Abstract
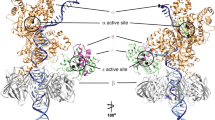
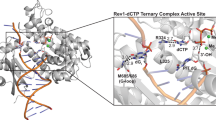
Many candidate unnatural DNA base pairs have been developed, but some of the best-replicated pairs adopt intercalated structures in free DNA that are difficult to reconcile with known mechanisms of polymerase recognition. Here we present crystal structures of KlenTaq DNA polymerase at different stages of replication for one such pair, dNaM-d5SICS, and show that efficient replication results from the polymerase itself, inducing the required natural-like structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
06 June 2012
In the version of this article initially published online, the fourth author was mistakenly credited with equal contribution to that of the first two authors. The error has been corrected in the print, PDF and HTML versions of this article.
References
Hollenstein, M., Hipolito, C.J., Lam, C.H. & Perrin, D.M. Nucleic Acids Res. 37, 1638–1649 (2009).
Seeman, N.C. Annu. Rev. Biochem. 79, 65–87 (2010).
Piccirilli, J.A., Krauch, T., Moroney, S.E. & Benner, S.A. Nature 343, 33–37 (1990).
Xie, J. & Schultz, P.G. Nat. Rev. Mol. Cell Biol. 7, 775–782 (2006).
Rothwell, P.J. & Waksman, G. Adv. Protein Chem. 71, 401–440 (2005).
Echols, H. & Goodman, M.F. Annu. Rev. Biochem. 60, 477–511 (1991).
Goodman, M.F. Proc. Natl. Acad. Sci. USA 94, 10493–10495 (1997).
Kool, E.T. Annu. Rev. Biochem. 71, 191–219 (2002).
Kunkel, T.A. J. Biol. Chem. 279, 16895–16898 (2004).
Seo, Y.J., Hwang, G.T., Ordoukhanian, P. & Romesberg, F.E. J. Am. Chem. Soc. 131, 3246–3252 (2009).
Hirao, I. et al. Nat. Methods 3, 729–735 (2006).
McMinn, D.L. et al. J. Am. Chem. Soc. 121, 11585–11586 (1999).
Lavergne, T., Malyshev, D.A. & Romesberg, F.E. Chemistry 18, 1231–1239 (2012).
Malyshev, D.A., Seo, Y.J., Ordoukhanian, P. & Romesberg, F.E. J. Am. Chem. Soc. 131, 14620–14621 (2009).
Seo, Y.J., Matsuda, S. & Romesberg, F.E. J. Am. Chem. Soc. 131, 5046–5047 (2009).
Malyshev, D.A. et al. Chemistry 16, 12650–12659 (2010).
Matsuda, S. et al. J. Am. Chem. Soc. 129, 10466–10473 (2007).
Wojciechowski, F. & Leumann, C.J. Chem. Soc. Rev. 40, 5669–5679 (2011).
Brotschi, C., Haberli, A. & Leumann, C.J. Angew. Chem. Int. Ed. Engl. 40, 3012–3014 (2001).
Wu, E.Y. & Beese, L.S. J. Biol. Chem. 286, 19758–19767 (2011).
Li, Y., Korolev, S. & Waksman, G. EMBO J. 17, 7514–7525 (1998).
Doublié, S., Tabor, S., Long, A.M., Richardson, C.C. & Ellenberger, T. Nature 391, 251–258 (1998).
Leconte, A.M. et al. J. Am. Chem. Soc. 130, 2336–2343 (2008).
Meyer, A.S., Blandino, M. & Spratt, T.E. J. Biol. Chem. 279, 33043–33046 (2004).
Moran, S., Ren, R.X.F., Rumney, S. & Kool, E.T. J. Am. Chem. Soc. 119, 2056–2057 (1997).
Acknowledgements
We thank the beamline staff of the Swiss Light Source at the Paul Scherrer Institute for their assistance during data collection. This work was supported by the Konstanz Research School Chemical Biology (to K.B.) and the US National Institutes of Health (GM060005 to F.E.R.).
Author information
Authors and Affiliations
Contributions
K.B., D.A.M., T.L., F.E.R. and A.M. conceived of the project, designed the experiments and analyzed the data. K.B., D.A.M., T.L. and P.O. performed chemical synthesis. K.B., W.W. and K.D. performed crystallography studies. T.J.D. performed the NMR experiments, and D.A.M. and T.J.D. performed modeling studies. K.B., D.A.M., A.M. and F.E.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 4818 kb)
Rights and permissions
About this article
Cite this article
Betz, K., Malyshev, D., Lavergne, T. et al. KlenTaq polymerase replicates unnatural base pairs by inducing a Watson-Crick geometry. Nat Chem Biol 8, 612–614 (2012). https://doi.org/10.1038/nchembio.966
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.966
This article is cited by
-
Transcriptional processing of an unnatural base pair by eukaryotic RNA polymerase II
Nature Chemical Biology (2021)
-
The mechanism of the nucleo-sugar selection by multi-subunit RNA polymerases
Nature Communications (2021)
-
New codons for efficient production of unnatural proteins in a semisynthetic organism
Nature Chemical Biology (2020)
-
The Toolbox for Modified Aptamers
Molecular Biotechnology (2016)
-
Cycloadditions for Studying Nucleic Acids
Topics in Current Chemistry (2016)