Abstract
Hit to lead (H2L) optimization is a key step for drug and agrochemical discovery. A critical challenge for H2L optimization is the low efficiency due to the lack of predictive method with high accuracy. We described a new computational method called Computational Substitution Optimization (CSO) that has allowed us to rapidly identify compounds with cytochrome bc1 complex inhibitory activity in the nanomolar and subnanomolar range. The comprehensively optimized candidate has proved to be a slow binding inhibitor of bc1 complex, ~73-fold more potent (Ki = 4.1 nM) than the best commercial fungicide azoxystrobin (AZ; Ki = 297.6 nM) and shows excellent in vivo fungicidal activity against downy mildew and powdery mildew disease. The excellent correlation between experimental and calculated binding free-energy shifts together with further crystallographic analysis confirmed the prediction accuracy of CSO method. To the best of our knowledge, CSO is a new computational approach to substitution-scanning mutagenesis of ligand and could be used as a general strategy of H2L optimisation in drug and agrochemical design.
Similar content being viewed by others
Introduction
Agrochemicals play a significant role in modern agriculture by protecting crops against yield loss. Due to the increasing demand for food supply and the explosive development of resistance, the discovery and development of new agrochemicals are continuously demanded1. However, agrochemical discovery remains a vital challenge2: the research and development (R&D) costs have been rising from U.S. $152 million in 1995 to $256 million in 2005 and the number of compounds needs to be synthesized have been rising from 52500 in 1995 to 140000 in 2005 to bring a new agrochemical to market. A new agrochemical is invented initially by the discovery of a bioactive lead compound. Thus, lead generation is a crucial step for the agrochemical discovery3, which requires at least two processes: hit identification and Hit-to-Lead (H2L) optimization4. Over the past decade, introduction of new technologies, such as high-throughput screening (HTS) and fragment-based design, have made hit identification become more efficient5,6. But, traditional H2L optimization is still carried out through a large number of analogues’ syntheses, which lead to a low efficient and resource-intensive research7. Although some techniques, such as quantitative structure-activity relationship (QSAR)8 and molecular dynamics (MD) simulations9,10, have been developed to help understand the structure-activity relationship for H2L optimization, they still have some drawbacks, such as dependent of sufficient biological data, highly time-consuming and low prediction accuracy11. Therefore, there is urgent demand in developing an effective tool enabling to eliminate undesirable lead compound classes/types early during H2L optimization process, prior to expensive and extensive experimental work12. Having this challenge in mind, we developed a new computational protocol called Computational Substitution Optimization (CSO) to rapidly assess how replacing hydrogen atoms with new substituent into a “hit” scaffold could improve binding free energies to their targets. To showcase this method, we applied CSO in the H2L optimization of cytochrome bc1 complex inhibitors to discover antifungal lead.
The cytochrome bc1 complex (EC 1.10.2.2, bc1) has been chosen as a model for the present study, because of its crucial role in the life cycle. The bc1 complex is an essential respiratory enzyme complex present in the inner mitochondrial membrane of eukaryotic organisms. It has been identified as a promising target for new drugs and agrochemicals13. For example, malaria, as a devastating tropical disease, remains a major threat to over 40% of the world’s population and approximately 660,000 deaths in 201014. Atovaquone is a major drug for prophylaxis and treatment of uncomplicated malaria patient15 and as alternative therapy in case of resistances against chloroquine or artemisin-based therapies16. But with a limited number of antimalarial drugs available, the emergence of multidrug-resistant Plasmodium falciparum malaria threatens the health of individual patients as well as eradication strategies for the disease17. In addition, some bc1 complex inhibitors have been introduced into the agrochemical market to provide effective control of important fungal diseases threatening global food security18. Azoxystrobin (AZ), as a representative inhibitor targeting bc1, is the biggest using fungicide in the whole word with a global annual sales of over USD1.4 billion19. However, the increasing number of resistance is also a big problem that AZ has to face19. Therefore, there is a wide research interest on the discovery of cytochrome bc1 complex inhibitor today, which could be used as a specific probe for novel functional study and as a new lead compound for future drug and agrochemical discovery20.
In this study, the application of CSO has led to a series of new bc1 inhibitors with nanomolar to subnanomolar potency. Excellent agreements between the predicted and experimental binding free energies were obtained, indicating the predictive capacity of CSO. Further, assays showed that compound 18 was the lead candidate from the new series of bc1 complex inhibitors because of its high potency (Ki = 4.10 nM which is up to 73-fold higher than that of AZ), outstanding physiochemical properties and excellent in vivo bioactivity. In order to investigate the underlying mechanisms responsible for the excellent in vivo bioactivity, the kinetic effects on cytochrome bc1 complex were studied and exhibited slow-binding characteristics, which is different from the classical fast-binding of AZ indicating a turning point for bc1 complex inhibitor discovery. Most importantly, the determined crystal structure of lead compound bound to bc1 complex at 3.23 Å provided a solid molecular basis for the prediction accuracy of CSO method. This work demonstrated that CSO method can significantly improve the efficiency of H2L optimization process and could be used as a general strategy for drug and agrochemical discovery.
Results
H2L Optimization through CSO
The workflow of the computational protocol used in this study is shown in Fig. 1. This new protocol is a combination of a MD simulation and a free-energy perturbation (FEP)-based scanning calculation. Due to E-Methyl 2-(2-((benzo- thiazol-2-ylthio)methyl)phenyl)-3-methoxyacrylate (compound 1) was identified previously as a hit compound targeting cytochrome bc1 complex (Ki = 31.10 nM, against porcine bc1)21, the MD simulation was first performed on compound 1-bound bc1 complex system. To evaluate the convergence of the MD simulation, plots of root-mean-square deviation (RMSD) and the distances of hydrogen bonds were examined (Fig. S1).
Diagrammatic representation of the workflow of CSO.
The CSO protocol was designed to automatically perform computational substitution, energy minimization and binding affinity evaluation: (1) The MD simulation of protein with original “hit” ligand was performed. (2) In order to obtain a representative ensemble of the binding structures, snapshots were collected from the MD trajectory at regular intervals and minimized. (3) The “Hit” ligand in each snapshot collected from the MD trajectory was mutated to new ligand with substitutions using the library of substitution and minimized. Each substitution group was set to link with the specific site of “Hit” compounds. (4) ΔΔG was calculated and the final binding free energy change is the average of the calculated values associated with each snapshot. New leads were finally determined according to the order of the binding free-energy shifts.
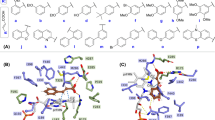
Due to direct interactions of the substituent with the phenyl ring of phenylalanine, electron-donating and electron-withdrawing substituent can effectively increase the π-π stacking interactions22. Hence, some typical groups (a: −F, b: −Cl, c: −Br, d: −NO2, e: −CH3, f: −CF3, g: −OCH3, h: −OH, i: −NH2, j: −COOH) were introduced one by one onto different sites of the benzothiazole ring of 1 to produce compounds 2–41 and its nitrogen analogues 42–81, respectively (Table 1). To avoid excessive perturbation in FEP-based calculation, only single substitutions were investigated in the present study. In total, 80 computational substitutions were made and the computational changes in binding free energies as a result of each substitution were calculated for each compound. Detailed results of the binding free-energy changes are provided in Table S1.
A 10-fold improvement of potency should be the minimum standard for an acceptable H2L optimization process. Thus, by setting the criterion of the calculated binding free-energy shift (∆∆Gcal) lower than −1.37 kcal/mol (~10-fold improvement in activity), 29 compounds were considered as candidates for further chemical synthesis. Considering the feasibility of organic synthesis, we only synthesized 10 out of them which are easy to synthesize. Their experimental binding free-energy shifts (∆∆Gexp) ranged from −2.69 to −1.40 kcal/mol. For an ideal method, it can predict not only compounds with higher potency, but also compounds with lower potency. Hence, a further 11 compounds those ∆∆Gcal larger than −1.37 kcal/mol and easier to synthesise were also selected randomly for chemical synthesis. The synthetic routes for the total 21 compounds are summarised in Scheme S1. Their chemical structures were characterised by 1H and 13C NMR spectroscopy, HRMS, elemental analysis and X-ray diffraction analysis (Fig. S2).
In Vitro Activities and Lead Identification
Inhibitory kinetics is of great importance to understand the inhibitory activity of compounds. The inhibition constants (Ki = 0.39–159.96 nM) of these compounds with the bc1 complex were determined according to previously established method23 and listed in Table 1. These results indicate that the newly synthesized compounds are specific and effective inhibitors of bc1 complex. Notably, the experimental and calculated binding free-energy shifts (ΔΔGexp and ΔΔGcal, respectively) of these compounds showed good linear correlation (r2 = 0.8), confirming the predictive accuracy of the CSO method.
Based on the above in vitro study, compounds with nanomolar and subnanomolar Ki values were selected from the total 21 newly synthezised compounds for further studies. Pesticide likeness24, defined as a complex balance of various physicochemical properties and structure features, can evaluate whether a molecule is similar to the known pesticides. After the filtration of pesticide likeness, seven compounds were kept to adhere to our previously defined rules for pesticide-likeness including molecular weight (MW) ≤ 435 Da, ClogP ≤ 6, number of H-bond acceptors (HBA) ≤ 6, number of H-bond donors (HBD) ≤ 2, number of rotatable bonds (ROB) ≤ 9 and number of aromatic bonds (ARB) ≤ 17 (Table 1)24. Finally, based on the results of in vivo test, three out of seven compounds were found to display good antifungal activity (>70%) against downy mildew and powdery mildew at the concentration of 200 mg/L (Table S2). Among these three compounds, compound 18 completely inhibited the growth of both species of fungus, outperforming the control fungicide, AZ (which inhibited the growth of powdery mildew, but only reduced the growth of downy mildew by 94%). Therefore, compound 18 was recognized as lead compound for subsequent studies.
In Vivo Antifungal Activity
The above results indicated that compound 18 shows excellent potency towards bc1 complex, pesticide-like properties and antifungal activity. For a further validation, the controlling efficacy of compound 18 against downy mildew and powdery mildew disease was tested in greenhouse and field models. Because the pathogens exhibit a specific host for cucumber, the inoculations of Pseudoperonospora cubensis (P. cubensis) and Sphaerotheca fuliginea (S. fuliginea) were carried out by spraying a conidial suspension on the seed leaves of cucumber. As expected, the greenhouse test indicated that compound 18 showed excellent fungicidal activity against both downy mildew and powdery mildew diseases. The protective and curative EC90 values of compound 18 against P. cubensis were 4.29 mg/L and 120.29 mg/L, respectively and against S. fuliginea are 1.89 mg/L and 32.89 mg/L, respectively. As a control, the protective and curative EC90 values of AZ against P. cubensis are 5.36 mg/L and 50.66 mg/L, respectively and against S. fuliginea are 254.30 mg/L and >500 mg/L, respectively.
To further study the potential of compound 18 against powdery mildew and downy mildew, field experiments were conducted during the growing season of summer squash and cucumber. The summer squash plants at a fruiting stage naturally infected by S. fuliginea were used. A foliar application of compound 18 at 75 g.ai/ha recorded 74.60% disease reduction over control. Almost no lesions were observed on the leaves of summer squash. If the treatment concentration of compound 18 was reduced by half to 37.5 g.ai/ha, lesions on the leaves of treated plants expanded very slowly, or ceased to expand, which demonstrates that compound 18 has a curative activity at 37.5 g.ai/ha against the expansion of lesions by S. fuliginea with a curative effect of approximately 70.92% (Fig. 2A,B). Lesions on the leaves of control plants, however, expanded quickly and intense fungal hyphae could be observed. The curative effect is approximately 64.85% for the AZ-treated (93.75 g.ai/ha) summer squash. In addition, field trials for the treatment of downy mildew of cucumber were also performed. Cucumber plants naturally infected by P. cubensis were used for the study. Fungi on the leaves of control plants expanded quickly and lesions could be observed. However, we observed that when compound 18 (60 g.ai/ha) was sprayed on the leaves of the cucumber at the primary infection location, only small or no lesions were observed and the statistical curative effect was approximately 82.59% (Fig. 2B). As a comparison, when leaves of cucumber were treated with AZ (75 g.ai/ha), a lower curative effect (59.59%) was observed. The field experiments show that the curative effect of compound 18 is superior to AZ as a treatment of powdery mildew and downy mildew.
Protection of summer squash and cucumber from attack by Sphaerotheca fuliginea and Pseudoperonospora cubensis, respectively, by compound 18.
(A) Compound 18 prevented S. fuliginea from infecting and killing summer squash. Compound 18, at different concentrations, were spread on leaves and photographs were taken after 7 days. (B) The relative control effect of compound 18 against powdery mildew and downy mildew in field trials.
Action Mechanism
Based on the above in vitro and in vivo studies, it is reasonable to speculate that compound 18 might have different action mechanism from AZ. Hence, the kinetic effects of compound 18 on cytochrome bc1 complex were investigated and compared with compound 1 (hit compound) and AZ (Fig. 3A).
Inhibitory kinetics of bc1 complex.
(A) Chemical structures of AZ (azoxystrobin) and its analogues, compound 1 and compound 18. These compounds have a common pharmacophore structure: methoxyacrylate (MOA). (B) AZ is a competitive inhibitor with respect to substrate DBH2. Each reaction mixture contains phosphate buffered saline (PBS; 100 mM, pH 6.5), ethylenediaminetetraacetic acid (EDTA; 2 mM), lauryl maltoside (750 μM), oxidised cytochrome c (100 μM), SCR (0.05 nM), DBH2 (20–120 μM) and a specified amount of AZ (1, 0 nM; 2, 500 nM; 3, 1000 nM; 4, 2000 nM; 5, 3000 nM; and 6, 5000 nM). (C) Compound 1 is non-competitive with respect to cytochrome c. The reaction mixture contains 100 mM, pH 7.4), EDTA (0.3 mM), succinate (20 mM), SCR (0.1 nM), cytochrome c (1.13–15.6 μM) and a series of concentrations of compound 1 (1, 0 nM; 2, 10 nM; 3, 20 nM; 4, 40 nM; and 5, 60 nM). (D) Compound 1 is competitive with respect to DBH2. Each reaction mixture contains 100 mM, pH 6.5), EDTA (2 mM), lauryl maltoside (750 μM), oxidised cytochrome c (100 μM), SCR (0.05 nM), DBH2 (20–120 μM) and a specified amount of compound 1 (1, 0 nM; 2, 50 nM; 3, 100 nM; 4, 200 nM; 5, 300 nM; and 6, 500 nM).
Our previous data showed that AZ inhibits the activity of SCR with Ki = 297.6 ± 7.9 nM and that AZ is non-competitive with respect to cytochrome c23. To address the competition of AZ with respect to ubiquinol, we conducted inhibition assays for bc1 complex (Assay 3) at varying concentrations of DBH2 in the absence and presence of the inhibitor AZ. Figure 3B shows a set of double-reciprocal plots of 1/v0 versus 1/[DBH2], which yields a series of straight lines with a common intercept on the 1/v axis with different slopes. Because the intercepts of each line on the 1/[DBH2] and 1/v axes are equal to −1/Km and 1/Vmax, respectively, at the indicated AZ concentration, we can conclude that Km for the DBH2 substrate increases with increasing AZ concentration, whereas Vmax is not affected. These results indicate that AZ is a competitive inhibitor of bc1 complex with respect to substrate DBH2 and its kinetic parameters are determined as Km(DBH2) = 80.0 ± 7.0 μM and Ki = 394.7 ± 19.6 nM.
To unravel the inhibitory mechanism of compound 1, we first examined the dependence of product formation on the concentration of the cytochrome c as substrate by monitoring the initial rates of SCR (Assay 1). When the succinate concentration was fixed and that of cytochrome c varied in the presence of various fixed concentrations of compound 1, the double-reciprocal plots yielded a non-competitive pattern (Fig. 3C). Analysis of the data yielded Km and Ki values of 4.3 ± 0.2 μM and 31.1 ± 0.9 nM, respectively. Next, we analysed the effect of ubiquinol as substrate in the DBH2-cytochrome c system (Assay 3). When DBH2 was used as the substrate, a competitive pattern was observed for compound 1, in which a series of straight lines converged to a point on the vertical axis (Fig. 3D). The Ki value was determined as 96.5 ± 5.6 nM, which is approximately 3-fold higher than that determined in the succinate-cytochrome c system. This difference likely results from the presence of lauryl maltoside in the assay system, as suggested in previous studies23. Taken together, compound 1 is a non-competitive inhibitor with respect to substrate cytochrome c, but a competitive inhibitor with respect to DBH2, similar to fungicide AZ.
Different with AZ and compound 1, compound 18 unexpectedly behaves as a slow-binding methoxyacrylate (MOA)-type inhibitor of bc1 complex (Figs 4A and S3). In the presence of compound 18, the product formation curve for the SCR-catalysed reaction begins linearly but falls off and approaches a steady state with increasing time. However, when the enzyme was pre-incubated with compound 18 to reach the binding equilibrium, the progress curve simply displays a linear product-versus-time relationship (line 3), with a rate approximately the same as the steady-state rate of line 2 (Fig. S3). These data suggest that compound 18 is a slow-binding inhibitor of bc1 complex.
Slow-binding inhibition mechanism.
(A,B) Effect of various concentrations of cytochrome c and compound 18 on the inhibition of SCR activity. Each reaction mixture contains 100 mM, pH 7.4), EDTA (0.3 mM), succinate (20 mM), SCR (0.1 nM) and indicated amounts of cytochrome c and compound 18. The assays were performed in the presence of (A) 18 (35 nM) and a specified amount of cytochrome c (1, 4 μM; 2, 8 μM; 3, 20 μM; and 4, 48 μM) or (B) cytochrome c (60 μM) and various concentrations of compound 18 (1, 0 nM; 2, 15 nM; 3, 20 nM; 4, 30 nM; 5, 40 nM; and 6, 50 nM). Experimental data are shown as coloured dots and theoretical values are represented as black solid lines. Insets: plots of Kobs against concentration of (A) cytochrome c and (B) compound 18. (C,D) Effect of various concentrations of DBH2 and 18 on the inhibition of bc1 complex activity. Each reaction mixture contains 100 mM, pH 6.5), EDTA (2 mM), lauryl maltoside (750 μM), oxidised cytochrome c (100 μM), SCR (0.05 nM) and indicated amounts of DBH2 and compound 18. Two representative sets of inhibitory assays were performed in the presence of (C) DBH2 (20 μM) and a specified amount of compound 18 (1, 20 nM; 2, 40 nM; 3, 60 nM; 4, 80 nM), or (D) DBH2 (120 μM) and a specified amount of compound 18 (1, 40 nM; 2, 80 nM; 3, 120 nM; 4, 180 nM). Insets: plots of Kobs against concentration of compound 18. (E) Plot of the apparent rate constant A against concentration of DBH2. Inset: Plot of 1/A against concentration of DBH2. (F) Plot of the apparent rate constant B against concentration of DBH2.
To reveal the kinetic mechanism underlying this inhibition, we first examined the effect of the concentration of the cytochrome c as substrate and the inhibitor compound 18 on product formation by following the SCR-catalysed reaction (Assay 1). In the presence of different concentrations of cytochrome c and a fixed concentration of compound 18, the time-dependence of the reactions follows the substrate reaction kinetic theory and the kinetic parameters (the initial [v0] and steady-state [vs] rate of the reaction and the observed first-order rate constant [kobs]) can be determined by nonlinear regression analysis (Fig. 4A). As shown in the inset, the plot of kobs against the concentration of compound 18 is a horizontal line, which clearly indicated that compound 18 is non-competitive with respect to cytochrome c23. Consequently, the values of the true association and dissociation rate constants k+0 and k−0 were equal to those of the apparent constants A and B, respectively and can be determined from one set of enzymatic assays by varying the concentration of compound 18 at a fixed cytochrome c concentration (Fig. 4B). The kinetic parameters of compound 18 were determined, in which the inset shows that kobs is proportional to the inhibitor concentration with the slope and the intercept giving k+0 (=0.00066 ± 0.00001 s−1nM−1) and k−0 (=0.00271 ± 0.00031 s−1), respectively23. The inhibition constant for compound 18 can be calculated as Ki = k−0/k+0 = 4.1 ± 0.5 nM.
Next, we assessed the relationship between compound 18 and the other substrate, ubiquinol, by using the DBH2-cytochrome c system (Assay 3). At a fixed concentration of compound 18 and different concentrations of DBH2, the progress curves exhibited similar curvilinear patterns as observed in the assays at varying cytochrome c concentration, as a result of the slow onset of inhibition (Figs S4 and 4A). However, the kobs values decreased with increasing concentrations of DBH2 (inset of Fig. S4) in contrast to the observation in the inset of Fig. 4A. To achieve a comprehensive understanding, we followed the rate of product formation at various concentrations of DBH2 and compound 18. Two representative sets of curves were shown in Fig. 4C,D and the kobs values at the indicated DBH2 concentrations were determined by using the procedures described above. As shown in the insets, both sets of kobs values were proportional to the concentration of compound 18 and the slopes and intercepts of these straight lines denote the apparent association and dissociation constants (A and B, respectively) at various DBH2 concentrations23. The characteristic signature of decreasing A values and invariable B values with increasing concentrations of DBH2 suggests that compound 18 is a competitive inhibitor with respect to DBH2 (Fig. 4E,F). With the determined Km(DBH2) = 80.0 μM, the microscopic inhibition rate constants k+0 = 0.00038 ± 0.00001 s−1nM−1 and k−0 = 0.00509 ± 0.00009 s−1 and inhibition constant Ki = k−0/k+0 = 13.4 ± 0.6 nM can be derived. Together, these kinetic analyses suggest that compound 18 is a slow-binding inhibitor of bc1 complex and is non-competitive with substrate cytochrome c but competitive with respect to DBH2.
Structural Basis of Cytochrome bc1 Complex
Elucidation of the structural basis of compound 18 inhibition is helpful to further understand its action mechanism and to validate the reliability of the predicted binding modes of inhibitors. Hence, we determined the crystal structures of compound 18 bounds to chicken bc1 complex at a resolution of 3.23 Å. The binding mode of compound 18 is clearly defined by the electron density map in Fig. S5. According to the crystal structure, compound 18 fits well in the pocket formed by side chains Phe275, Phe129, Ile147, Pro271, Glu272, Leu295, Met125 and Val299. The structural similarity between the predicted and X-ray crystal binding models was found to be RMSD = 0.635 Å (Fig. S6), further validated the prediction capability of CSO method.
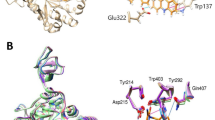
A structural comparison of the binding modes of compound 18 and AZ has revealed interesting similarities and differences. The overall arrangement of the compound 18-bc1 complex resembles the structure of the AZ-bc1 complex with no large conformational changes of the pocket, especially the side chain of the surrounding residues (Fig. 5A,B). Compound 18 mimics the binding mode of AZ, with the methoxyacrylate and the bridging phenyl ring overlapping extensively with that of AZ, both of which bind in a sub-pocket formed by Tyr132, Glu272, Pro271, Tyr279 and Gly143 (Fig. 5C,D). However, notable differences appear in the binding mode of the side chain. The π-π interactions between Phe275 and the benzothiazole side-chain group of compound 18 are improved greatly, relative to that of AZ, because of its large co-planar structure (Fig. 5E,F), which can induce ~73-fold improvement of the in vitro activity based on the Ki value of AZ.
Comparison of binding between compound 18 and AZ.
(A) Detailed description of the interactions of compound 18 generated with the program Ligplot. Hemispheres represent hydrophobic interactions, whereas lines represent polar interactions. (B) Detailed description of the interactions of AZ generated with the program Ligplot. (C) The methoxyacrylate and the bridging phenyl ring of compound 18 that binds in a sub-pocket formed by Tyr132, Glu272, Pro271, Tyr279 and Gly143. (D) The methoxyacrylate and the bridging phenyl ring of AZ binding in the same sub-pocket with a similar mode. (E) The π-π interactions between Phe275 and the benzothiazole side-chain group of compound 18. (F) Distorted π-π interactions between Phe275 and the side-chain group of AZ. Stereo views of the X-ray structure of compound 18 bound to chicken bc1 complex are available in Fig. S5.
Discussion
An effective tool to eliminate undesirable compounds prior to expensive and extensive experimental work is crucial for improving the efficiency of H2L optimization. Upon this challenge, CSO method was developed and proved to be an effective method to improve the efficiency of H2L process. Compared with the traditional methods, the CSO method has several advantages. First, CSO method can significantly reduce the numbers of compounds, need to be synthesized. For the present study, we need to synthesize 80 compounds theoretically, but only 21 compounds were synthesized (actually, only ten compounds needed to be synthesized and the other 11 compounds were just for testing) and 12 compounds with nanomolar to subnanomolar potency were obtained. That means, taking the present study as an example, CSO method improved the efficiency of H2L by about 75%. Second, the CSO method is independent of biological data and molecular descriptors and has the ability to predict a more expansive chemical space. The high correlation of ΔΔG (r2 = 0.8 for the linear correlation) and the high correspondence of the predicted and crystallographic poses (RMSD = 0.635 Å) support the reliability and accuracy of the CSO method. On the contrary, the traditional QSAR models are highly dependent on biological data and molecular descriptors and always used to explain the existing results and have very poor ability to predict the results outside the training sets. Third, because the CSO method is based on the assumption that the binding modes of substituted hits are similar to that of the original hit, it is possible to perform adequate conformational sampling based on the energy-minimized proteins in complex with substituted hits. Therefore, the CSO method is significantly time-saving compared with MD models.
It should be noted that the CSO method also have limitations. Clearly, this method is based on the assumption that the binding modes of substituted hits are similar to that of the original hit. This assumption is reasonable for small substituents. For a more bulky substituent, it may cause a considerable change in the binding mode and hence leads to significantly overestimate of the binding free-energy shift. Therefore, further improvement should be done to make CSO method suitable for broader application (more bulky groups).
The application of CSO method resulted in a candidate (compound 18). Compared with the commercial fungicide (AZ), the candidate has several advantages as follows: First, compound 18 has a high inhibiting potency for bc1 complex (Ki = 4.10 nM), which is up to ~73-fold more potent than AZ (Ki = 297.6 nM). In the view of ligand efficiency (LE)25, compound 18 (LE = 0.45 kcal mol−1 per non-H atom) is prior to AZ (LE = 0.3 kcal mol−1 per non-H atom). Second, compound 18 exhibited stronger antifungal activity against downy mildew and powdery mildew than AZ. In the greenhouse test and field trial, the protective and curative effect of compound 18 against downy mildew and powdery mildew are much higher than that of AZ. Third, the inhibitory kinetics studies showed that compound 18 exhibit slow-binding characteristics, which is great different from the classical fast-binding of AZ. This significant characteristic may help to overcome the increasing resistance problem that AZ has to face. Finally, the chemical structure of compound 18 is much simple and easily to be synthesized. The successful design of a highly efficient fungicide candidate also demonstrates a promising future of CSO method. In fact, the commercial development of compound 18 in China is in progress.
Materials and Methods
Computational protocol
Based on the modification and combination of AutoGrow and the Amber 9.0 program26,27, the CSO protocol was designed to perform automatically computational substitution, energy minimization and binding affinity evaluation. Depicted in Fig. 1 is the workflow of the CSO calculations.
Kinetic assays
The porcine succinate-cytochrome c reductase (SCR, the mixture of complex II and bc1) was prepared essentially according to the previously reported method28. The enzymatic activities of SCR, complex II and bc1 complex were analyzed in respective reaction mixtures as reported previously29. Kinetic analyses for the inhibition mechanism were performed as previously described23.
Greenhouse Fungicidal Activity
The fungicidal activity of compound 18 against P. cubensis and S. fuliginea was evaluated according to a previously published method30 and a potted-plant test method was adopted.
Field Trials
In the field trials, summer squash plants which is naturally infected by S. fuliginea and cucumber plants which is naturally infected by P. cubensis were used.
Crystallization and structure determination
Orthorhombic crystal of chicken bc1 in the space group P212121, containing a dimer in the asymmetric unit was prepared under optimized initial crystallization conditions. The structure was validated using the online tools at the molprobity site31 and deposited at the protein data bank with ID 4U3F.
Additional Information
How to cite this article: Hao, G.-F. et al. Rational Design of Highly Potent and Slow-Binding Cytochrome bc1 Inhibitor as Fungicide by Computational Substitution Optimization. Sci. Rep. 5, 13471; doi: 10.1038/srep13471 (2015).
References
Guan, A. Y., Liu, C. L., Yang, X. P. & Dekeyser, M. Application of the Intermediate Derivatization Approach in Agrochemical Discovery. Chem Rev 114, 7079–7107 (2014).
Lamberth, C., Jeanmart, S., Luksch, T. & Plant, A. Current Challenges and Trends in the Discovery of Agrochemicals. Science 341, 742–746 (2013).
Delaney, J., Clarke, E., Hughes, D. & Rice, M. Modern agrochemical research: a missed opportunity for drug discovery? Drug Discov Today 11, 839–845 (2006).
Keseru, G. M. & Makara, G. M. Hit discovery and hit-to-lead approaches. Drug Discov Today 11, 741–748 (2006).
Morley, A. D. et al. Fragment-based hit identification: thinking in 3D. Drug Discov Today 18, 1221–1227 (2013).
Whalen, K. L., Chau, A. C. & Spies, M. A. In silico Optimization of a Fragment-Based Hit Yields Biologically Active, High-Efficiency Inhibitors for Glutamate Racemase. ChemMedChem 8, 1681–1689 (2013).
Wunberg, T. et al. Improving the hit-to-lead process: data-driven assessment of drug-like and lead-like screening hits. Drug Discov Today 11, 175–180 (2006).
Topliss, J. G. & Edwards, R. P. Chance factors in studies of quantitative structure-activity relationships. J Med Chem 22, 1238–1244 (1979).
Zhao, H. & Caflisch, A. Molecular dynamics in drug design. Eur J Med Chem 91, 4–14 (2015).
Nichols, S. E., Swift, R. V. & Amaro, R. E. Rational Prediction with Molecular Dynamics for Hit Identification. Curr Top Med Chem 12, 2002–2012 (2012).
Poulain, R. et al. From hit to lead. Analyzing structure-profile relationships. J Med Chem 44, 3391–3401 (2001).
Manly, C. J., Chandrasekhar, J., Ochterski, J. W., Hammer, J. D. & Warfield, B. B. Strategies and tactics for optimizing the Hit-to-Lead process and beyond—A computational chemistry perspective. Drug Discov Today 13, 99–109 (2008).
Barton, V., Fisher, N., Biagini, G. A., Ward, S. A. & O’Neill, P. M. Inhibiting Plasmodium cytochrome bc1: a complex issue. Curr Opin Chem Biol 14, 440–446 (2010).
Birth, D., Kao, W.-C. & Hunte, C. Structural analysis of atovaquone-inhibited cytochrome bc1 complex reveals the molecular basis of antimalarial drug action. Nat Commun 5, 4029 (2014).
Hudson, A. T. Atovaquone-a novel broad-spectrum anti-infective drug. Parasitol Today 9, 66–68 (1993).
Laufer, M. K. et al. A Longitudinal Trial Comparing Chloroquine as Monotherapy or in Combination with Artesunate, Azithromycin or Atovaquone-Proguanil to Treat Malaria. PLoS ONE 7, e42284 (2012).
Petersen, I., Eastman, R. & Lanzer, M. Drug-resistant malaria: Molecular mechanisms and implications for public health. FEBS Lett 585, 1551–1562 (2011).
Leadbeater, A. & Gisi, U. The Challenges of Chemical Control of Plant Diseases. in Recent Developments in Management of Plant Diseases. Vol. 1 3–17 (Springer-Verlag Berlin, Heidelberger Platz 3, D-14197 Berlin, Germany, 2009).
Mavroeidi, V. I. & Shaw, M. W. Sensitivity distributions and cross-resistance patterns of Mycosphaerella graminicola to fluquinconazole, prochloraz and azoxystrobin over a period of 9 years. Crop Prot 24, 259–266 (2005).
Cape, J. L., Bowman, M. K. & Kramer, D. M. A semiquinone intermediate generated at the Q(o) site of the cytochrome bc(1), complex: Importance for the Q-cycle and superoxide production. Proc Natl Acad Sci USA 104, 7887–7892 (2007).
Hao, G. F. et al. Computational discovery of picomolar Q(o) site inhibitors of cytochrome bc1 complex. J Am Chem Soc 134, 11168–11176 (2012).
Wheeler, S. E. & Houk, K. N. Substituent Effects in the Benzene Dimer are Due to Direct Interactions of the Substituents with the Unsubstituted Benzene. J Am Chem Soc 130, 10854–10855 (2008).
Zhao, P. L. et al. Subnanomolar inhibitor of cytochrome bc1 complex designed by optimizing interaction with conformationally flexible residues. J Am Chem Soc 132, 185–194 (2010).
Hao, G., Dong, Q. & Yang, G. A Comparative Study on the Constitutive Properties of Marketed Pesticides. Mol Inform 30, 614–622 (2011).
Hopkins, A. L., Groom, C. R. & Alex, A. Ligand efficiency: a useful metric for lead selection. Drug Discov Today 9, 430–431 (2004).
Case, D. A. et al. AMBER9. (University of California, San Francisco, CA, 2006).
Durrant, J. D., Amaro, R. E. & McCammon, J. A. AutoGrow: a novel algorithm for protein inhibitor design. Chem Biol Drug Des 73, 168–178 (2009).
Yu, L. & Yu, C. A. Quantitative resolution of succinate-cytochrome c reductase into succinate-ubiquinone and ubiquinol-cytochrome c reductases. J Biol Chem 257, 2016–2021 (1982).
Fisher, N. et al. Modeling the Qo site of crop pathogens in Saccharomyces cerevisiae cytochrome b. Eur J Biochem 271, 2264–2271 (2004).
Wang, B.-L. et al. Synthesis and Biological Activity of Some Novel Trifluoromethyl-Substituted 1,2,4-Triazole and Bis(1,2,4-Triazole) Mannich Bases Containing Piperazine Rings. J Agric Food Chem 58, 5515–5522 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr Sect D: Biol Crystallogr 66, 12–21 (2010).
Acknowledgements
We would like to thank Prof. Baoju Li at The Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences and Prof. Jie Chen at ZheJiang Research Institute of Chemical Industry for help with the in vivo antifungal activity assay. We are grateful to the beamline staff at CHESS for help in setting up the data collection. The research was supported in part by the National Key Technologies R&D Program (2011BAE06B05) and the NSFC (No. 21332004).
Author information
Authors and Affiliations
Contributions
G.F.Y. conceived and designed the study. J.W.W. designed the kinetic assays experiment and analyzed the data. E.A.B. designed the crystallization experiment and analyzed the data. G.F.H. and S.G.Y. developed the computational protocol and performed the computational design. W.H. and W.L.T. performed organic synthesis. L.W., Y.Q.S. and H.L. performed kinetic assays experiment. L.S.H. performed crystallization experiment. G.F.Y. and G.F.H. analyzed the data and contributed in manuscript writing. G.F.Y., J.W.W., E.A.B. and G.F.H. made contributions in interpreting results and writing improvement of this paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hao, GF., Yang, SG., Huang, W. et al. Rational Design of Highly Potent and Slow-Binding Cytochrome bc1 Inhibitor as Fungicide by Computational Substitution Optimization. Sci Rep 5, 13471 (2015). https://doi.org/10.1038/srep13471
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13471
This article is cited by
-
Drug repurposing strategy part 1: from approved drugs to agri-bactericides leads
The Journal of Antibiotics (2023)
-
Supramolecular associations between atypical oxidative phosphorylation complexes of Euglena gracilis
Journal of Bioenergetics and Biomembranes (2021)
-
Synthesis and Herbicidal Activity of Novel β-Methoxyacrylate Derivatives Containing a Substituted Phenylpyridine Moiety
Chemical Research in Chinese Universities (2019)
-
The atypical subunit composition of respiratory complexes I and IV is associated with original extra structural domains in Euglena gracilis
Scientific Reports (2018)
-
Identification of a New Isoindole-2-yl Scaffold as a Qo and Qi Dual Inhibitor of Cytochrome bc 1 Complex: Virtual Screening, Synthesis, and Biochemical Assay
Interdisciplinary Sciences: Computational Life Sciences (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.