Article
|
Open Access
Featured
-
-
Article
| Open Access DELVE: feature selection for preserving biological trajectories in single-cell data
DELVE: feature selection for preserving biological trajectories in single-cell data
Characteristic genes or proteins driving continuous biological processes are difficult to uncover from noisy single-cell data. Here, authors present DELVE, an unsupervised feature selection method to identify core molecular features driving cell fate decisions.
- Jolene S. Ranek
- , Wayne Stallaert
- & Jeremy E. Purvis
-
Article
| Open Access Multi-dimensional scaling techniques unveiled gain1q&loss13q co-occurrence in Multiple Myeloma patients with specific genomic, transcriptional and adverse clinical features
Multi-dimensional scaling techniques unveiled gain1q&loss13q co-occurrence in Multiple Myeloma patients with specific genomic, transcriptional and adverse clinical features
The characterisation of the molecular features of multiple myeloma (MM) remains challenging. Here, the authors identify a subset of MM patients with a dismal clinical outcome, harbouring both chromosomes 1q CN gain and 13 CN loss and overexpressing CCND2.
- Carolina Terragna
- , Andrea Poletti
- & Michele Cavo
-
Article
| Open Access PheWAS-based clustering of Mendelian Randomisation instruments reveals distinct mechanism-specific causal effects between obesity and educational attainment
PheWAS-based clustering of Mendelian Randomisation instruments reveals distinct mechanism-specific causal effects between obesity and educational attainment
Mendelian Randomisation estimates causal effects between risk factors and complex outcomes using genetic variants as instrumental variables, however it can be affected by certain biases. To alleviate these biases the authors propose an approach based on clustering genetic instruments according to the types of trait they are associated with, and apply this method to revisit the surprisingly large apparent causal effect of body mass index on educational attainment.
- Liza Darrous
- , Gibran Hemani
- & Zoltán Kutalik
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Acral melanoma (AM) is a rare melanoma subtype with unique features, where lymph node metastasis is closely associated with clinical outcomes. Here, the authors use single-cell and spatial transcriptomics to analyse early dissemination, tumour microenvironment, and heterogeneity in AM, and infer metabolic shifts with therapeutic implications.
- Chuanyuan Wei
- , Wei Sun
- & Jianying Gu
-
Article
| Open Access Dissecting the human leptomeninges at single-cell resolution
Dissecting the human leptomeninges at single-cell resolution
The meninges protect the central nervous system at the brain border, and its dysfunction can lead to neural inflammation and cell damage. Here, the authors uncover the gene signatures of diverse cell types in the aged human leptomeninges and highlight their changes in Alzheimer’s Disease.
- Nicola A. Kearns
- , Artemis Iatrou
- & Yanling Wang
-
Article
| Open Access Super-resolved trajectory-derived nanoclustering analysis using spatiotemporal indexing
Super-resolved trajectory-derived nanoclustering analysis using spatiotemporal indexing
Existing single-molecule localization microscopy analyses overlook important temporal information in living cells. Here, the authors report nanoscale spatiotemporal indexing clustering (NASTIC), which leverages a video game algorithm to fast-track the investigation of the complex temporal dynamics of protein clustering.
- Tristan P. Wallis
- , Anmin Jiang
- & Frédéric A. Meunier
-
Article
| Open Access Cell-attribute aware community detection improves differential abundance testing from single-cell RNA-Seq data
Cell-attribute aware community detection improves differential abundance testing from single-cell RNA-Seq data
Single-cell technologies allow quantification of cell-types in human tissues, yet detecting if their abundance changes in aging or disease is challenging. By using cell-attribute aware clustering, this work presents a differential abundance testing algorithm with increased power.
- Alok K. Maity
- & Andrew E. Teschendorff
-
Article
| Open Access ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
RNA-sequencing data analysis is difficult for non-model species that have no reference genome. ExpressAnalyst enables RNA-sequencing analysis for any eukaryotic species in less than 24 h, on a laptop, and without any programming.
- Peng Liu
- , Jessica Ewald
- & Jianguo Xia
-
Article
| Open Access Latent generative landscapes as maps of functional diversity in protein sequence space
Latent generative landscapes as maps of functional diversity in protein sequence space
In this work, the authors study protein families’ VAE latent manifolds and coevolutionary Hamiltonians. These Latent Generative Landscapes predict phylogenetic groupings, fitness & functional properties for several systems with clear protein engineering/design potential.
- Cheyenne Ziegler
- , Jonathan Martin
- & Faruck Morcos
-
Article
| Open Access Reversion mutations in germline BRCA1/2-mutant tumors reveal a BRCA-mediated phenotype in non-canonical histologies
Reversion mutations in germline BRCA1/2-mutant tumors reveal a BRCA-mediated phenotype in non-canonical histologies
Mutations in BRCA1/2 are associated with a homologous recombination deficiency phenotype in BRCA-associated cancers. Reversion mutations can restore BRCA1/2 function and result in treatment resistance in these cancer-types. Here, the authors show that, in select cases, reversion mutations in BRCA1/2 can indicate prior BRCA-mediated tumorigenesis in non-canonical histologies.
- Yonina R. Murciano-Goroff
- , Alison M. Schram
- & Alexander Drilon
-
Article
| Open Access Proteo-genomic characterization of virus-associated liver cancers reveals potential subtypes and therapeutic targets
Proteo-genomic characterization of virus-associated liver cancers reveals potential subtypes and therapeutic targets
The molecular heterogeneity of primary liver cancer remains to be characterised. Here, the authors perform multi-omics analysis of 259 primary liver cancer tissues and identify three distinct molecular subtypes.
- Masashi Fujita
- , Mei-Ju May Chen
- & Hidewaki Nakagawa
-
Article
| Open Access Alignment of single-cell trajectory trees with CAPITAL
Alignment of single-cell trajectory trees with CAPITAL
Global alignment of complex cell state trajectories between single-cell datasets remains challenging. Here, the authors present a computational method called CAPITAL to compare branching trajectories, and demonstrate that this method achieves accurate and robust alignments.
- Reiichi Sugihara
- , Yuki Kato
- & Yukio Kawahara
-
Article
| Open Access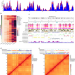 Regulation associated modules reflect 3D genome modularity associated with chromatin activity
Regulation associated modules reflect 3D genome modularity associated with chromatin activity
Here the authors report histone modifications show a modular pattern referred to as ‘regulation associated modules’ (RAMs) that reflect the spatial modularity of chromatin. They find enhancer-promoter interactions and extrachromosomal DNAs (ecDNAs) occur more often within the same RAMs than within the same TADs, indicating stronger insulation of the RAM boundaries.
- Lina Zheng
- & Wei Wang
-
Article
| Open Access Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
In the era of single-cell sequencing, there is a growing need to extract insights from data with clustering methods. Here, inspired by forest fire dynamics, the authors devise an algorithm that can cluster single-cell data with minimal prior assumptions and can compute a non-parametric posterior probability for each data point.
- Zhanlin Chen
- , Jeremy Goldwasser
- & Mark Gerstein
-
Article
| Open Access Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder
Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder
Breakthrough technologies for spatially resolved transcriptomics have enabled genome-wide profiling of gene expressions in captured locations. Here the authors integrate gene expressions and spatial locations to identify spatial domains using an adaptive graph attention auto-encoder.
- Kangning Dong
- & Shihua Zhang
-
Article
| Open Access Trajectory of immune evasion and cancer progression in hepatocellular carcinoma
Trajectory of immune evasion and cancer progression in hepatocellular carcinoma
In order to design cancer immune therapies, it is important to understand how tumours evade the immune response that is mounted against them. Authors here analyse the distribution and properties of immune cells in hepatocellular carcinoma and describe a progressive tumour-immune co-evolution programme from early to late stage cancer.
- Phuong H. D. Nguyen
- , Martin Wasser
- & Valerie Chew
-
Article
| Open Access Pluripotency factors determine gene expression repertoire at zygotic genome activation
Pluripotency factors determine gene expression repertoire at zygotic genome activation
Zygotic genome activation in zebrafish relies on pluripotency transcription factors Pou5f3 and Sox19b. Here the authors investigate how these factors interact in vivo by analyzing the changes in chromatin state and time-resolved transcription in Pou5f3 and Sox19b single and double mutant embryos.
- Meijiang Gao
- , Marina Veil
- & Daria Onichtchouk
-
Article
| Open Access Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Idiopathic pulmonary arterial hypertension is a rare and fatal disease with a heterogeneous treatment response. Here the authors show that unsupervised machine learning of whole blood transcriptomes from 359 patients with idiopathic pulmonary arterial hypertension identifies 3 subgroups (endophenotypes) that improve risk stratification and provide new molecular insights.
- Sokratis Kariotis
- , Emmanuel Jammeh
- & Richard C. Trembath
-
Article
| Open Access Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Normalisr removes technical bias in single-cell RNA-seq and detects gene differential and coexpression accurately and efficiently. It also infers gene regulatory and co-expression networks from conventional and CRISPR screen single-cell RNA-seq datasets.
- Lingfei Wang
-
Article
| Open Access Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions
Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions
While transcriptomics have enhanced our understanding for cancer, spatial transcriptomics enable the characterisation of cellular interactions. Here, the authors integrate single cell data with spatial information for HER2 + tumours and develop tools for the prediction of interactions between tumour-infiltrating cells.
- Alma Andersson
- , Ludvig Larsson
- & Joakim Lundeberg
-
Article
| Open Access DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
Cell-type-specific genes are often strongly correlated in expression - an informative yet underexplored property of single-cell data. Here, the authors leverage gene expression correlations to develop DUBStepR, a feature selection method for accurately clustering single-cell data.
- Bobby Ranjan
- , Wenjie Sun
- & Shyam Prabhakar
-
Article
| Open Access Learning interpretable cellular and gene signature embeddings from single-cell transcriptomic data
Learning interpretable cellular and gene signature embeddings from single-cell transcriptomic data
Computational single-cell RNA-seq analyses often face challenges in scalability, model interpretability, and confounders. Here, we show a new model to address these challenges by learning meaningful embeddings from the data that simultaneously refine gene signatures and cell functions in diverse conditions.
- Yifan Zhao
- , Huiyu Cai
- & Yue Li
-
Article
| Open Access Towards omics-based predictions of planktonic functional composition from environmental data
Towards omics-based predictions of planktonic functional composition from environmental data
Advances in omics approaches could enable quantitative predictions of microbial functional composition. Here the authors re-analyze 885 metagenome-assembled genomes from Tara Oceans, and use a network approach to quantify protein functional clusters and explore their biogeography.
- Emile Faure
- , Sakina-Dorothée Ayata
- & Lucie Bittner
-
Article
| Open Access Coordination of endothelial cell positioning and fate specification by the epicardium
Coordination of endothelial cell positioning and fate specification by the epicardium
It remains unclear how spatial information controls endothelial cell identity and behavior in the developing heart. Here the authors perform single cell RNA sequencing at key developmental timepoints in mice to interrogate cellular contributions to coronary vessel patterning and maturation in the epicardium.
- Pearl Quijada
- , Michael A. Trembley
- & Eric M. Small
-
Article
| Open Access Chromatin states shaped by an epigenetic code confer regenerative potential to the mouse liver
Chromatin states shaped by an epigenetic code confer regenerative potential to the mouse liver
Few studies have provided functional analysis of the epigenetic landscape in the regenerating liver. Here the authors define chromatin states in the quiescent vs. regenerating mouse liver through integration of genome wide profiles of DNA methylation, histone modifications, and chromatin accessibility, identifying H3K27me3 as an epigenetic mark conferring regenerative potential.
- Chi Zhang
- , Filippo Macchi
- & Kirsten C. Sadler
-
Article
| Open Access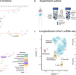 Evolution of core archetypal phenotypes in progressive high grade serous ovarian cancer
Evolution of core archetypal phenotypes in progressive high grade serous ovarian cancer
High-grade serous ovarian cancer (HGSOC) is prone to developing resistance to treatment. Here, the authors use single-cell RNA-seq and an analysis of archetypes, and find that shifts in metabolism and proliferation are associated with the response to treatment and clonal heterogeneity in HGSOC.
- Aritro Nath
- , Patrick A. Cosgrove
- & Andrea H. Bild
-
Article
| Open Access DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires
The advent of high-throughput T-cell receptor sequencing has allowed for the rapid and thorough characterization of the adaptive immune response. Here the authors show how deep learning can reveal both descriptive and predictive sequence concepts within the immune repertoire.
- John-William Sidhom
- , H. Benjamin Larman
- & Alexander S. Baras
-
Article
| Open Access Improving gene function predictions using independent transcriptional components
Improving gene function predictions using independent transcriptional components
Our understanding of the function of many transcripts is still incomplete, limiting the interpretability of transcriptomic data. Here the authors use consensus-independent component analysis, together with a guilt-by-association approach, to improve the prediction of gene function.
- Carlos G. Urzúa-Traslaviña
- , Vincent C. Leeuwenburgh
- & Rudolf S. N. Fehrmann
-
Article
| Open Access Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations
Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations
Genomic prediction of phenotype may be improved by using DNA mutations with functional, evolutionary, and pleiotropic consequences. Here the authors describe a method for genome-wide fine-mapping of QTLs and develop a genotyping array for improved prediction of genetic values for cattle traits.
- Ruidong Xiang
- , Iona M. MacLeod
- & Michael E. Goddard
-
Article
| Open Access Dissecting the cellular specificity of smoking effects and reconstructing lineages in the human airway epithelium
Dissecting the cellular specificity of smoking effects and reconstructing lineages in the human airway epithelium
Chronic lung diseases are characterized by molecular and cellular composition changes. Here the authors use single-cell RNA sequencing to map cell type-specific changes in human tracheal epithelium related to smoking, and to provide evidence for a tuft-like progenitor for pulmonary neuroendocrine cells and ionocytes.
- Katherine C. Goldfarbmuren
- , Nathan D. Jackson
- & Max A. Seibold
-
Article
| Open Access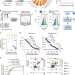 Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
5p and 3p miRNA strands have different mRNA-targeting sequences and may both functionally impact gene expression in cancer. Here, the authors undertake a pan-cancer analysis that indicates 5p/3p miRNA strands function together to regulate tumorigenic processes.
- Ramkrishna Mitra
- , Clare M. Adams
- & Christine M. Eischen
-
Article
| Open Access The SIGMA rat brain templates and atlases for multimodal MRI data analysis and visualization
The SIGMA rat brain templates and atlases for multimodal MRI data analysis and visualization
Magnetic resonance imaging (MRI) is widely used to study the rat brain. Here, the authors provide standardized MRI brain templates and descriptive atlases for the rat, incorporating both structural and functional MRI data, along with associated resources.
- D. A. Barrière
- , R. Magalhães
- & S. Mériaux
-
Article
| Open Access Macrophage-associated wound healing contributes to African green monkey SIV pathogenesis control
Macrophage-associated wound healing contributes to African green monkey SIV pathogenesis control
Here, the authors compare gene expression signatures in rectal tissues of African green monkeys (AGMs) and rhesus macaques (RMs) acutely infected with simian immunodeficiency virus and find that AGMs rapidly activate and maintain evolutionarily conserved regenerative wound healing mechanisms.
- Fredrik Barrenas
- , Kevin Raehtz
- & Michael Gale Jr
-
Article
| Open Access SCALE method for single-cell ATAC-seq analysis via latent feature extraction
SCALE method for single-cell ATAC-seq analysis via latent feature extraction
Single-cell ATAC-seq data is challenging to analyse for reasons such as high dimensionality and sparsity. Here, the authors develop SCALE, a deep learning method that leverages latent feature extraction for various tasks of scATACseq data analysis.
- Lei Xiong
- , Kui Xu
- & Qiangfeng Cliff Zhang
-
Article
| Open Access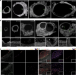 Single-cell reconstruction of follicular remodeling in the human adult ovary
Single-cell reconstruction of follicular remodeling in the human adult ovary
In the ovary, follicular degeneration occurs after folliculogenesis, with one dominant follicle reaching maturity every month. Here, by performing single cell RNA-seq of adult human follicles, the authors identify subclasses of granulosa and theca cell lineages, which correlate with the growth or degeneration of the follicles.
- X. Fan
- , M. Bialecka
- & S. M. Chuva de Sousa Lopes
-
Article
| Open Access Single cell RNA-seq and ATAC-seq analysis of cardiac progenitor cell transition states and lineage settlement
Single cell RNA-seq and ATAC-seq analysis of cardiac progenitor cell transition states and lineage settlement
Cardiac progenitor cells (CPCs) form cardiomyocytes, pericytes, smooth muscle and endothelial cells during embryonic development. Here, the authors characterize mouse CPCs marked by Nkx2.5 and Isl1 from E7.5 to E9.5 by single cell RNA-seq and ATAC-seq, showing fate transitions involve distinct open chromatin state.
- Guangshuai Jia
- , Jens Preussner
- & Thomas Braun
-
Article
| Open Access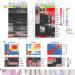 Single cell RNA-seq reveals profound transcriptional similarity between Barrett’s oesophagus and oesophageal submucosal glands
Single cell RNA-seq reveals profound transcriptional similarity between Barrett’s oesophagus and oesophageal submucosal glands
Barrett’s oesophagus is associated with an increased risk of oseophageal cancer, but its cell of origin is unclear. Here the authors show, using single-cell RNA sequencing of biopsies from six patients and two unaffected subjects, that cells in Barrett’s oesophagus show a transcriptional profile that is similar to that of cells in oesophageal submucosal glands.
- Richard Peter Owen
- , Michael Joseph White
- & Xin Lu
-
Article
| Open Access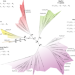 Broad phylogenetic analysis of cation/proton antiporters reveals transport determinants
Broad phylogenetic analysis of cation/proton antiporters reveals transport determinants
Cation/proton antiporters (CPAs) play a major role in maintaining living cells’ homeostasis and are divided in two main groups: CPA1 and CPA2. Here authors use a comprehensive evolutionary analysis of 6537 representative CPAs and reveal a sequence motif that determines central phenotypic characteristics.
- Gal Masrati
- , Manish Dwivedi
- & Nir Ben-Tal
-
Article
| Open Access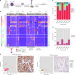 Unravelling subclonal heterogeneity and aggressive disease states in TNBC through single-cell RNA-seq
Unravelling subclonal heterogeneity and aggressive disease states in TNBC through single-cell RNA-seq
Triple-negative breast cancer is highly heterogeneous and aggressive. Here, the authors utilise single-cell RNA sequencing to investigate this heterogeneity, and discover a subpopulation of cells associated with metastasis and treatment resistance signatures, and linked to long term survival outcomes.
- Mihriban Karaayvaz
- , Simona Cristea
- & Leif W. Ellisen
-
Article
| Open Access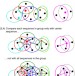 Clustering huge protein sequence sets in linear time
Clustering huge protein sequence sets in linear time
Billions of metagenomic and genomic sequences fill up public datasets, which makes similarity clustering an important and time-critical analysis step. Here, the authors develop Linclust, an algorithm with linear time complexity that can cluster over a billion sequences within hours on a single server.
- Martin Steinegger
- & Johannes Söding
-
Article
| Open Access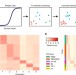 Unsupervised clustering and epigenetic classification of single cells
Unsupervised clustering and epigenetic classification of single cells
Single cell ATAC-seq (scATAC-seq) data reveals cellular level epigenetic heterogeneity but its application in delineating distinct subpopulations is still challenging. Here, the authors develop scABC, a statistical method for unsupervised clustering of scATAC-seq data and identification of open chromatin regions specific to cell identity.
- Mahdi Zamanighomi
- , Zhixiang Lin
- & Wing Hung Wong
-
Article
| Open Access Functionally distinct disease-associated fibroblast subsets in rheumatoid arthritis
Functionally distinct disease-associated fibroblast subsets in rheumatoid arthritis
Synovial fibroblasts are thought to be central mediators of joint destruction in rheumatoid arthritis (RA). Here the authors use single-cell transcriptomics and flow cytometry to identify synovial fibroblast subsets that are expanded and display distinct tissue distribution and function in patients with RA.
- Fumitaka Mizoguchi
- , Kamil Slowikowski
- & Michael B. Brenner
-
Article
| Open Access Evolutionary action and structural basis of the allosteric switch controlling β2AR functional selectivity
Evolutionary action and structural basis of the allosteric switch controlling β2AR functional selectivity
Ligand-induced biased signaling is thought to result in part from ligand-specific receptor conformations that cause the engagement of distinct effectors. Here the authors trace and evaluate the impact of mutations of the β2–adrenergic receptor on multiple signaling outputs to provide structural-level insight into the determinants of GPCR functional selectivity.
- Anne-Marie Schönegge
- , Jonathan Gallion
- & Michel Bouvier
-
Article
| Open Access Functional reconstruction of a eukaryotic-like E1/E2/(RING) E3 ubiquitylation cascade from an uncultured archaeon
Functional reconstruction of a eukaryotic-like E1/E2/(RING) E3 ubiquitylation cascade from an uncultured archaeon
In eukaryotic cells, the ubiquitylation system regulates several cellular processes central to protein homoeostasis. Here the authors demonstrate the existence of an eukaryotic-like ubiquitylation cascade requiring E1, E2 and E3-like enzymes in the archaeon C. subterraneum, shedding light on the evolution of the ubiquitin-proteasome system.
- Rory Hennell James
- , Eva F. Caceres
- & Nicholas P. Robinson
-
Article
| Open Access Integrative analysis of genomic and epigenomic regulation of the transcriptome in liver cancer
Integrative analysis of genomic and epigenomic regulation of the transcriptome in liver cancer
Hepatocellular carcinoma is known to harbour numerous genomic and epigenomic aberrations, driving transcriptomic deregulation. Here, the authors integrate genomic, epigenomic, and expression data to reveal three prognostic subtypes, providing insight to the pathobiology of hepatocellular carcinoma.
- Hyun Goo Woo
- , Ji-Hye Choi
- & Yoon Jun Kim
-
Article
| Open Access Defining functional interactions during biogenesis of epithelial junctions
Defining functional interactions during biogenesis of epithelial junctions
Formation and reinforcement of E-cadherin-mediated adhesion depends on intracellular trafficking and interactions with the actin cytoskeleton, but how these are coordinated is not known. Here the authors conduct a focused phenotypic screen to identify new pathways regulating cell–cell junction homeostasis.
- J. C. Erasmus
- , S. Bruche
- & V. M. M. Braga

 Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences in linear time by relatedness sorting