Abstract
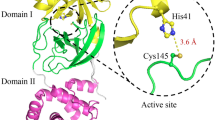
HUMAN cytomegalovirus (hCMV), a herpesvirus, infects up to 70% of the general population in the United States and can cause morbidity and mortality in immunosuppressed individuals (organ-transplant recipients and AIDS patients) and congenitally infected newborns1. hCMV protease is essential for the production of mature infectious virions, as it performs proteolytic processing near the carboxy terminus (M-site) of the viral assembly protein precursor (for a review, see ref. 2). hCMV protease is a serine protease2, although it has little homology to other clans of serine proteases2,3 Here we report the crystal structure of hCMV protease at 2.0Å resolution, and show that it possesses a new polypeptide backbone fold. Ser 132 and His 63 are found in close proximity in the active site, confirming earlier biochemical and mutagenesis studies2. The structure suggests that the third member of the triad is probably His 157. A dimer of the protease with an extensive interface is found in the crystal structure. This structure information will help in the design and optimization of inhibitors against herpesvirus proteases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Fields, B. N. et al. (eds) Virology Vol. 2, ch. 64–73 (Raven, New York, 1990).
Gibson, W., Welch, A. R. & Hall, M. R. T. Perspect. Drug Discov. Design 2, 413–426 (1995).
Rawlings, N. D. & Barrett, A. J. Methods Enzymol. 244, 19–61 (1994).
Pinko, C. et al. J. Biol. Chem. 270, 23634–23640 (1995).
Ollis, D. L. et al. Protein Eng. 5, 197–211 (1992).
Perona, J. J. & Craik, C. S. Protein Sci. 4, 337–360 (1995).
Murzin, A. G., Brenner, S. E., Hubbard, T. & Chothia, C. J. Mol. Biol. 247, 536–540 (1995).
Yamamoto, A. et al. Biochemistry 31, 11305–11309 (1992).
Darke, P. L. et al. J. Biol. Chem. 271, 7445–7449 (1996).
Margosiak, S. A., Vanderpool, D. L., Sisson, W., Pinko, C. & Kan, C.-C. Biochemistry 35, 5300–5307 (1996).
Burck, P. J. et al. J. Virol. 68, 2937–2946 (1994).
Hall, D. L. & Darke, P. L. J. Biol. Chem. 270, 22697–22700 (1995).
Hendrickson, W. A. Science 254, 51–58 (1991).
Jancarik, J. & Kim, S.-H. J. Appl. Crystallogr. 24, 409–411 (1991).
Otwinowski, Z. In Data Collection and Processing (eds Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (SERC Laboratory, Daresbury, 1993).
Tong, L. & Rossmann, M. G. J. Appl. Crystallogr. 26, 15–21 (1993).
Tong, L. J. Appl. Crystallogr. 26, 748–751 (1993).
Jones, T. A. J. Appl. Crystallogr. 11, 268–272 (1978).
Brünger, A. T. The X-PLOR Manual, Version 3.0 (Yale University, New Haven, CT, 1992).
Read, R. J. Acta Crystallogr. A42, 140–149 (1986).
Brünger, A. T. Nature 355, 472–475 (1992).
Carson, M. J. Mol. Graph. 5, 103–106 (1987).
Nicholls, A., Sharp, K. A. & Honig, B. Proteins 11, 281–296 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Tong, L., Qian, C., Massariol, MJ. et al. A new serine-protease fold revealed by the crystal structure of human cytomegalovirus protease. Nature 383, 272–275 (1996). https://doi.org/10.1038/383272a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/383272a0
This article is cited by
-
Extensive subunit contacts underpin herpesvirus capsid stability and interior-to-exterior allostery
Nature Structural & Molecular Biology (2016)
-
Inhibition of a viral enzyme by a small-molecule dimer disruptor
Nature Chemical Biology (2009)
-
Antiherpesvirus drugs: a promising spectrum of new drugs and drug targets
Nature Reviews Drug Discovery (2003)
-
Design of potent selective zinc-mediated serine protease inhibitors
Nature (1998)
-
Conserved mode of peptidomimetic inhibition and substrate recognition of human cytomegalovirus protease
Nature Structural Biology (1998)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.