Abstract
Although often located at a distance from their target gene promoters, enhancers are the primary genomic determinants of temporal and spatial transcriptional specificity in metazoans. Since the discovery of the first enhancer element in simian virus 40, there has been substantial interest in unraveling the mechanism(s) by which enhancers communicate with their partner promoters to ensure proper gene expression. These research efforts have benefited considerably from the application of increasingly sophisticated sequencing- and imaging-based approaches in conjunction with innovative (epi)genome-editing technologies; however, despite various proposed models, the principles of enhancer–promoter interaction have still not been fully elucidated. In this review, we provide an overview of recent progress in the eukaryotic gene transcription field pertaining to enhancer–promoter specificity. A better understanding of the mechanistic basis of lineage- and context-dependent enhancer–promoter engagement, along with the continued identification of functional enhancers, will provide key insights into the spatiotemporal control of gene expression that can reveal therapeutic opportunities for a range of enhancer-related diseases.
Similar content being viewed by others
Introduction
Transcriptional regulation is a crucial feature of gene expression control during development, homeostasis, and signal-dependent responses. In eukaryotes, DNA sequences located proximal or distal to the transcription start site (TSS), referred to as cis-regulatory elements (CREs), play pivotal roles in the regulation of RNA polymerase II (Pol II)-dependent gene expression. The CRE that is closest to the TSS (typically <1 kb) is the promoter, which, when active, recruits transcriptional machinery and is marked by trimethylation of histone H3 on lysine 4 (H3K4me3). More remotely positioned CREs (>1 kb from the TSS) serve as enhancers, the active versions of which also feature open or accessible chromatin occupied by transcriptional complexes. Active enhancers often display higher levels of H3K4 monomethylation (H3K4me1) than promoters, and they tend to show differential engagement of lineage-determining and cell type-specific transcription factors (TFs), based on the presence of specific DNA motifs, in addition to certain transcriptional coactivators, such as p300 and MED11. Despite these differences, functional enhancers and promoters share some common characteristics, including nucleosome-free DNase hypersensitivity, enrichment of H3K27ac as well as other active histone marks, recruitment of general and specific TFs, and bidirectional transcription. Indeed, it is now apparent that the classification of promoters and enhancers, at least from a functional perspective, can be arbitrary2. Nevertheless, a comprehensive understanding of enhancer and promoter grammar and functionality, including their interplay, is necessary to elucidate gene regulatory strategies in normal physiology and potential pathological alterations.
A number of large-scale research projects have sought to catalog and characterize CREs. Notably, the ENCODE consortium has interrogated nearly a million putative CREs in the human genome by examining multiple epigenomic features3. Recent advances in single-cell technologies have allowed for the analysis of cell type-specific CREs. While application of single-cell ATAC-seq has proven particularly powerful for annotation of putative CREs, additional single-cell-based strategies have begun to provide more extensive epigenomic profiling of CREs in distinct cell types from various tissues during different developmental stages or in certain disease contexts, even though technical challenges regarding cell number and quality as well as in data processing remain4,5. In addition, high-throughput, sequencing-based reporter assays, such as self-transcribing active regulatory region sequencing (STARR-seq) and massively parallel reporter assays (MPRAs), have provided complementary insights into the functional potential and properties of natural as well as artificial CREs6.
Validation of putative CREs requires assignment of their target genes, which is critical for discerning any role in transcriptional regulation. Although this process is typically straightforward for promoter CREs, given their location at the TSS(s) of annotated genes, it is a considerably more challenging task for distal CREs that constitute enhancers. High-throughput derivatives of chromatin conformation capture (3 C), which detect physical contacts between genomic regions, and approaches based on expression quantitative trait loci (eQTL) that correlate genetic variants to gene transcript levels have helped in identifying instances of enhancer–promoter interaction (EPI), which is generally considered to be a key feature of enhancer-mediated regulation7,8. Nevertheless, a better understanding of the molecular mechanisms by which enhancers interact with their target genes to activate transcription is required to elucidate regulatory strategies. In this review, we provide an overview of recent insights into the molecular underpinnings of EPI in the context of 3D genome architecture and discuss the disease implications of EPI alterations. In addition, we briefly introduce the experimental methods used to uncover EPIs and comment on issues with data interpretation as well as discrepancies arising from different approaches.
Identification of enhancer-promoter interactions
It is now apparent that eukaryotic genomes are highly organized within the nucleus and that the prevailing 3D structure, which is relatively stable but amenable to alterations, impacts transcription and ultimately cellular phenotype. Fundamental features of genomic organization, including topologically associated domains (TADs) and the polymeric nature of chromatin, have been elucidated by utilization of 3C-based techniques combined with next-generation sequencing (NGS) and super-resolution microscopy tools9,10. EPIs occur within the context of hierarchical 3D genomic architecture. Although functional validation of most EPIs is lacking, the regulatory role of enhancer–promoter looping has been confirmed in many studies of individual loci and can be addressed in a high-throughput manner with clustered regularly interspaced short palindromic repeats (CRISPR)-based epigenome editing approaches.
3C-based assays and complementary approaches
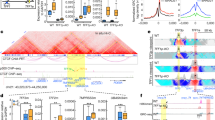
A large collection of methods has been introduced to enable the study of chromosome architecture that broadly includes 3C and its derivatives, which involve proximity ligation of digested chromosomes in crosslinked cells, as well as alternative procedures with sequencing readouts that do not rely on a ligation step, namely split-pool recognition of interactions by tag extension (SPRITE) and genome architecture mapping (GAM) (Fig. 1a)11. 3C-based techniques include 4C-seq and Hi-C, which has various derivatives itself, such as PLAC-seq, Capture-C, and micro-C11, and also has been optimized12. A crosslinking-independent, 3C-based assay, known as intrinsic 3C, has also been developed13. Collectively, these 3C-based techniques have revealed the presence of territories in metazoan chromosomes that are further organized into A/B compartments, TADs, and chromatin loops that include EPIs14. Despite substantial advancements in the understanding of 3D genome architecture afforded by 3C-based assays, proximity ligation methods have inherent limitations. First, in instances that involve contacts connecting three or more genomic loci, these methods cannot clearly distinguish whether the interaction is simultaneous within the same cell or occurs in a pairwise manner in different cells. Second, these strategies are not able to detect long-range DNA interactions that cannot be efficiently ligated15. To overcome these challenges, several modified procedures, such as C-walks (molecular barcoding), MC-4C (long-read sequencing), and Pore-C (nanopore sequencing), attempt to capture both pairwise and multiway chromosomal contacts by sequencing large proximity-ligated concatemers to reveal high-order conformations. Recently developed genome-wide, ligation-free approaches, including SPRITE, GAM, and ChIA-Drop, can also survey multiway chromosomal contacts and are not affected by artifacts arising from proximity ligation; however, these assays still require crosslinking. SPRITE involves sequencing barcoded DNA following multiple rounds of split-pool tagging, with the expectation that each interacting chromatin complex will have a distinct barcode. Thus, an interaction map can be determined by compiling the retrieved DNA segments with the same barcode. In the GAM technique, interaction data are gathered from thin cryosections of fixed cell nuclei that are collected by laser microdissection. Contact frequencies can be inferred because adjacent DNA loci are more likely to be present in the same nuclear slice. In ChIA-Drop, a chromatin complex isolated by ChIP is subjected to droplet-based sequencing. As the contents of each droplet are uniquely barcoded, the interaction map can be determined in a manner similar to that of SPRITE but with single-molecule resolution. Notably, several years after their respective development, SPRITE and GAM have yet to be widely adopted, possibly due to distinct technical challenges associated with implementation of each technique.
3D genome architectural features, including EPIs, have been studied in bulk or in single cells by combining methods that probe DNA‒DNA, DNA‒RNA, and DNA‒protein interactions with sequencing and microscopy approaches. a Hi-C, a genome-wide 3C-based technique, and its derivatives have profoundly impacted investigation of the 3D genome. Contacts between genomic loci can be captured by Hi-C at kilobase-scale resolution, but this method requires massive sequencing reads. To achieve high resolution with lower sequencing costs, protein-centric immunoprecipitation was combined with Hi-C, yielding PLAC-seq, which is also known as HiChIP. Ligation-free methods, such as SPRITE and GAM, were developed to minimize artifacts arising from proximity ligation. b Multiplexed oligo-based FISH allows for systematic direct tracing of 3D genome structure and has been adapted to visualize promoters and enhancers by targeting histone modifications. c Actively transcribing genomic loci, such as genes and enhancers, can be monitored simultaneously with two-color, live-cell imaging by visualizing genomically integrated MS2/PP7 repeats via fluorescently tagged-versions of their respective RNA-binding proteins, MCP and PCP.
Imaging-based results
In parallel, application of quantitative imaging methods has revealed molecular features of EPI in single cells. Aspects of genome structure, including the spatial distance between two or more chromosomal loci, can be directly visualized in both fixed and live cells by high-resolution microscopy.
Advancements in DNA/RNA detection and microscopy techniques have facilitated imaging of EPI. While standard fluorescence in situ hybridization (FISH), in which fluorescently labeled probes target DNA and RNA in fixed cells, has limited resolution and can detect a small number of probes, super-resolution microscopy in combination with multiplexed probes allows for visualization of interactions involving >1000 genomic loci, including those located within a proximity of 10–100 kb16,17. By separating fluorescent signals in time that are too close to resolve in space, advanced microscopic techniques such as STORM/PALM overcome the diffraction limit to increase resolution9. Limitations on the number of genomic loci that can be visualized simultaneously have also been overcome by barcoding and sequential, combinatorial labeling approaches, including OligoFISSEQ18 and MERFISH (Fig. 1b)17. An analysis of chromosome 21 by MERFISH revealed a high correlation with previously published Hi-C data for various features of 3D genomic structure, including the distribution of A/B compartments as well as the location of TADs and TAD boundaries, despite substantial heterogeneity at the single-cell level17. MERFISH has been further adapted to image particular epigenomic features in a high-throughput manner by inclusion of TN5-mediated tagmentation targeting H3K4me3, H3K27ac, or H3K27me3 histone modifications that mark active promoters, active genomic loci, and silenced genomic loci, respectively. Epigenetic regions of interest were tagged with the T7 promoter for amplification of the targeted DNA fragments by in situ transcription, and the resulting RNAs were detected via MERFISH, revealing hundreds of genomic loci decorated with the specific histone marks19. This epigenomic MERFISH study showed the spatial distribution of putative enhancers, enhancer–promoter pairs, and enhancer hubs during mouse brain development. To benchmark the modified technique, images for a subset of labeled promoters were compared to the expression patterns of the corresponding genes reported in the Allen brain in situ hybridization (ISH) atlas, which demonstrated high congruence.
Live-cell imaging has extended an understanding of 3D genome structure to a fourth dimension by recording interaction dynamics over time. Visualization of chromatin loci in live cells can be achieved by imaging repeated sequences of DNA or RNA bound by fluorescently- labeled, sequence-specific nucleic acid binding proteins. Bacterial operon systems, such as the lactose operon (LacO/LacR), tetracycline operon (TetO/TetR) and cumate gene-switch (cuO/CymR), have been employed to visualize targeted genomic loci via insertion of multiple copies of an operator sequence and expression of the cognate repressor protein with a fluorescent tag. These systems can be combined for visualization of two different genomic loci with distinct fluorophores20. MS2 and its analog PP7, which are derived from RNA bacteriophages, are currently the most widely used RNA-imaging system. MS2 and PP7 sequences recruit their respective binding proteins, MCP and PCP, which can be differentially labeled by fluorescent protein fusion to enable dual-color, live tracking of RNA dynamics (Fig. 1c). The MS2/PP7 system also allows for detection of genomic loci that are actively transcribed. Accordingly, MS2/PP7 live-imaging studies have visualized EPIs and demonstrated the critical role of enhancers in controlling the transcriptional bursting dynamics of cognate gene promoters21,22,23. While targeting exogenous DNA or RNA sequences can confer robust signal strength, it is technically challenging due to the laborious nature of the requisite genetic manipulation(s).
CRISPR-based technologies, which allow for sequence-specific recognition without genetic editing, offer an alternative strategy that can be employed independently or in combination with MS2/PP7 labeling. An enzymatically-dead version of CRISPR-associated protein 9 (dCas9) has been used in imaging studies of native genomic regions to evaluate EPI dynamics. In this approach, signal amplification and/or multiplexed gRNA delivery is typically needed to improve the signal-to-background ratio and overcome the inefficient labeling of nonrepetitive genomic loci24. The discovery of the Cas13 family, comprising Cas13a-d, X, and Y, which specifically target RNA, has enabled the introduction of additional experimental tools that can be applied to EPI studies25. Recently, several reports have demonstrated the utility of enzymatically-dead CRISPR-Cas13 (dCas13) proteins as a new platform for RNA imaging in live cells26. Dual-color imaging has been achieved by employing orthologous dCas13 proteins and by coupling the dCas13 system with either MS2 RNA labeling or dCas9 genomic locus detection27,28. Although dCas13-mediated visualization of eRNAs has not been reported, this potential application would be valuable for further elucidation of the molecular underpinnings of EPI dynamics.
Methods for imaging 3D genome structure have been empowered by the development of super-resolution fluorescence microscopy technologies, especially single-molecule localization microscopy, such as SIM, SMLM, and STED, which have improved resolution to the low-nm range29. A technique called MINFLUX, which offers a superior resolution of 1–3 nm in fixed and live cell imaging, was recently combined with DNA-PAINT for multicolor labeling of mitochondria30. MINFLUX has not been applied to studies of genome architecture, but this nanoscopy approach may eventually yield additional insights into the molecular details of EPIs.
Functional validation of enhancers
While strategies that provide 3D genome information and decipher the epigenetic landscape are useful for evaluating presumed CREs and their interactions, these approaches cannot validate enhancer activity in the native chromatin context. Genome editing techniques have been used to delete or alter certain enhancers in cell lines and animal models, but traditional approaches are laborious and low-throughput. Development of CRISPR-based, epigenome-editing technologies has allowed for relatively simple targeting of transcriptional repressive machinery to specific regions of interest, including enhancers and promoters, by coupling dCas9 to different repressors/repressor domains, such as the KRAB domain, methyl-CpG binding protein 2 (MeCP2), and DNA methyltransferases (DNMTs) (CRISPRi). Alternatively, dCas9 can be outfitted with activator domains to achieve the opposite effect (CRISPRa). KRAB repressor domain-fused dCas9 efficiently silences 200–500 base pair (bp) genomic regions in a sequence-specific manner by promoting deposition of suppressive histone modifications that induce heterochromatin formation via the recruitment of additional factors, such as KAP1 and HP1. The repressive capacity of dCas9-KRAB is enhanced by attachment of MeCP2 (dCas9-KRAB-MeCP2). In the case of DNMT fusions, DNA hypermethylation of target regulatory regions causes transcriptional down-regulation. Furthermore, when dCas9 is connected to KRAB, Dnmt3A, and Dnmt3L simultaneously, the system can confer a heritable repressive effect that persists even after cell division and differentiation (CRISPRoff)31. These CRISPR tools allow for validation of functional enhancers of particular target genes in a case-by-case fashion, but they also have been employed for multiplexed and genome-wide interrogation of enhancer function32,33. For example, a CRISPRi approach integrated with FISH and flow cytometry, called CRISPRi-FlowFISH, was used to identify functional noncoding variants from genome-wide association studies (GWASs)34. In addition, an integrative method termed STING-seq, which combines CRISPRi-based perturbation of GWAS-linked CREs and single-cell RNA sequencing, allowed for identification of causal variants that affect disease-related gene expression in a high-throughput manner35.
Molecular mechanisms underlying EPI specificity
Although the role of enhancers in the regulation of gene transcription is well established, the fundamental question of how enhancers communicate with their distal gene promoter target(s) remains largely unresolved. In this section, we describe several molecular mechanisms that contribute to the enhancer–promoter interactome.
Looping-dependent models of EPI
CTCF binding and loop extrusion by the molecular motor cohesin create a 3D framework for functional networks of dynamic long-range interactions between promoters and enhancers in the nucleus7,36. In this model, the DNA loop-extruding movement of cohesin along chromatin is halted at CTCF binding sites (CBSs) positioned in a convergent orientation. A higher frequency of physical interaction within the region demarcated by convergent CTCF sites, as measured by Hi-C, constitutes the basis of TADs or loop domains. Flanking CTCF sites serve as loop anchors/TAD boundaries that often have an insulator function (for a detailed review, see refs. 37,38). Targeted degradation of the cohesin machinery or CTCF compromises local TAD structure and TAD insulation, while higher-order chromatin structures (i.e., chromosome compartments) are not necessarily affected39,40,41. Conversely, transcription enforces TAD strength, long-range interactions and boundary site insulation42, possibly through weak CTCF-RNA interactions, biomolecular condensate formation, and/or the process of transcription itself43,44.
While discrepant results have emerged from degron-targeted CTCF degradation in cultured cells, such depletion experiments have generally revealed a surprisingly subtle immediate impact on transcription, affecting only a limited subset of genes initially despite the near complete loss of TADs (Fig. 2a)41,45,46. This may be attributed, at least in part, to the extent of CTCF protein degradation that is achieved by a degron system within a particular cell type. Although the degron approach routinely renders CTCF undetectable in nuclear lysates and even in chromatin fractions by immunoblotting, the amount of residual CTCF at individual binding sites can be highly variable, with some retaining 90% of CTCF binding upon acute depletion47,48. These differences may result in temporally nuanced outcomes for transcriptional regulation41,49, as not all CTCF binding sites are targeted with the same kinetics48. Stronger CTCF binding sites, which are functionally better insulators, seem to retain CTCF longer than weaker sites41,48,50,51, perhaps reflecting slower turnover of CTCF at the former or the effect of chromatin-based differential accessibility on the degron system. These findings also raise the possibility of context- and cell type-specific roles for CTCF in gene expression. However, limited transcriptional changes were observed in mouse embryonic stem cells (ESCs), even in the context of almost 100% degron-mediated depletion of CTCF or cohesin genome-wide at an early (3 h) time point46. Thus, a “molecular memory,” mediated by various chromatin features, might render CTCF, cohesin, and other looping factors dispensable for established EPI at least initially and buffer against any immediate transcriptional changes. This possibility is consistent with micro-C data demonstrating that global EPIs are largely maintained upon acute depletion of CTCF, cohesin or the looping-associated chromatin protein Ying Yang 1 (YY1), although they may weaken over time46. While nonessential for enhancer–promoter pairs that are closely positioned, more distant interactions rely increasingly on loop extrusion by cohesin or nearby boundary CTCF sites51,52. In addition, application of a tiled micro-C approach, allowing for even finer resolution of sub-TAD scale alterations, has revealed a decreased efficiency in EPIs and associated subtle transcriptional effects upon degron-mediated removal of cohesin. In this setting, weakened EPIs presumably persevere due to DNA contacts that involve transcription factors or CTCF via cohesin-independent looping mechanisms39,53.
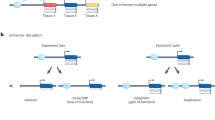
a The effect of acute CTCF/cohesin depletion on transcription and EPI is remarkably limited. Despite widespread loss of chromatin loops anchored at CBSs and weakening of chromatin domain borders, only subtle changes in gene expression are detected. A subset of genes with increased transcription after CTCF degradation is characterized by relatively close positioning of its promoters to enhancer elements, while the other subset, comprising downregulated genes, shows loss of promoter-bound CTCF. b Two nearby TADs can combine in the absence of CTCF binding at a TAD boundary due to mutation, epigenetic perturbation of the CBS, or release of CTCF by RNA interaction. TAD fusion can lead to ectopic contact between promoters and enhancers, which is known as enhancer hijacking. c Genetic variants located in promoter CTCF binding sites prevent interaction of an enhancer with its preferred promoter, allowing the enhancer to interact with and activate other gene promoters in the same TAD, a phenomenon referred to as enhancer release and retargeting (ERR).
Deletion of CTCF binding sites in TAD boundaries results in local transcriptional dysregulation due to altered EPI54, whereas complete genetic knockout (KO) of CTCF at the organismal level causes widespread changes in gene expression in addition to reduced TAD insulation. The latter manipulation was accomplished in zebrafish, which, unlike mice, can seemingly tolerate loss of endogenous CTCF until a relatively late stage of development due to the provision of maternal CTCF protein55. Among the limited set of genes that is differentially expressed after degron-mediated depletion of CTCF, upregulated genes tend to be located closer to enhancers41,56, likely indicating their activation by ectopic enhancer–promoter contacts that are also enabled by disruption of TAD insulation45,53. Genes that are down-regulated after CTCF degradation tend to display loss of CTCF binding in their promoters41,45,47,49,56, potentially reflecting the role of promoter-bound CTCF in chromatin accessibility and/or in enabling EPI (Fig. 2a)47,56,57.
Like many other chromatin-associated proteins, CTCF is thought to be functionally regulated by RNA interactions. CTCF zinc finger (ZnF) 1, ZnF10, and a C-terminal RNA-binding region have been shown to bind a number of lncRNAs with low affinity (Fig. 3)58,59,60,61,62, and modeling suggests the presence of shared structural motifs in the RNA interaction partners63. RNA binding contributes to CTCF self-association and its sub-nuclear dynamics58,59,60,64. In addition, genome-wide CTCF occupancy is altered by mutations that disrupt CTCF-RNA binding59,60, raising the intriguing possibility that CTCF-RNA interactions also shape the 3D genome. Notably, the X chromosome-encoded lncRNA Jpx has been reported to function as a CTCF release factor. Jpx selectively dismisses CTCF from loop anchor regions, which alters TAD structures and concomitantly impacts transcription due to newly formed EPIs (Fig. 2b)61. Thus, the emerging CTCF-RNA interactome might influence local interactions as well as global nuclear organization by fine-tuning TAD architecture. Disruption of these looping events by non-coding SNPs/mutations that affect expression or secondary structure of CTCF-interacting RNAs may have phenotypic consequences as a result of altered EPIs and consequent gene expression changes. However, it should be noted that the in vivo specificity of RNA interactions involving numerous chromatin-associated proteins, including CTCF and YY1, has recently been challenged, possibly necessitating further validation to confirm functional significance in many cases65.
CTCF is characterized by a central 11 zinc-finger (ZnF) DNA-binding domain. ZnF3-7 (green) mediate binding to the core CTCF DNA motif. The IUPred3 disorder plot168 indicates the presence of an intrinsically disordered region (IDR) in the C-terminus. ZnF1, ZnF10, and part of the IDR domain are involved in RNA binding (orange). The relative position of a subset of documented disease-linked CTCF mutations is also indicated, with those implicated in neurodevelopmental disorders and diverse cancers colored in blue and red, respectively132,134,135,169,170,171,172.
Signal-induced nuclear receptor activation showcases the specificity of EPIs66,67,68. In the well-studied glucocorticoid receptor (GR) system, NIPBL is employed by GR to facilitate enhancer–promoter looping69. EPI specificity in estrogen receptor (ER)-dependent transcription is mediated by TF binding to enhancers that encourages condensate formation, the presence of cohesin, and the activity of topoisomerase I7,70,71. Nuclear hormone receptors themselves have been suggested to serve as anchors for intra-TAD looping72. The Mediator complex, which physically links nuclear hormone receptors and other transcriptional activators to the basal transcription machinery, also functionally connects enhancers to promoters73, although its architectural role in EPIs might not be direct and may involve the formation of condensates74,75.
Looping-independent models of EPI
While genome-wide analyses have demonstrated that eukaryotic transcription arising from promoters and enhancers is subject to common regulatory controls that frequently involve their physical interaction or spatial proximity, looping-independent mechanisms of enhancer–promoter interplay are not mutually exclusive and may even prevail in some instances. Alternative models of EPI that do not necessarily require looping events include tracking, whereby enhancer-bound RNA Pol II scans intervening DNA in a linear fashion to seek out its cognate promoter, and linking, which entails oligomerization of a regulatory protein that binds interacting CREs to establish a linkage76. In the scanning model, RNA Pol II, if not continuously transcribing, might be aided by a DNA motor protein, such as cohesin, possibly helping to explain the effects of CTCF-bound insulator elements that, when situated intermediately within TAD boundaries or otherwise, restrain the capacity of enhancers to activate neighboring promoters76,77. The linking model derives from the observation that oligomerized bacteriophage lambda repressor connects bound regulatory sites, or operators, with target promoters78, but there is limited supporting evidence for this phenomenon in eukaryotic systems76.
Non-looping models of EPI do not stipulate increased 3D colocalization of regulatory elements. In this regard, super-resolution imaging of sonic hedgehog gene (Shh) activation in the context of neural differentiation has revealed decreased enhancer–promoter proximity that can be disrupted by DNA-tethered protein impediments79. The increased enhancer-promoter spatial separation was ascribed to the enzymatic activity of the transcriptional cofactor poly(ADP-ribose) polymerase 1 (PARP1), which synthesizes long, branched chains of the nucleic acid PAR as a covalent modification of histones and other proteins to promote chromatin decompaction and non-covalent protein recruitment. These observations are consistent with a tracking or linking model79. However, a different developmental enhancer seems to rely on looping-dependent spatial proximity, albeit largely invariant, to achieve appropriate spatiotemporal Shh expression80, and other examples in which enhancer–promoter distance shows a positive correlation with gene activation have not been rigorously characterized. Nevertheless, it is plausible that for some instances in which looping mediates EPI, linking and/or tracking mechanisms contribute to the physical or functional interplay of regulatory elements brought into spatial proximity76.
Transcriptional control involving EPI
Bursting
Expression of most, if not all, genes in prokaryotes and eukaryotes occurs in a discontinuous fashion, involving episodes of activity separated by refractory intervals of complete or near-complete inactivity. In metazoans, this pulsatile nature of gene expression, dubbed transcriptional bursting, may be crucial for regulation of developmental programs as well as for signal-dependent responses in differentiated cells or tissues. Mechanistically, transcriptional bursting is regulated by EPI as well as other DNA sequence and chromatin features known to impact gene expression, such as core promoter elements81, nucleosome positioning/density82, TF occupancy/dwell time82, and epigenetic modifications21,83,84,85. The frequency of bursting, as opposed to the amplitude or duration, is the parameter most acutely linked to enhancer activity (Fig. 4a)81,85. Strong enhancers increase bursting frequency at their cognate promoters, which corresponds to higher levels of gene expression81,85. Moreover, an artificially imposed linkage of the β-globin gene promoter with its distal enhancer results in transcriptional activation by selectively increasing bursting frequency21.
a Gene expression occurs in a discontinuous fashion, referred to as bursting. Enhancers can increase bursting frequency to augment cognate gene expression via increased enhancer–promoter contact. Enhancer bursting, unlike promoter bursting, and synchronization of enhancer/promoter activation have yet to be visualized by imaging techniques. b Interaction of IDR-containing proteins (TFs, cofactors, and other chromatin-associated proteins) and their association with eRNAs results in formation of biocondensates. These phase-separated structures are thought to form at specific genomic loci to facilitate robust transcriptional activation. c Multiple mechanisms may allow enhancers to induce pause release at promoters by discharging pausing factors. Anti-pause enhancers (A-PEs) increase gene expression from cognate promoters via JMJD6-dependent dismissal of 7SK snRNA and HEXIM1/2. Alternatively, eRNAs can cause disassociation of promoter-bound NELF.
Bursting was first observed more than four decades ago in electron micrographs as ribonucleoprotein (RNP) fibers emanating in a discontinuous pattern from Drosophila melanogaster sister chromatids. In recent years, increasingly sophisticated techniques, including live cell imaging of nascent transcripts, single-molecule FISH (smFISH), and sc- or snRNA-seq, have been employed to study the phenomenon at static time points and in real time86. As the field continues to develop, there are still many unanswered questions, especially regarding EPIs. While Hi-C data indicate that EPI reliably predicts gene activation87, how activation of one or more enhancers, based on eRNA production, is coordinated with that of a target gene requires further inquiry. As non-coding transcription is also presumed to be discontinuous, it remains unclear to what extent bursting at promoters and enhancers is synchronized, although the latter is more challenging to investigate for biological as well as technical reasons. In addition, a biochemical dissection of the requisite components and molecular mechanism(s) governing bursting kinetics at promoters and enhancers in the context of EPIs, as might be feasible with cell-free assays88,89, would be particularly instructive.
Pause release
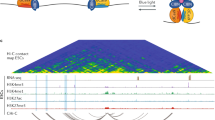
Key points of regulatory control of promoter-driven transcription include recruitment of the pre-initiation complex (PIC), comprising general transcription factors as well as RNA Pol II, and subsequent release of proximally paused RNA Pol II90, both of which may be subject to modulation via EPI88. A study using yeast nuclear extracts and an upstream activating sequence (UAS)-promoter system showed the spatiotemporal dynamics of enhancer-promoter interplay in PIC assembly by single-molecule visualization91, suggesting sequential enhancer-promoter activation. In this context, RNA Pol II pre-assembles at the UAS/enhancer with other PIC components before the complexes are transferred to the core promoter. Similarly, at the α-globin gene locus in differentiating erythroid cells, the PIC is first recruited to distal enhancers prior to its promoter deposition in concert with increased enhancer–promoter association92. ChIP-seq assays have substantiated the widespread enhancer presence of the PIC93 along with that of the large Mediator coactivator complex94 in metazoans74. Although 3C assays indicate that Mediator can collaborate with cohesin to facilitate EPI94, promoter-based capture Hi-C data suggest that this may not be a global phenomenon95. Similarly, the role of the PIC in EPI might be limited globally95. Further genome-wide interrogation of various cell types and/or conditions is needed to elucidate any potential reciprocal relationship between RNA Pol II transcriptional machinery loading and EPI.
Following initiation, RNA Pol II typically transcribes less than 100 nucleotides before pausing, which is mediated by the pausing components DRB sensitivity-inducing factor (DSIF) and negative elongation factor (NELF). Paused RNA Pol II is released by the kinase activity of positive transcription elongation factor b (P-TEFb), which phosphorylates DSIF and NELF, altering the suppressive function of the former and causing dissociation of the latter to allow for resumption of transcription96. In multiple cell types, the arginine demethylase jumonji C-domain-containing protein 6 (JMJD6) and bromodomain-containing protein 4 (BRD4), an epigenetic reader of acetylated histones, collaborate at a collection of distal CREs, termed anti-pause enhancers (A-PEs), to facilitate pause release and thus gene activation at a large group of regulated promoters. Detailed mechanistic analysis supports a model in which JMJD6 is recruited to A-PEs by BRD4 to demethylate both 7SK snRNA and the modified histone with which it associates, H4R3me2(s0). Accordingly, JMJD6 secures the dismissal of 7SK snRNA and HEXIM1/2, which collaborate to inhibit P-TEFb-mediated pause release (Fig. 4c). A-PEs associate with target gene promoters, as demonstrated by 3 C analysis, suggesting that EPI confers specificity in this regulatory strategy. Alternatively, EPI may also ensure local accumulation of trans-acting eRNAs, which can bind and dissociate NELF from regulated promoters to elicit pause release97. Notably, pausing factors, including NELF94 and PAF198, bind to enhancers, and transcriptional pausing has been detected at enhancers genome-wide (Fig. 4c)99, but it is unknown whether pause release at functionally linked enhancers and promoters occurs in synchrony via their interaction.
Phase separation
Recent appreciation of the potential biological role(s) of condensates formed by liquid‒liquid phase separation (LLPS) has altered views on the formation and function of membraneless structures within cells, such as nucleoli, nuclear speckles, and stress granules, while affording new insights into many cellular activities, including transcription71,100,101,102,103,104,105 as well as dynamic changes in 3D chromatin architecture43,103. The activation domain of TFs often features one or more intrinsically disordered regions (IDRs), stretches of amino acids, sometimes with low sequence complexity, that do not fold into stable structures but rather can assume various conformations and bind diverse partners. CTCF and other architectural proteins are also predicted to contain IDRs (Fig. 3), which may have functional implications for the (re)organization of genome topology. IDR-containing proteins tend to self-associate and interact with each other via numerous IDR-mediated, low-affinity, multivalent interactions106. Depending on their sequence composition, many IDRs can also bind RNA, either alone or in collaboration with RNA-binding motifs that are frequently found in the same protein107. Accordingly, formation of condensates, which are nucleated by enhancer sequences that attract various IDR-containing TFs and induce short promoter-derived ncRNAs that act in concert with eRNAs, has been proposed to facilitate transcriptional initiation (Fig. 4b)101,102,108. However, high levels of RNA produced by elongating RNA Pol II may promote condensate dissolution via a negative feedback effect, possibly resulting in the refractory period that follows a transcriptional burst. Thus, LLPS offers a potential biophysical explanation for the pulsatile nature of transcription promoted by EPI101,102. Condensate formation may also help to explain the coordinated bursting of two genes by a single enhancer85,109 as well as cooperative chromosomal enhancer assembly71. However, while in vitro assays using purified proteins have allowed for exquisitely detailed analysis of the biochemical determinants of condensate formation as well as their functional potential, the tools available for studying phase separation within intact cells remain limited, and, consequently, definitive proof of its role(s) in transcriptional regulation and genome structural organization has not been achieved110.
EPIs in disease
Changes in enhancer function or activity with pathological consequences have been ascribed to multiple molecular mechanisms, including SNPs, epigenetic modifications, genomic structural variants (Table 1), and mutations in architectural proteins (Table 2). These ‘enhanceropathies’ clearly involve altered EPI in some instances, but this aspect has not been rigorously investigated in many cases.
Enhancer variants
Small insertions (1–31 bp), which are readily detected in cancer cell genomes, can alter enhancer activity or potentially generate de novo enhancers, leading to transcriptional changes that can affect oncogene expression111. Disease-linked SNPs detected by GWASs are predominantly located in non-coding sequences, including intronic and intergenic regions that often constitute enhancers; however, enhancer–promoter pairs require experimental identification/validation, and disease-relevant enhancers typically show cell type-specific activity, complicating straightforward pathological assignments for SNPs (as well as de novo mutations) found in putative enhancers.
In addition to genetic perturbations, epigenetic changes can impact enhancer activity/function with direct or indirect effects on EPIs. In this regard, enhancer-specific alterations in DNA methylation or in the histone modification landscape, including H3K27ac profiles, have been linked to specific disease phenotypes (Table 1)112. Comprehensive analysis of enhancer activity based on DNA hypersensitivity and H3K27ac for almost 9000 samples representing 33 cancer types revealed genome-wide activation of enhancers accompanied by tumor aneuploidy113, suggesting that enhancer activity not only promotes oncogene expression but also facilitates chromosomal structural rearrangements. Human endogenous retroviral loci constitute a potential alternative source of enhancer activity impacting proximal gene promoters in cancer114. Reactivation of human endogenous retrovirus (HERV) subfamily K was recently shown to contribute to senescence-associated inflammation and aging115, indicating even broader (patho)physiological relevance, but the role of an HERV-derived enhancer program as well as EPI in adjacent and distal gene expression requires further investigation in diverse cell types. Notably, CRISPRi strategies, which feature high specificity and scalability, have greatly enabled identification of functional EPIs and evaluation of enhancer epigenomic landscapes in looping events32,33,34,35.
Enhancer mistargeting by alteration of 3D genomic organization
Changes in 3D genome structure can cause alterations in EPIs that induce expression of genes other than the original target(s). Genomic rearrangements, which are commonly observed in most cancer types and sometimes may arise from a single catastrophic event116, can reposition CREs to allow for enhancer hijacking, a scenario in which an enhancer regulates one or more new target gene promoters112. Copy number variation that alters TAD structure due to disruption of TAD boundaries, allowing for enhancer retargeting, has been implicated in a few congenital diseases and several cancers112. Analysis of structural variants (SVs) in more than 1200 cancer samples by whole-genome sequencing revealed hundreds of genes located ≤100 kb of an SV breakpoint with altered expression, most of which were up-regulated, including the cancer-associated genes TERT, MDM2, CDK4, ERBB2, CD274, PDCD1LG2, and IGF2117. Remarkably, while TAD disruption, which can also be caused by point mutations in boundary-localized CTCF-binding motifs118, may be a frequent feature of cancer genomes, its effects on proximal gene expression are usually limited112,119. Overall, 3D genome studies have convincingly implicated enhancer mistargeting and hijacking as disease mechanisms, but there are currently few validated examples due to the traditional challenges of identifying functional enhancers and determining their cancer-driving potential.
Promoter variants
Alterations in promoter sequences can impact nearby gene expression beyond the associated gene body. Recent reports suggest that functional loss of a particular promoter due to mutation results in release of its cognate enhancer, which can then engage and activate alternative promoters in the same TAD by a mechanism called enhancer release and retargeting (ERR) (Fig. 2c)57. At the TFF1 locus in MCF7 cells, deletion of the TFF1 promoter, which is the preferred target of the proximal TFF1 enhancer (TFF1e), allows for an ectopic looping event between TFF1e and a different promoter, namely the TFF3 promoter that is located 50 kb away. This retargeting causes a > 20-fold increase in TFF3 transcription. In addition to cancer-associated SNPs in the TFF1 promoter that correlate with TFF3 expression, activation of other proto-oncogenes, such as MYC, can be induced by alteration of neighboring gene promoters57,120.
Additional examples of ERR in cancer and development have been reported. In B-ALL, deletion of the FLT3 promoter correlates with ectopic upregulation of the CDX2 gene, which is positioned 30 kb away, and coincides with retargeting of an upstream enhancer to the promoter of the latter121. In gastric cancer, analysis of ENCODE and GTEx eQTL data revealed that a gene encoding the class A orphan GPCR protein GPR35 is activated by the ERR mechanism122. GPR35 expression is induced by a retargeted upstream enhancer upon mutation of the CTCF-binding region in the promoter of the neighboring gene, CAPN10. During mesoderm formation, enhancer-dependent upregulation of Mesp2 was observed in KO embryos for the nearby Mesp1 gene123. Furthermore, promoter competition resembling ERR has been documented for the β-globin locus control region (LCR)124. In one scenario, deletion of the α-globin genes augmented expression of the NME4 gene seemingly due to increased interaction of the latter with the major upstream regulatory element MCS-R2, despite a linear distance of 300 kb. However, it is unclear why ectopic contact with MCS-R2 was insufficient to elicit expression of several other genes in the region. At the same locus, impairment of the HBB promoter redirected the LCR from the adult HBB gene to the fetal HBG genes, resulting in increased production of fetal HBG125. ERR-like phenotypes are also observed during transvection, a phenomenon involving EPI between paired homologous chromosomes in Drosophila. In this context, several cases have been described in which deletion of a promoter for an allele on the same homolog as its enhancer results in increased promoter activity of the allele on the other chromosome126. Finally, a potential contribution of ERR in autism spectrum disorder (ASD) has recently been suggested. Rare promoter de novo variants (DNVs) identified by whole-genome sequencing of ASD individuals are enriched in TADs that contain known ASD risk genes but are often located in the promoters of constituent non-ASD risk genes. Transcriptomic analysis of modified iPSC lines harboring selected DNVs revealed gene dysregulation within the TAD as well as global expression changes, including that of multiple ASD risk genes. While these observations are suggestive of altered 3D genome architecture and CRE interactions in this neurodevelopmental disease setting, definitive proof of the ERR mechanism remains to be established127.
The role of architectural proteins in disease-associated 3D genome structure
As a major regulator of 3D genome structure in most metazoans, CTCF influences gene expression under physiological and pathological conditions (Table 2)128,129,130. Haploinsufficiency in a Ctcf+/- mouse model leads to increased tumorigenesis due to disrupted CpG methylation131. In humans, CTCF mutations have been implicated in various cancers and also contribute to neurological disorders132,133,134,135. CTCF mutations associated with human diseases are often located in the zinc finger (ZnF) DNA-binding domain (DBD), which can have pleiotropic effects as a result of altered transcriptional regulation and TAD organization (Fig. 3)134,135. In addition, CTCF binding site mutations are frequent in cancers118,130,136,137,138, leading to loss of insulator sites and aberrant transcription. When promoter CTCF sites are mutated, enhancer retargeting to neighboring gene promoters can occur57.
Somatic mutations in components of the cohesin complex have been reported in several cancer types whereas inherited mutations cause human developmental disorders that feature profound genome instability without any cancer predisposition (Table 2). The latter group, collectively referred to as ‘cohesinopathies’, includes Cornelia de Lange syndrome (CdLS)139. In a cell model of CdLS harboring a mutation in the cohesin-loading factor NIPBL, gene clusters with differential cohesin binding display disease-associated gene expression changes that may arise from disrupted promoter-promoter and enhancer-promoter contacts140. Induced proteolytic cleavage of the cohesin subunit RAD21 in postmitotic neurons recapitulates the transcriptional changes observed in CdLS patients141. In some solid and hematopoietic cancers, cohesin component expression levels correlate with prognosis and metastasis142,143. Although involvement of cohesin in cancer was initially ascribed to chromosome segregation defects caused by aberrant sister chromatid separation during the cell cycle and associated aneuploidy143, recent studies have demonstrated the relevance of its roles in TAD organization and transcriptional dysregulation. Cancer-associated mutations in cohesin subunits are overrepresented in STAG2142. Degron-mediated depletion of STAG2 in hematopoietic stem cells does not cause widespread disruption of chromosomal compartments or TADs, likely due to partial compensation by STAG1144. However, loss or downregulation of STAG2 specifically disrupts the local structure of a subset of TADs associated with stem cell self-renewal and differentiation, leading to the formation of new, long-range DNA loops to more distal sites and concomitant transcriptional dysregulation144,145, consistent with a tumor-suppressor role of cohesin in some cancers146.
Rare mutations in genes encoding components of condensin complexes have also been linked to human disease (Table 2). ‘Condensinopathies’ caused by inactivating mutations in condensin subunit genes typically manifest as neurodevelopmental disorders147. Mutations in the condensin II subunits NCAPD2, NCAPD3, and NCAPH result in loss of chromosome structural integrity and impaired chromosome segregation148. Cohesin and condensin have distinct spatially- and temporally-defined roles in loop extrusion during the cell cycle37,38,149, which is also the case for the two condensin complexes. Condensin II is located in the nucleus throughout the cell cycle, whereas condensin I is largely restricted to the cytoplasm during interphase150. In fission yeast, Hi-C interrogation has revealed chromosomal interactions that are quantitatively dependent on condensin, and degron-induced loss of condensin increases DNA mobility151. Similar to cohesin disruption, some condensin mutations may impact transcriptional activity as a result of alterations in local chromatin compaction and TAD structure.
The zinc-finger protein YY1 is a context-dependent activator/repressor that mediates EPI and selectively regulates pluripotency-related gene expression152,153. The ubiquitously expressed transcription factor may regulate as much as 10% of the human transcriptome154, and altered YY1 expression has been linked to various cancers and neurological diseases (Table 2)155,156. Since YY1 interacts with chromatin modifiers as well as chromatin-remodeling complexes and has roles in DNA repair, it remains largely unknown whether YY1 looping function is a salient feature of its diverse disease associations155.
MeCP2 is a ubiquitously expressed epigenetic regulator that is present at high levels in neurons and has been associated with multiple neurological disorders (Table 2). Notably, MeCP2 is mutated in 95% of individuals with Rett syndrome (RTT)157. Although MeCP2 can serve as a transcriptional activator in certain contexts, its role in inhibiting transcription via recruitment of the NCoR1/2 corepressor complex to specific sites of methylated DNA is crucially compromised in RTT158. As evidence of its relevance to disease-related genomic architecture, the RTT-associated Dlx5-Dlx6 locus is derepressed in Mecp2-/- mice, and a repressive chromatin loop normally mediated by Mecp2 is replaced with longer-distance activating chromatin-associated loops159.
Conclusion and outlook
Our understanding of enhancer function in gene expression has dramatically evolved with the increased capacity to probe and precisely perturb the 3D structure of the genome. Over 25% of sub-TAD cohesin-dependent chromatin loops are cell type-specific and tend to correlate, albeit weakly, with variations in gene expression between cell types160. Cohesin-mediated loops are enriched for enhancers, and cohesin-bound enhancers have a propensity to interact with other enhancers as well as TSSs160. These findings, published in phase III of the ENCODE project, underscore the importance of EPIs but also illuminate ongoing challenges, which largely pertain to the issue of identifying the functional enhancer(s) of a gene. Indeed, a given gene may have multiple enhancers of varying functional significance. While the search space for EPIs has been proposed to be limited to the size of a TAD (i.e., ~1 Mb), this restriction has been questioned. Moreover, a given enhancer does not necessarily contact the nearest gene promoter and may have limited or no functional significance to its neighboring gene even if there is evidence of looping161. Finally, some or all EPIs for a particular gene may be cell type specific. Despite this complexity, there are now a number of tools, including 3C-based and imaging modalities, that make it feasible to determine functional EPI for one or many loci when combined with CRISPR genome or epigenome editing.
The role of altered EPI in human disease phenotypes is becoming increasingly apparent, and multiple molecular mechanisms have been described. Changes in EPI due to structural variation in enhancers and promoters as well as mutations in the TFs and architectural proteins that associate with them, or alterations in their binding sites, have all been implicated as drivers of oncogenic transformation112,162. Furthermore, the preponderance of GWAS SNPs are located in non-coding intronic and intergenic regions that are putative enhancers or other regulatory sites112, consistent with the possibility that alterations in EPI contribute to a multitude of pathological conditions emanating from various tissues112,163.
While dysregulation of EPIs can cause transcriptional changes in disease, they also present a promising target for therapeutic interventions112,163. A few epigenetic therapies that may impact EPI are already clinically available, including hypomethylating agents (HMAs) that inhibit DNMTs and histone deacetylase inhibitors (HDACis), for treatment of specific hematological malignancies11. These drugs broadly affect the epigenomic landscape, which can be drastically altered in the context of malignant transformation due to chromosomal rearrangements or other genetic alterations in addition to the emergence of cancer-specific super enhancers (SEs). Cancer cells rely crucially on SEs that engage in EPIs to drive oncogenic transcriptional dysregulation164. Accordingly, epigenetic agents targeting various SE components, including bromodomain and extraterminal domain (BET) proteins such as BRD4, have been extensively evaluated as potential anticancer therapies, yet none have attained clinical approval. Broader application of these different classes of epigenetic drugs in cancer treatment may require multimodal regimens instead of their use as monotherapies165.
The limited specificity of traditional epigenetic drugs may restrict their therapeutic utility beyond cancer treatments. Ultimately, enhancer-based therapies using CRISPR-derived epigenome-modulating or base-editing tools will likely offer alternative options with unparalleled specificity for many conditions. The first CRISPR-based drug, dubbed Casgevy, which was approved for use in the United Kingdom in November 2023 and shortly thereafter by the U.S. Food and Drug Administration (FDA), actually targets an erythroid-specific BCL11A enhancer initially implicated in the control of fetal hemoglobin (HbF) levels by the presence of GWAS SNPs166,167. Cas9-dependent disruption of this enhancer element in isolated CD34+ hematopoietic stem cells causes marked downregulation of the BCL11A repressor and concomitant upregulation of its target genes, which include those encoding the γ-globin subunits, allowing for stable resumption of HbF production that is protective against sickle cell disease and β-thalassemia. Furthermore, multiple clinical trials using base editors to introduce specific genetic changes, without a requirement for DNA double-strand breaks, in cells ex vivo or following in vivo delivery are currently underway. These tools may eventually be employed for precise modification of other GWAS enhancer or promoter SNPs that contribute to disease susceptibility through altered EPI. It is also possible to envision future therapeutic applications of Cas13-mediated eRNA depletion as well as dCas9/12-imposed enhancer/promoter repression or activation in EPI-linked pathologies. Development of effective CRISPR-based therapies will benefit from integration of single-cell and spatial multiomic strategies, providing genomic, epigenomic, and transcriptomic data167, in studies of functional EPIs and the consequences of their perturbation. These approaches will also allow for systematic evaluation of seemingly rare CRISPR-related off-target effects, a lingering concern that is a particularly important consideration for treatments involving in vivo modification(s). Nevertheless, based on recent progress, therapies targeting disease-associated EPI are now within the purview of precision medicine.
References
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Andersson, R. & Sandelin, A. Determinants of enhancer and promoter activities of regulatory elements. Nat. Rev. Genet. 21, 71–87 (2020).
Moore, J. E. et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Preissl, S., Gaulton, K. J. & Ren, B. Characterizing cis-regulatory elements using single-cell epigenomics. Nat. Rev. Genetm 24, 21–43 (2023).
Calderon, D. et al. Landscape of stimulation-responsive chromatin across diverse human immune cells. Nat. Genetm 51, 1494–1505 (2019).
Dao, L. T. M. & Spicuglia, S. Transcriptional regulation by promoters with enhancer function. Transcription 9, 307–314 (2018).
Nair, S. J. et al. Transcriptional enhancers at 40: evolution of a viral DNA element to nuclear architectural structures. Trends Genet. 38, 1019–1047 (2022).
Consortium, G. The GTEx consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Boettiger, A. & Murphy, S. Advances in chromatin imaging at kilobase-scale resolution. Trends Genet. 36, 273–287 (2020).
Friedman, M. J., Lee, H., Kwon, Y. C. & Oh, S. Dynamics of viral and Host 3D genome structure upon infection. J. Microbiol. Biotechnol. 32, 1515–1526 (2022).
Friedman, M. J., Lee, H., Lee, J. Y. & Oh, S. Transcriptional and epigenetic regulation of context-dependent plasticity in T-Helper Lineages. Immune Netw. 23, e5 (2023).
Lafontaine, D. L., Yang, L., Dekker, J. & Gibcus, J. H. Hi-C 3.0: improved protocol for genome-wide chromosome conformation capture. Curr. Protoc. 1, e198 (2021).
Brant, L. et al. Exploiting native forces to capture chromosome conformation in mammalian cell nuclei. Mol. Syst. Biol. 12, 891 (2016).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Quinodoz, S. A. et al. SPRITE: a genome-wide method for mapping higher-order 3D interactions in the nucleus using combinatorial split-and-pool barcoding. Nat. Protoc. 17, 36–75 (2022).
Liu, S. & Zhao, K. The toolbox for untangling chromosome architecture in immune cells. Front. Immunol. 12, 670884 (2021).
Su, J. H., Zheng, P., Kinrot, S. S., Bintu, B. & Zhuang, X. Genome-scale imaging of the 3D organization and transcriptional activity of chromatin. Cell 182, 1641–1659.e1626 (2020).
Nguyen, H. Q. et al. 3D mapping and accelerated super-resolution imaging of the human genome using in situ sequencing. Nat. Methods 17, 822–832 (2020).
Lu, T., Ang, C. E. & Zhuang, X. Spatially resolved epigenomic profiling of single cells in complex tissues. Cell 185, 4448–4464.e4417 (2022).
Alexander, J. M. et al. Live-cell imaging reveals enhancer-dependent. Elife 8 (2019). https://doi.org/10.7554/eLife.41769
Bartman, C. R., Hsu, S. C., Hsiung, C. C., Raj, A. & Blobel, G. A. Enhancer regulation of transcriptional bursting parameters revealed by forced chromatin looping. Mol. Cell 62, 237–247 (2016).
Levo, M. et al. Transcriptional coupling of distant regulatory genes in living embryos. Nature 605, 754–760 (2022).
Chen, H. et al. Dynamic interplay between enhancer-promoter topology and gene activity. Nat. Genet. 50, 1296–1303 (2018).
Van Tricht, C., Voet, T., Lammertyn, J. & Spasic, D. Imaging the unimaginable: leveraging signal generation of CRISPR-Cas for sensitive genome imaging. Trends Biotechnol. 41, 769–784 (2023).
Bot, J. F., van der Oost, J. & Geijsen, N. The double life of CRISPR-Cas13. Curr. Opin. Biotechnol. 78, 102789 (2022).
Cao, H. et al. Progress of CRISPR-Cas13 Mediated Live-Cell RNA imaging and detection of RNA-Protein interactions. Front. Cell Dev. Biol. 10, 866820 (2022).
Yang, L. Z. et al. Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems. Mol. Cell 76, 981–997.e987 (2019).
Wang, H. et al. CRISPR-mediated live imaging of genome editing and transcription. Science 365, 1301–1305 (2019).
Jerkovic, I. & Cavalli, G. Understanding 3D genome organization by multidisciplinary methods. Nat. Rev. Mol. Cell Biol. 22, 511–528 (2021).
Ostersehlt, L. M. et al. DNA-PAINT MINFLUX nanoscopy. Nat. Methods 19, 1072–1075 (2022).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519.e2517 (2021).
Carleton, J. B., Berrett, K. C. & Gertz, J. Multiplex enhancer interference reveals collaborative control of gene regulation by estrogen receptor α-bound enhancers. Cell Syst. 5, 333–344.e335 (2017).
Xie, S., Duan, J., Li, B., Zhou, P. & Hon, G. C. Multiplexed engineering and analysis of combinatorial enhancer activity in single cells. Mol. Cell 66, 285–299.e285 (2017).
Nasser, J. et al. Genome-wide enhancer maps link risk variants to disease genes. Nature 593, 238–243 (2021).
Morris, J. A. et al. Discovery of target genes and pathways at GWAS loci by pooled single-cell CRISPR screens. Science 380, eadh7699 (2023).
Merkenschlager, M. & Odom, D. T. CTCF and cohesin: linking gene regulatory elements with their targets. Cell 152, 1285–1297 (2013).
Higashi, T. L. & Uhlmann, F. SMC complexes: lifting the lid on loop extrusion. Curr. Opin. Cell Biol. 74, 13–22 (2022).
Davidson, I. F. & Peters, J. M. Genome folding through loop extrusion by SMC complexes. Nat. Rev. Mol. Cell Biol. 22, 445–464 (2021).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e324 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944.e922 (2017).
Ulianov, S. V. et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 26, 70–84 (2016).
Gamliel, A. et al. Long-distance association of topological boundaries through nuclear condensates. Proc. Natl. Acad. Sci. USA 119, e2206216119 (2022).
Islam, Z. et al. Active enhancers strengthen insulation by RNA-mediated CTCF binding at chromatin domain boundaries. Genome Res. 33, 1–17 (2023).
Zhang, H. et al. CTCF and transcription influence chromatin structure re-configuration after mitosis. Nat. Commun. 12, 5157 (2021).
Hsieh, T. S. et al. Enhancer-promoter interactions and transcription are largely maintained upon acute loss of CTCF, cohesin, WAPL or YY1. Nat. Genet. 54, 1919–1932 (2022).
Hyle, J. et al. Acute depletion of CTCF directly affects MYC regulation through loss of enhancer-promoter looping. Nucleic Acids Res. 47, 6699–6713 (2019).
Luan, J. et al. Distinct properties and functions of CTCF revealed by a rapidly inducible degron system. Cell Rep. 34, 108783 (2021).
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl. Acad. Sci. USA 111, 996–1001 (2014).
Huang, H. et al. CTCF mediates dosage- and sequence-context-dependent transcriptional insulation by forming local chromatin domains. Nat. Genet. 53, 1064–1074 (2021).
Rinzema, N. J. et al. Building regulatory landscapes reveals that an enhancer can recruit cohesin to create contact domains, engage CTCF sites and activate distant genes. Nat. Struct. Mol. Biol. 29, 563–574 (2022).
Kane, L. et al. Cohesin is required for long-range enhancer action at the Shh locus. Nat. Struct. Mol. Biol. 29, 891–897 (2022).
Aljahani, A. et al. Analysis of sub-kilobase chromatin topology reveals nano-scale regulatory interactions with variable dependence on cohesin and CTCF. Nat. Commun. 13, 2139 (2022).
Ushiki, A. et al. Deletion of CTCF sites in the SHH locus alters enhancer-promoter interactions and leads to acheiropodia. Nat. Commun. 12, 2282 (2021).
Franke, M. et al. CTCF knockout in zebrafish induces alterations in regulatory landscapes and developmental gene expression. Nat. Commun. 12, 5415 (2021).
Xu, B. et al. Acute depletion of CTCF rewires genome-wide chromatin accessibility. Genome Biol. 22, 244 (2021).
Oh, S. et al. Enhancer release and retargeting activates disease-susceptibility genes. Nature 595, 735–740 (2021).
Hansen, A. S. et al. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell 76, 395–411.e313 (2019).
Saldaña-Meyer, R. et al. CTCF regulates the human p53 gene through direct interaction with its natural antisense transcript, Wrap53. Genes Dev. 28, 723–734 (2014).
Saldaña-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422.e415 (2019).
Oh, H. J. et al. Jpx RNA regulates CTCF anchor site selection and formation of chromosome loops. Cell 184, 6157–6173.e6124 (2021).
He, C. et al. High-resolution mapping of RNA-binding regions in the nuclear proteome of embryonic stem cells. Mol. Cell 64, 416–430 (2016).
Kuang, S. & Wang, L. Identification and analysis of consensus RNA motifs binding to the genome regulator CTCF. NAR Genom. Bioinform. 2, lqaa031 (2020).
Hansen, A. S., Amitai, A., Cattoglio, C., Tjian, R. & Darzacq, X. Guided nuclear exploration increases CTCF target search efficiency. Nat. Chem. Biol. 16, 257–266 (2020).
Guo, J. K. et al. Denaturing purifications demonstrate that PRC2 and other widely reported chromatin proteins do not appear to bind directly to RNA in vivo. Mol. Cell. https://doi.org/10.1016/j.molcel.2024.01.026 (2024).
Yang, F. et al. Glucocorticoid receptor:megatrans switching mediates the repression of an ERα-regulated transcriptional program. Mol. Cell 66, 321–331.e326 (2017).
Johnson, T. A. et al. Conventional and pioneer modes of glucocorticoid receptor interaction with enhancer chromatin in vivo. Nucleic Acids Res. 46, 203–214 (2018).
Warwick, T., Schulz, M. H., Gilsbach, R., Brandes, R. P. & Seuter, S. Nuclear receptor activation shapes spatial genome organization essential for gene expression control: lessons learned from the vitamin D receptor. Nucleic Acids Res. 50, 3745–3763 (2022).
Rinaldi, L. et al. The glucocorticoid receptor associates with the cohesin loader NIPBL to promote long-range gene regulation. Sci. Adv. 8, eabj8360 (2022).
Tan, Y. et al. Signal-induced enhancer activation requires Ku70 to read topoisomerase1-DNA covalent complexes. Nat. Struct. Mol. Biol. 30, 148–158 (2023).
Nair, S. J. et al. Phase separation of ligand-activated enhancers licenses cooperative chromosomal enhancer assembly. Nat. Struct. Mol. Biol. 26, 193–203 (2019).
Le Dily, F. et al. Hormone-control regions mediate steroid receptor-dependent genome organization. Genome Res. 29, 29–39 (2019).
El Khattabi, L. et al. A pliable mediator acts as a functional rather than an architectural bridge between promoters and enhancers. Cell 178, 1145–1158.e1120 (2019).
Soutourina, J. Transcription regulation by the mediator complex. Nat. Rev. Mol. Cell Biol. 19, 262–274 (2018).
André, K. M., Sipos, E. H. & Soutourina, J. Mediator roles going beyond transcription. Trends Genet. 37, 224–234 (2021).
Furlong, E. E. M. & Levine, M. Developmental enhancers and chromosome topology. Science 361, 1341–1345 (2018).
Burgess-Beusse, B. et al. The insulation of genes from external enhancers and silencing chromatin. Proc. Natl. Acad. Sci. USA 99, 16433–16437 (2002).
Lewis, A. E. et al. Identification of nuclear phosphatidylinositol 4,5-bisphosphate-interacting proteins by neomycin extraction. Mol. Cell Proteom. 10, M110.003376 (2011).
Benabdallah, N. S. et al. Decreased enhancer-promoter proximity accompanying enhancer activation. Mol. Cell 76, 473–484.e477 (2019).
Williamson, I., Lettice, L. A., Hill, R. E. & Bickmore, W. A. Shh and ZRS enhancer colocalisation is specific to the zone of polarising activity. Development 143, 2994–3001 (2016).
Larsson, A. J. M. et al. Genomic encoding of transcriptional burst kinetics. Nature 565, 251–254 (2019).
Donovan, B. T. et al. Live-cell imaging reveals the interplay between transcription factors, nucleosomes, and bursting. EMBO J. 38. https://doi.org/10.15252/embj.2018100809 (2019).
Senecal, A. et al. Transcription factors modulate c-Fos transcriptional bursts. Cell Rep. 8, 75–83 (2014).
Bartman, C. R. et al. Transcriptional burst initiation and polymerase pause release are key control points of transcriptional regulation. Mol. Cell 73, 519–532.e514 (2019).
Fukaya, T., Lim, B. & Levine, M. Enhancer control of transcriptional bursting. Cell 166, 358–368 (2016).
Tunnacliffe, E. & Chubb, J. R. What is a transcriptional burst? Trends Genet. 36, 288–297 (2020).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Panigrahi, A. & O’Malley, B. W. Mechanisms of enhancer action: the known and the unknown. Genome Biol. 22, 108 (2021).
Panigrahi, A. K. et al. SRC-3 coactivator governs dynamic estrogen-induced chromatin looping interactions during transcription. Mol. Cell 70, 679–694.e677 (2018).
Core, L. J. et al. Defining the status of RNA polymerase at promoters. Cell Rep. 2, 1025–1035 (2012).
Baek, I., Friedman, L. J., Gelles, J. & Buratowski, S. Single-molecule studies reveal branched pathways for activator-dependent assembly of RNA polymerase II pre-initiation complexes. Mol. Cell 81, 3576–3588.e3576 (2021).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J. A., Wood, W. G. & Higgs, D. R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Koch, F. et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nat. Struct. Mol. Biol. 18, 956–963 (2011).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Sun, F. H. et al. HPF1 remodels the active site of PARP1 to enable the serine ADP-ribosylation of histones. Nat. Commun. 12, 1028 (2021).
Adelman, K. & Lis, J. T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Schaukowitch, K. & Kim, T. K. Emerging epigenetic mechanisms of long non-coding RNAs. Neuroscience 264, 25–38 (2014).
Chen, F. X. et al. PAF1 regulation of promoter-proximal pause release via enhancer activation. Science 357, 1294–1298 (2017).
Henriques, T. et al. Widespread transcriptional pausing and elongation control at enhancers. Genes Dev. 32, 26–41 (2018).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Sharp, P. A., Chakraborty, A. K., Henninger, J. E. & Young, R. A. RNA in formation and regulation of transcriptional condensates. RNA 28, 52–57 (2022).
Henninger, J. E. et al. RNA-mediated feedback control of transcriptional condensates. Cell 184, 207–225.e224 (2021).
Shrinivas, K. et al. Enhancer features that drive formation of transcriptional condensates. Mol. Cell 75, 549–561.e547 (2019).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361. https://doi.org/10.1126/science.aar3958 (2018).
Ferrie, J. J., Karr, J. P., Tjian, R. & Darzacq, X. “Structure”-function relationships in eukaryotic transcription factors: the role of intrinsically disordered regions in gene regulation. Mol. Cell 82, 3970–3984 (2022).
Ottoz, D. S. M. & Berchowitz, L. E. The role of disorder in RNA binding affinity and specificity. Open Biol. 10, 200328 (2020).
Bosch, J. et al. Effects of blood pressure and lipid lowering on cognition: Results from the HOPE-3 study. Neurology 92, e1435–e1446 (2019).
Hnisz, D. & Young, R. A. New insights into genome structure: genes of a feather stick together. Mol. Cell 67, 730–731 (2017).
Graham, T. G. W., Ferrie, J. J., Dailey, G. M., Tjian, R. & Darzacq, X. Detecting molecular interactions in live-cell single-molecule imaging with proximity-assisted photoactivation (PAPA). Elife 11. https://doi.org/10.7554/eLife.76870 (2022).
Abraham, B. J. et al. Small genomic insertions form enhancers that misregulate oncogenes. Nat. Commun. 8, 14385 (2017).
Claringbould, A. & Zaugg, J. B. Enhancers in disease: molecular basis and emerging treatment strategies. Trends Mol. Med. 27, 1060–1073 (2021).
Chen, H. et al. A pan-cancer analysis of enhancer expression in nearly 9000 patient samples. Cell 173, 386–399.e312 (2018).
Deniz, Ö. et al. Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia. Nat. Commun. 11, 3506 (2020).
Liu, X. et al. Resurrection of endogenous retroviruses during aging reinforces senescence. Cell 186, 287–304.e226 (2023).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Zhang, Y. et al. High-coverage whole-genome analysis of 1220 cancers reveals hundreds of genes deregulated by rearrangement-mediated cis-regulatory alterations. Nat. Commun. 11, 736 (2020).
Katainen, R. et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 47, 818–821 (2015).
Akdemir, K. C. et al. Disruption of chromatin folding domains by somatic genomic rearrangements in human cancer. Nat. Genet. 52, 294–305 (2020).
Cho, S. W. et al. Promoter of lncRNA gene PVT1 is a tumor-suppressor DNA boundary element. Cell 173, 1398–1412.e1322 (2018).
Kimura, S. et al. Enhancer retargeting of CDX2 and UBTF::ATXN7L3 define a subtype of high-risk B-progenitor acute lymphoblastic leukemia. Blood 139, 3519–3531 (2022).
Shu, C. et al. ERR-activated GPR35 promotes immune infiltration level of macrophages in gastric cancer tissues. Cell Death Discov. 8, 444 (2022).
Ajima, R., Sakakibara, Y., Sakurai-Yamatani, N., Muraoka, M. & Saga, Y. Formal proof of the requirement of MESP1 and MESP2 in mesoderm specification and their transcriptional control via specific enhancers in mice. Development 148. https://doi.org/10.1242/dev.194613 (2021).
Lower, K. M. et al. Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc. Natl. Acad. Sci. USA 106, 21771–21776 (2009).
Topfer, S. K. et al. Disrupting the adult globin promoter alleviates promoter competition and reactivates fetal globin gene expression. Blood 139, 2107–2118 (2022).
Bateman, J. R. & Johnson, J. E. Altering enhancer-promoter linear distance impacts promoter competition in cis and in trans. Genetics 222. https://doi.org/10.1093/genetics/iyac098 (2022).
Nakamura, T. et al. Topologically associating domains define the impact of de novo promoter variants on autism spectrum disorder risk. Cell Genom. 4, 100488 (2024).
Ong, C. T. & Corces, V. G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genet. 15, 234–246 (2014).
Zheng, H. & Xie, W. The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol. 20, 535–550 (2019).
Dehingia, B., Milewska, M., Janowski, M. & Pękowska, A. CTCF shapes chromatin structure and gene expression in health and disease. EMBO Rep. 23, e55146 (2022).
Kemp, C. J. et al. CTCF haploinsufficiency destabilizes DNA methylation and predisposes to cancer. Cell Rep. 7, 1020–1029 (2014).
Konrad, E. D. H. et al. CTCF variants in 39 individuals with a variable neurodevelopmental disorder broaden the mutational and clinical spectrum. Genet Med. 21, 2723–2733 (2019).
Davis, L., Onn, I. & Elliott, E. The emerging roles for the chromatin structure regulators CTCF and cohesin in neurodevelopment and behavior. Cell Mol. Life Sci. 75, 1205–1214 (2018).
Filippova, G. N. et al. Tumor-associated zinc finger mutations in the CTCF transcription factor selectively alter tts DNA-binding specificity. Cancer Res. 62, 48–52 (2002).
Bailey, C. G. et al. Structure-function relationships explain CTCF zinc finger mutation phenotypes in cancer. Cell Mol. Life Sci. 78, 7519–7536 (2021).
Poulos, R. C. et al. Functional mutations form at CTCF-cohesin binding sites in melanoma due to uneven nucleotide excision repair across the Motif. Cell Rep. 17, 2865–2872 (2016).
Kaiser, V. B., Taylor, M. S. & Semple, C. A. Mutational biases drive elevated rates of substitution at regulatory sites across cancer types. PLoS Genet. 12, e1006207 (2016).
Guo, Y. A. et al. Mutation hotspots at CTCF binding sites coupled to chromosomal instability in gastrointestinal cancers. Nat. Commun. 9, 1520 (2018).
Cheng, H., Zhang, N. & Pati, D. Cohesin subunit RAD21: from biology to disease. Gene 758, 144966 (2020).
Boudaoud, I. et al. Connected gene communities underlie transcriptional changes in cornelia de lange syndrome. Genetics 207, 139–151 (2017).
Weiss, F. D. et al. Neuronal genes deregulated in Cornelia de Lange Syndrome respond to removal and re-expression of cohesin. Nat. Commun. 12, 2919 (2021).
Waldman, T. Emerging themes in cohesin cancer biology. Nat. Rev. Cancer 20, 504–515 (2020).
Losada, A. Cohesin in cancer: chromosome segregation and beyond. Nat. Rev. Cancer 14, 389–393 (2014).
Viny, A. D. et al. Cohesin members Stag1 and Stag2 display distinct roles in chromatin accessibility and topological control of HSC self-renewal and differentiation. Cell Stem Cell 25, 682–696.e688 (2019).
Wutz, G. et al. ESCO1 and CTCF enable formation of long chromatin loops by protecting cohesin. Elife 9. https://doi.org/10.7554/eLife.52091 (2020).
Cuadrado, A. et al. Specific contributions of cohesin-SA1 and Cohesin-SA2 to TADs and polycomb domains in embryonic stem cells. Cell Rep. 27, 3500–3510.e3504 (2019).
Pang, D., Yu, S. & Yang, X. A mini-review of the role of condensin in human nervous system diseases. Front. Mol. Neurosci. 15, 889796 (2022).
Martin, C. A. et al. Mutations in genes encoding condensin complex proteins cause microcephaly through decatenation failure at mitosis. Genes Dev. 30, 2158–2172 (2016).
Golfier, S., Quail, T., Kimura, H. & Brugués, J. Cohesin and condensin extrude DNA loops in a cell cycle-dependent manner. Elife 9. https://doi.org/10.7554/eLife.53885 (2020).
Cutts, E. E. & Vannini, A. Condensin complexes: understanding loop extrusion one conformational change at a time. Biochem. Soc. Trans. 48, 2089–2100 (2020).
Kakui, Y. et al. Fission yeast condensin contributes to interphase chromatin organization and prevents transcription-coupled DNA damage. Genome Biol. 21, 272 (2020).
Verheul, T. C. J., van Hijfte, L., Perenthaler, E. & Barakat, T. S. The Why of YY1: mechanisms of transcriptional regulation by Yin Yang 1. Front. Cell Dev. Biol. 8, 592164 (2020).
Dong, X. et al. YY1 safeguard multidimensional epigenetic landscape associated with extended pluripotency. Nucleic Acids Res. 50, 12019–12038 (2022).
Schug, J. et al. Promoter features related to tissue specificity as measured by Shannon entropy. Genome Biol. 6, R33 (2005).
Khachigian, L. M. The Yin and Yang of YY1 in tumor growth and suppression. Int. J. Cancer 143, 460–465 (2018).
Pabian-Jewuła, S., Bragiel-Pieczonka, A. & Rylski, M. Ying Yang 1 engagement in brain pathology. J. Neurochem. 161, 236–253 (2022).
Amir, R. E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 23, 185–188 (1999).
Tillotson, R. & Bird, A. The molecular basis of MeCP2 function in the brain. J. Mol. Biol. 432, 1602–1623 (2020).
Horike, S., Cai, S., Miyano, M., Cheng, J. F. & Kohwi-Shigematsu, T. Loss of silent-chromatin looping and impaired imprinting of DLX5 in Rett syndrome. Nat. Genet. 37, 31–40 (2005).
Grubert, F. et al. Landscape of cohesin-mediated chromatin loops in the human genome. Nature 583, 737–743 (2020).
van Arensbergen, J., van Steensel, B. & Bussemaker, H. J. In search of the determinants of enhancer-promoter interaction specificity. Trends Cell Biol. 24, 695–702 (2014).
Ahn, J. H. et al. Phase separation drives aberrant chromatin looping and cancer development. Nature 595, 591–595 (2021).
Chen, M., Liu, X., Liu, Q., Shi, D. & Li, H. 3D genomics and its applications in precision medicine. Cell Mol. Biol. Lett. 28, 19 (2023).
Jia, Q., Chen, S., Tan, Y., Li, Y. & Tang, F. Oncogenic super-enhancer formation in tumorigenesis and its molecular mechanisms. Exp. Mol. Med. 52, 713–723 (2020).
Feehley, T., O’Donnell, C. W., Mendlein, J., Karande, M. & McCauley, T. Drugging the epigenome in the age of precision medicine. Clin. Epigenet. 15, 6 (2023).
Bauer, D. E. et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342, 253–257 (2013).
Frangoul, H. et al. CRISPR-Cas9 gene editing for sickle cell disease and beta-Thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Erdős, G., Pajkos, M. & Dosztányi, Z. IUPred3: prediction of protein disorder enhanced with unambiguous experimental annotation and visualization of evolutionary conservation. Nucleic Acids Res. 49, W297–W303 (2021).
Gregor, A. et al. De novo mutations in the genome organizer CTCF cause intellectual disability. Am. J. Hum. Genet. 93, 124–131 (2013).
Bastaki, F. et al. Identification of a novel CTCF mutation responsible for syndromic intellectual disability - a case report. BMC Med. Genet. 18, 68 (2017).
Guo, J., Cao, B., Xu, X., Wu, F. & Zhu, B. Novel CTCF mutations in Chinese patients with ovarian endometriosis. Mol. Med. Rep. 18, 1031–1036 (2018).
Marshall, A. D. et al. CTCF genetic alterations in endometrial carcinoma are pro-tumorigenic. Oncogene 36, 4100–4110 (2017).
Chatterjee, S. et al. Enhancer variants synergistically drive dysfunction of a gene regulatory network in hirschsprung disease. Cell 167, 355–368.e310 (2016).
Zhang, M. et al. Long-range. Proc. Natl. Acad. Sci. USA 116, 22692–22698 (2019).
Zhang, Z. et al. SNP rs4971059 predisposes to breast carcinogenesis and chemoresistance via TRIM46-mediated HDAC1 degradation. EMBO J. 40, e107974 (2021).
Gao, P. et al. Biology and clinical implications of the 19q13 aggressive prostate cancer susceptibility locus. Cell 174, 576–589.e518 (2018).
Yu, C. Y. et al. A 16q22.1 variant confers susceptibility to colorectal cancer as a distal regulator of ZFP90. Oncogene 39, 1347–1360 (2020).
Giorgio, E. et al. A large genomic deletion leads to enhancer adoption by the lamin B1 gene: a second path to autosomal dominant adult-onset demyelinating leukodystrophy (ADLD). Hum. Mol. Genet. 24, 3143–3154 (2015).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Lupiáñez, D. G., Spielmann, M. & Mundlos, S. Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet. 32, 225–237 (2016).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Gröschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Ji, X. et al. 3D chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell 18, 262–275 (2016).
de Bruijn, S. E. et al. Structural variants create new topological-associated domains and ectopic retinal enhancer-gene contact in dominant retinitis pigmentosa. Am. J. Hum. Genet. 107, 802–814 (2020).
Laugsch, M. et al. Modeling the pathological long-range regulatory effects of human structural variation with patient-specific hiPSCs. Cell Stem Cell 24, 736–752.e712 (2019).
Vicente-García, C. et al. Regulatory landscape fusion in rhabdomyosarcoma through interactions between the PAX3 promoter and FOXO1 regulatory elements. Genome Biol. 18, 106 (2017).
Chai, P. et al. Generation of onco-enhancer enhances chromosomal remodeling and accelerates tumorigenesis. Nucleic Acids Res. 48, 12135–12150 (2020).
Meinel, J. A. et al. Disruption of the topologically associated domain at Xp21.2 is related to 46,XY gonadal dysgenesis. J. Med. Genet. https://doi.org/10.1136/jmg-2022-108635 (2022).
Sun, J. H. et al. Disease-associated short tandem repeats co-localize with chromatin domain boundaries. Cell 175, 224–238.e215 (2018).
Valton, A. L. & Dekker, J. TAD disruption as oncogenic driver. Curr. Opin. Genet. Dev. 36, 34–40 (2016).
Kaiser, V. B. & Semple, C. A. When TADs go bad: chromatin structure and nuclear organisation in human disease. F1000Res 6. https://doi.org/10.12688/f1000research.10792.1 (2017).
Anania, C. & Lupiáñez, D. G. Order and disorder: abnormal 3D chromatin organization in human disease. Brief. Funct. Genom. 19, 128–138 (2020).
Tena, J. J. & Santos-Pereira, J. M. Topologically associating domains and regulatory landscapes in development, evolution and disease. Front. Cell Dev. Biol. 9, 702787 (2021).
Chen, F. et al. Three additional de novo CTCF mutations in Chinese patients help to define an emerging neurodevelopmental disorder. Am. J. Med. Genet C. Semin. Med. Genet. 181, 218–225 (2019).
Oh, S., Oh, C. & Yoo, K. H. Functional roles of CTCF in breast cancer. BMB Rep. 50, 445–453 (2017).
Price, E. et al. An updated catalog of CTCF variants associated with neurodevelopmental disorder phenotypes. Front. Mol. Neurosci. 16, 1185796 (2023).
Zhang, B. N. et al. A cohesin subunit variant identified from a peripheral sclerocornea pedigree. Dis. Markers 2019, 8781524 (2019).
Bonora, E. et al. Mutations in RAD21 disrupt regulation of APOB in patients with chronic intestinal pseudo-obstruction. Gastroenterology 148, 771–782.e711 (2015).
Shah, R. R. & Bird, A. P. MeCP2 mutations: progress towards understanding and treating Rett syndrome. Genome Med. 9, 17 (2017).
Gabriele, M. et al. YY1 haploinsufficiency causes an intellectual disability syndrome featuring transcriptional and chromatin dysfunction. Am. J. Hum. Genet. 100, 907–925 (2017).
Acknowledgements
Soohwan Oh is supported by a Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2022R1C1C1010699) and the MSIT (Ministry of Science and ICT), Korea under the ITRC (Information Technology Research Center) support program (IITP-2023-RS-2023-00258971) supervised by the IITP (Institute for Information & Communications Technology Planning & Evaluation). Figures were created with BioRender.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.J.F., S.O., M.G.R. Funding acquisition: S.O. Supervision: S.O., M.G.R. Visualization: H.L. Writing - original draft: M.J.F., S.O., H.L., T.W. Writing - review & editing: M.J.F., S.O., M.G.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Friedman, M.J., Wagner, T., Lee, H. et al. Enhancer–promoter specificity in gene transcription: molecular mechanisms and disease associations. Exp Mol Med 56, 772–787 (2024). https://doi.org/10.1038/s12276-024-01233-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-024-01233-y
This article is cited by
-
4D nucleome: dynamic three-dimensional genome organization over time
Experimental & Molecular Medicine (2024)