RNeasy mini kit. RNA from control groups (transfected cells that received no treatment and untransfected cells) was also extracted.
Human blood coagulation Factor IX chromogenic assay, immunocytochemistry and RT-PCR
FIX chromogenic assay (Aniara, Biophen) was used to measure the concentration of FIX secreted by HEK-293T cells transfected with Tsase-puro dTK-Tson(hFIX). Tson(hFIX) was used as a control. 25,000 HEK-293T cells were seeded into each of the 6-well plates and transfected with the plasmids. The seeding and transfection was done in duplicates; one to passage and the other to condition the media for the chromogenic assay. On the following day after transfection, 2 ml of 6 µg/ml of Vitamin K- supplemented serum-free medium was applied to the cells for 24 hours. The conditioned media was then collected and later used for the chromogenic assay performed according to manufacturer's protocol. The process was continued for 5 passages to measure sustained secretion of FIX.
Immunocytochemistry to stain for FIX in transfected cells was done using 20 µg/ml of mouse anti-human FIX primary antibody (Hematologic technologies, AHIX-5041). Alexa Fluor 488 Goat Anti-Mouse IgG (Life technologies, A-11001) was utilized as secondary antibody.
RT-PCR was performed to validate the presence of hFIX expression and elimination of transposase under HSV-tk/ganciclovir selection. Primers used for the detection of hFIX expression were 5′-TCCATCGTGAACGAGAAGTG-3′ and 5′-TAGTTGTGGTGGGGGATGAT-3′. Cells were transfected with Tsase-puro dTK-Tson(hFIX) and one group was treated with ganciclovir while the other was left untreated. Untransfected cells were used as a negative control. Total RNA was extracted using the Qiagen RNeasy mini kit. RT-PCR was performed using the MyTaqTM One-Step RT-PCR Kit (Bioline) and the cycling conditions were: 45°C for 20 minutes (reverse transcription), 95°C for 1 minute (polymerase activation), 29 (22 for GAPDH) cycles of 95°C for 10 seconds (denaturation), 58°C for 10 seconds (annealing) and 72°C for 30 seconds (extension). The gels were run under the same experimental conditions. The gels were cropped and the bands are presented using Microsoft PowerPoint software.
GraphPad Prism 5.0 statistical analysis software was utilized for all the statistical analysis. All the error bars are standard error of means. Statistical significance was determined at a P value of ≤ 0.05. Two-way ANOVA with an appropriate post hoc test (mentioned in the figure legends) was used to analyze the effect of transposon plasmid type with passaging of the transfected cells on the parameters of median fluorescence intensity and percentage fluorescence.
References
WyssCoray, T. et al. Amyloidogenic role of cytokine TGF-beta 1 in transgenic mice and in Alzheimer's disease. Nature 389, 603–606 (1997).
Moretti, A. et al. Multipotent embryonic isl1+ progenitor cells lead to cardiac, smooth muscle and endothelial cell diversification. Cell 127, 1151–1165 (2006).
Keen, H. et al. Human insulin produced by recombinant DNA technology: safety and hypoglycaemic potency in healthy men. Lancet 2, 398–401 (1980).
Wood, W. I. et al. Expression of active human factor VIII from recombinant DNA clones. Nature 312, 330–337 (1984).
Li, H. et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 475, 217–221 (2011).
Pawliuk, R. et al. Correction of sickle cell disease in transgenic mouse models by gene therapy. Science 294, 2368–2371 (2001).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010).
Kolls, J. K. et al. Use of transient CD4 lymphocyte depletion to prolong transgene expression of E1-deleted adenoviral vectors. Hum. Gene Ther. 7, 489–497 (1996).
Kumar, M., Keller, B., Makalou, N. & Sutton, R. E. Systematic determination of the packaging limit of lentiviral vectors. Hum. Gene Ther. 12, 1893–1905 (2001).
Huang, X. & Yang, Y. Innate immune recognition of viruses and viral vectors. Hum. Gene Ther. 20, 293–301 (2009).
Pfeifer, A. Lentiviral transgenesis. Transgenic Res. 13, 513–522 (2004).
Segall, H. I., Yoo, E. & Sutton, R. E. Characterization and detection of artificial replication-competent lentivirus of altered host range. Mol. Ther. 8, 118–129 (2003).
Warnock, J. N., Daigre, C. & Al-Rubeai, M. Introduction to viral vectors. Methods Mol. Biol. 737, 1–25 (2011).
Flotte, T. R. Gene therapy: The first two decades and the current state-of-the-art. J. Cell. Physiol. 213, 301–305 (2007).
Ivics, Z., Hackett, P. B., Plasterk, R. H. & Izsvak, Z. Molecular reconstruction of Sleeping beauty, a Tc1-like transposon from fish and its transposition in human cells. Cell 91, 501–510 (1997).
Hackett, P. B., Ekker, S. C., Largaespada, D. A. & McIvor, R. S. Sleeping beauty transposon-mediated gene therapy for prolonged expression. Adv. Genet. 54, 189–232 (2005).
Ivics, Z. & Izsvak, Z. Transposons for gene therapy! Curr. Gene Ther. 6, 593–607 (2006).
Wilson, M. H., Coates, C. J. & George, A. L., Jr PiggyBac transposon-mediated gene transfer in human cells. Mol. Ther. 15, 139–145 (2007).
Wu, S. C. Y. et al. piggyBac is a flexible and highly active transposon as compared to Sleeping Beauty, Tol2 and Mos1 in mammalian cells. Proc. Natl. Acad. Sci. USA 103, 15008–15013 (2006).
Feschotte, C. The piggyBac transposon holds promise for human gene therapy. Proc. Natl. Acad. Sci. USA 103, 14981–14982 (2006).
Huang, X. et al. Gene Transfer Efficiency and Genome-Wide Integration Profiling of Sleeping Beauty, Tol2 and PiggyBac Transposons in Human Primary T Cells (vol 18, pg 1803, 2010). Mol. Ther. 18, 2038–2038 (2010).
Kahlig, K. M. et al. Multiplexed transposon-mediated stable gene transfer in human cells. Proc. Natl. Acad. Sci. USA 107, 1343–1348 (2010).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009).
Kaji, K. et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 458, 771–775 (2009).
Dupuy, A. J. et al. Mammalian germ-line transgenesis by transposition. Proc. Natl. Acad. Sci. USA 99, 4495–4499 (2002).
Ding, S. et al. Efficient transposition of the piggyBac resource (PB) transposon in mammalian cells and mice. Cell 122, 473–483 (2005).
Suganuma, R. et al. Tn5 transposase-mediated mouse transgenesis. Biol. Reprod. 73, 1157–1163 (2005).
Urschitz, J. et al. Helper-independent piggyBac plasmids for gene delivery approaches: strategies for avoiding potential genotoxic effects. Proc. Natl. Acad. Sci. USA 107, 8117–8122 (2010).
Marh, J. et al. Hyperactive self-inactivating piggyBac for transposase-enhanced pronuclear microinjection transgenesis. Proc. Natl. Acad. Sci. USA 109, 19184–19189 (2012).
Liang, Q., Kong, J., Stalker, J. & Bradley, A. Chromosomal Mobilization and Reintegration of Sleeping Beauty and PiggyBac Transposons. Genesis 47, 404–408 (2009).
Wang, W. et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 105, 9290–9295 (2008).
Wurtele, H., Little, K. C. E. & Chartrand, P. Illegitimate DNA integration in mammalian cells. Gene Ther. 10, 1791–1799 (2003).
Wang, Z. et al. Detection of integration of plasmid DNA into host genomic DNA following intramuscular injection and electroporation. Gene Ther. 11, 711–721 (2004).
Eszterhas, S. K., Bouhassira, E. E., Martin, D. I. K. & Fiering, S. Transcriptional interference by independently regulated genes occurs in any relative arrangement of the genes and is influenced by chromosomal integration position. Mol. Cell. Biol. 22, 469–479 (2002).
Hacien-Bey-Abina, S. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1 (vol 302, pg 415, 2003). Science 302, 568–568 (2003).
Shimizu, A. & Shimizu, N. Dual promoter expression system with insulator ensures a stringent tissue-specific regulation of two reporter genes in the transgenic fish. Transgenic Res. 22, 435–444 (2013).
Barese, C. N. et al. Thymidine kinase suicide gene-mediated ganciclovir ablation of autologous gene-modified rhesus hematopoiesis. Mol. Ther. 20, 1932–1943 (2012).
Yusa, K., Zhou, L. Q., Li, M. A., Bradley, A. & Craig, N. L. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl. Acad. Sci. USA 108, 1531–1536 (2011).
Galla, M. et al. Avoiding cytotoxicity of transposases by dose-controlled mRNA delivery. Nucleic Acids Res. 39, 7147–7160 (2011).
Kita-Matsuo, H. et al. Lentiviral vectors and protocols for creation of stable hESC lines for fluorescent tracking and drug resistance selection of cardiomyocytes. PloS one 4, e5046 (2009).
Stanford, W. L., Cohn, J. B. & Cordes, S. P. Gene-trap mutagenesis: past, present and beyond. Nat. Rev. Genet. 2, 756–768 (2001).
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PloS one 6, e18556 (2011).
Chen, Z. Y., He, C. Y., Ehrhardt, A. & Kay, M. A. Minicircle DNA vectors devoid of bacterial DNA result in persistent and high-level transgene expression in vivo. Mol. Ther. 8, 495–500 (2003).
Chakraborty, S., Christoforou, N., Fattahi, A., Herzog, R. W. & Leong, K. W. A robust strategy for negative selection of Cre-loxP recombination-based excision of transgenes in induced pluripotent stem cells. PloS one 8, e64342 (2013).
Acknowledgements
We thank Pablo Perez-Pinera for helpful discussions and preliminary experiments. We also acknowledge a) Keisuke Kaji for Addgene Plasmid 20866: pCAG2LMKOSimO; b) Bert Vogelstein for Addgene Plasmid 16616: pIRES-puro-GFP and c) Mark Magnuson for Addgene Plasmid 22733: pLCA.66/2272. Support from NIH (KWL: EB015300, AI096305, UH2TR000505; CAG: DP2OD008586,), NSF (CBET-1151035), American Heart Association (10SDG3060033) and the Muscular Dystrophy Association (MDA277360) is acknowledged.
Author information
Authors and Affiliations
Contributions
S.C., C.A.G. and K.W.L. designed the experiments, S.C., H.J. and J.C. performed the research, S.C. and K.W.L. analyzed the data, S.C., C.A.G. and K.W.L. wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figure Legends
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Chakraborty, S., Ji, H., Chen, J. et al. Vector modifications to eliminate transposase expression following piggyBac-mediated transgenesis. Sci Rep 4, 7403 (2014). https://doi.org/10.1038/srep07403
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07403
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.