Abstract
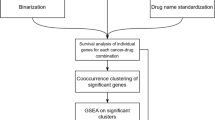
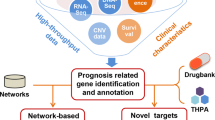
Screening for drug compounds that exhibit therapeutic properties in the treatment of various diseases remains a challenge even after considerable advancements in biomedical research. Here, we introduce an integrated platform that exploits gene expression compendia generated from drug-treated cell lines and primary tumor tissue to identify therapeutic candidates that can be used in the treatment of acute myeloid leukemia (AML). Our framework combines these data with patient survival information to identify potential candidates that presumably have a significant impact on AML patient survival. We use a drug regulatory score (DRS) to measure the similarity between drug-induced cell line and patient tumor gene expression profiles, and show that these computed scores are highly correlated with in vitro metrics of pharmacological activity. Furthermore, we conducted several in vivo validation experiments of our potential candidate drugs in AML mouse models to demonstrate the accuracy of our in silico predictions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hurle MR, Yang L, Xie Q, Rajpal DK, Sanseau P, Agarwal P . Computational drug repositioning: from data to therapeutics. Clin Pharmacol Ther 2013; 93: 335–341.
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006; 313: 1929–1935.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 2013; 41: D955–D961.
Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 2012; 150: 264–278.
Burnett A, Wetzler M, Lowenberg B . Therapeutic advances in acute myeloid leukemia. J Clin Oncol 2011; 29: 487–494.
Iorio F, Bosotti R, Scacheri E, Belcastro V, Mithbaokar P, Ferriero R et al. Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc Natl Acad Sci USA 2010; 107: 14621–14626.
Hassane DC, Guzman ML, Corbett C, Li X, Abboud R, Young F et al. Discovery of agents that eradicate leukemia stem cells using an in silico screen of public gene expression data. Blood 2008; 111: 5654–5662.
Sirota M, Dudley JT, Kim J, Chiang AP, Morgan AA, Sweet-Cordero et al. Discovery and preclinical validation of drug indications using compendia of public gene expression data. Sci Transl Med 2011; 3: 96ra77.
Dudley JT, Sirota M, Shenoy M, Pai RK, Roedder S, Chiang AP et al. Computational repositioning of the anticonvulsant topiramate for inflammatory bowel disease. Sci Transl Med 2011; 3 96ra76.
Wang K, Sun J, Zhou S, Wan C, Qin S, Li C et al. Prediction of drug-target interactions for drug repositioning only based on genomic expression similarity. PLoS Comput Biol 2013; 9: e1003315.
Pacini C, Iorio F, Goncalves E, Iskar M, Klabunde T, Bork P et al. DvD: an R/Cytoscape pipeline for drug repurposing using public repositories of gene expression data. Bioinformatics 2013; 29: 132–134.
Ung MH, Varn FS, Cheng C . IDEA: integrated drug expression analysis - integration of gene expression and clinical data for the identification of therapeutic candidates. CPT Pharmacometrics Syst Pharmacol 2015; 4: 415–425.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003; 4: 249–264.
Herold T, Metzeler KH, Vosberg S, Hartmann L, Rollig C, Stolzel F et al. Isolated trisomy 13 defines a homogeneous AML subgroup with high frequency of mutations in spliceosome genes and poor prognosis. Blood 2014; 124: 1304–1311.
Appelbaum FR, Gundacker H, Head DR, Slovak ML, Willman CL, Godwin JE et al. Age and acute myeloid leukemia. Blood 2006; 107: 3481–3485.
Wilson CS, Davidson GS, Martin SB, Andries E, Potter J, Harvey R et al. Gene expression profiling of adult acute myeloid leukemia identifies novel biologic clusters for risk classification and outcome prediction. Blood 2006; 108: 685–696.
Valk PJ, Verhaak RG, Beijen MA, Erpelinck CA, Barjesteh van Waalwijk van Doorn-Khosrovani S, Boer JM et al. Prognostically useful gene-expression profiles in acute myeloid leukemia. N Engl J Med 2004; 350: 1617–1628.
Wouters BJ, Lowenberg B, Erpelinck-Verschueren CA, van Putten WL, Valk PJ, Delwel R . Double CEBPA mutations, but not single CEBPA mutations, define a subgroup of acute myeloid leukemia with a distinctive gene expression profile that is uniquely associated with a favorable outcome. Blood 2009; 113: 3088–3091.
Lo M, Ling V, Low C, Wang YZ, Gout PW . Potential use of the anti-inflammatory drug, sulfasalazine, for targeted therapy of pancreatic cancer. Curr Oncol 2010; 17: 9–16.
Kannen V, Hintzsche H, Zanette DL, Silva WA Jr., Garcia SB, Waaga-Gasser AM et al. Antiproliferative effects of fluoxetine on colon cancer cells and in a colonic carcinogen mouse model. PLoS One 2012; 7: e50043.
Pisha E, Chai H, Lee IS, Chagwedera TE, Farnsworth NR, Cordell GA et al. Discovery of betulinic acid as a selective inhibitor of human melanoma that functions by induction of apoptosis. Nat Med 1995; 1: 1046–1051.
Massari NA, Medina VA, Cricco GP, Martinel Lamas DJ, Sambuco L, Pagotto R et al. Antitumor activity of histamine and clozapine in a mouse experimental model of human melanoma. J Dermatol Sci 2013; 72: 252–262.
Elso CM, Roberts LJ, Smyth GK, Thomson RJ, Baldwin TM, Foote SJ et al. Leishmaniasis host response loci (lmr1-3) modify disease severity through a Th1/Th2-independent pathway. Genes Immun 2004; 5: 93–100.
Baldwin T, Sakthianandeswaren A, Curtis JM, Kumar B, Smyth GK, Foote SJ et al. Wound healing response is a major contributor to the severity of cutaneous leishmaniasis in the ear model of infection. Parasite Immunol 2007; 29: 501–513.
Cox DR, Oakes D . Analysis of Survival Data. Chapman and Hall: London, New York, 1984 viii, 201 p.pp.
Hochberg Y, Benjamini Y . More powerful procedures for multiple significance testing. Stat Med 1990; 9: 811–818.
Huang, da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44–57.
Huang, da W, Sherman BT, Lempicki RA . Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2009; 37: 1–13.
Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res 2014; 42: D1091–D1097.
Vigushin DM, Ali S, Pace PE, Mirsaidi N, Ito K, Adcock I et al. Trichostatin A is a histone deacetylase inhibitor with potent antitumor activity against breast cancer in vivo. Clin Cancer Res 2001; 7: 971–976.
Johnstone RW . Histone-deacetylase inhibitors: novel drugs for the treatment of cancer. Nat Rev Drug Discov 2002; 1: 287–299.
Shaker S, Bernstein M, Momparler LF, Momparler RL . Preclinical evaluation of antineoplastic activity of inhibitors of DNA methylation (5-aza-2'-deoxycytidine) and histone deacetylation (trichostatin A, depsipeptide) in combination against myeloid leukemic cells. Leukemia Res 2003; 27: 437–444.
Mrozek K, Heinonen K, de la Chapelle A, Bloomfield CD . Clinical significance of cytogenetics in acute myeloid leukemia. Semin Oncol 1997; 24: 17–31.
Slovak ML, Kopecky KJ, Cassileth PA, Harrington DH, Theil KS, Mohamed et al. Karyotypic analysis predicts outcome of preremission and postremission therapy in adult acute myeloid leukemia: a Southwest Oncology Group/Eastern Cooperative Oncology Group Study. Blood 2000; 96: 4075–4083.
Zhou Y, Zhang Q, Stephens O, Heuck CJ, Tian E, Sawyer JR et al. Prediction of cytogenetic abnormalities with gene expression profiles. Blood 2012; 119: e148–e150.
Merino D, Khaw SL, Glaser SP, Anderson DJ, Belmont LD, Wong C et al. Bcl-2, Bcl-x(L), and Bcl-w are not equivalent targets of ABT-737 and navitoclax (ABT-263) in lymphoid and leukemic cells. Blood 2012; 119: 5807–5816.
Tahir SK, Wass J, Joseph MK, Devanarayan V, Hessler P, Zhang H et al. Identification of expression signatures predictive of sensitivity to the Bcl-2 family member inhibitor ABT-263 in small cell lung carcinoma and leukemia/lymphoma cell lines. Mol Cancer Ther 2010; 9: 545–557.
Yu C, Dasmahapatra G, Dent P, Grant S . Synergistic interactions between MEK1/2 and histone deacetylase inhibitors in BCR/ABL+ human leukemia cells. Leukemia 2005; 19: 1579–1589.
Ozaki K, Kosugi M, Baba N, Fujio K, Sakamoto T, Kimura S et al. Blockade of the ERK or PI3K-Akt signaling pathway enhances the cytotoxicity of histone deacetylase inhibitors in tumor cells resistant to gefitinib or imatinib. Biochem Biophys Res Commun 2010; 391: 1610–1615.
Chang-Yew Leow C, Gerondakis S, Spencer A . MEK inhibitors as a chemotherapeutic intervention in multiple myeloma. Blood Cancer J 2013; 3: e105.
Nishioka C, Ikezoe T, Yang J, Koeffler HP, Yokoyama A . Inhibition of MEK/ERK signaling synergistically potentiates histone deacetylase inhibitor-induced growth arrest, apoptosis and acetylation of histone H3 on p21waf1 promoter in acute myelogenous leukemia cell. Leukemia 2008; 22: 1449–1452.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25: 25–29.
Gupta AK, Grober JS, Hamilton TA, Ellis CN, Siegel MT, Voorhees JJ et al. Sulfasalazine therapy for psoriatic arthritis: a double blind, placebo controlled trial. J Rheumatol 1995; 22: 894–898.
Wong DT, Bymaster FP, Engleman EA . Prozac (fluoxetine, Lilly 110140), the first selective serotonin uptake inhibitor and an antidepressant drug: twenty years since its first publication. Life Sci 1995; 57: 411–441.
Kane J, Honigfeld G, Singer J, Meltzer H . Clozapine for the treatment-resistant schizophrenic. A double-blind comparison with chlorpromazine. Arch Gen Psychiatry 1988; 45: 789–796.
Chowdhury AR, Mandal S, Mittra B, Sharma S, Mukhopadhyay S, Majumder HK . Betulinic acid, a potent inhibitor of eukaryotic topoisomerase I: identification of the inhibitory step, the major functional group responsible and development of more potent derivatives. Med Sci Monit 2002; 8: BR254–BR265.
Campoli-Richards DM, Lackner TE, Monk JP . Ceforanide. A review of its antibacterial activity, pharmacokinetic properties and clinical efficacy. Drugs 1987; 34: 411–437.
Acknowledgements
We would like to thank Tobias Herold, Wolfgang Hiddemann, Thomas Büchner, Karsten Spiekermann, Stephanie Schneider and Maria Cristina Sauerland for providing us with the patient survival information for the Herold data set. We also thank Roel Veerhak for providing us with the patient survival information for the Wouter’s data set. This work was supported by the American Cancer Society Research Grant, #IRG-82-003-30, the National Center For Advancing Translational Sciences of the National Institutes of Health under Award Number UL1TR001086, and by the start-up funding package provided to CC by the Geisel School of Medicine at Dartmouth College.
Author contributions
CC, C-CL (Liu) and C-CL (Lin) conceived of the project; MHU, C-HS, C-WW, C-CL (Liu) and CC performed the computational drug prediction analysis; C-CH and C-CL (Lin) performed the mouse experiments; all authors contributed to the writing of the manuscript and interpretation of results.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Rights and permissions
About this article
Cite this article
Ung, M., Sun, CH., Weng, CW. et al. Integrated Drug Expression Analysis for leukemia: an integrated in silico and in vivo approach to drug discovery. Pharmacogenomics J 17, 351–359 (2017). https://doi.org/10.1038/tpj.2016.18
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2016.18
This article is cited by
-
An in-silico study of cancer cell survival and spatial distribution within a 3D microenvironment
Scientific Reports (2020)