Abstract
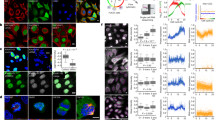
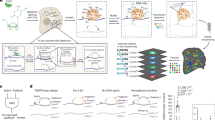
We examined cell cycle–dependent changes in the proteome of human cells by systematically measuring protein dynamics in individual living cells. We used time-lapse microscopy to measure the dynamics of a random subset of 20 nuclear proteins, each tagged with yellow fluorescent protein (YFP) at its endogenous chromosomal location. We synchronized the cells in silico by aligning protein dynamics in each cell between consecutive divisions. We observed widespread (40%) cell-cycle dependence of nuclear protein levels and detected previously unknown cell cycle–dependent localization changes. This approach to dynamic proteomics can aid in discovery and accurate quantification of the extensive regulation of protein concentration and localization in individual living cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Andersen, J.S. et al. Nucleolar proteome dynamics. Nature 433, 77–83 (2005).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Ideker, T. et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292, 929–934 (2001).
Rosenfeld, N., Young, J.W., Alon, U., Swain, P.S. & Elowitz, M.B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005).
Perlman, Z.E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Bannasch, D. et al. LIFEdb: a database for functional genomics experiments integrating information from external sources, and serving as a sample tracking system. Nucleic Acids Res. 32, D505–D508 (2004).
Gerlich, D., Beaudouin, J., Gebhard, M., Ellenberg, J. & Eils, R. Four-dimensional imaging and quantitative reconstruction to analyse complex spatiotemporal processes in live cells. Nat. Cell Biol. 3, 852–855 (2001).
Tvarusko, W. et al. Time-resolved analysis and visualization of dynamic processes in living cells. Proc. Natl. Acad. Sci. USA 96, 7950–7955 (1999).
Conrad, C. et al. Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 14, 1130–1136 (2004).
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002).
Huh, W.K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Chen, X. & Murphy, R.F. Objective clustering of proteins based on subcellular location patterns. J. Biomed. Biotechnol. 2005, 87–95 (2005).
Menges, M., de Jager, S.M., Gruissem, W. & Murray, J.A. Global analysis of the core cell-cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell-cycle control. Plant J. 41, 546–566 (2005).
Spellman, P.T. et al. Comprehensive identification of cell-cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell 9, 3273–3297 (1998).
Cho, R.J. et al. A genome-wide transcriptional analysis of the mitotic cell-cycle. Mol. Cell 2, 65–73 (1998).
Rustici, G. et al. Periodic gene expression program of the fission yeast cell-cycle. Nat. Genet. 36, 809–817 (2004).
Cho, R.J. et al. Transcriptional regulation and function during the human cell-cycle. Nat. Genet. 27, 48–54 (2001).
Whitfield, M.L. et al. Identification of genes periodically expressed in the human cell-cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002).
Jarvik, J.W. et al. In vivo functional proteomics: mammalian genome annotation using CD-tagging. Biotechniques 33, 852–860 (2002).
Jarvik, J.W., Adler, S.A., Telmer, C.A., Subramaniam, V. & Lopez, A.J. CD-tagging: a new approach to gene and protein discovery and analysis. Biotechniques 20, 896–904 (1996).
Scheel, J.R., Ray, J., Gage, F.H. & Barlow, C. Quantitative analysis of gene expression in living adult neural stem cells by gene trapping. Nat. Methods 2, 363–370 (2005).
Whitney, M. et al. A genome-wide functional assay of signal transduction in living mammalian cells. Nat. Biotechnol. 16, 1329–1333 (1998).
Friedrich, G. & Soriano, P. Promoter traps in embryonic stem cells: a genetic screen to identify and mutate developmental genes in mice. Genes Dev. 5, 1513–1523 (1991).
Skarnes, W.C., Auerbach, B.A. & Joyner, A.L. A gene trap approach in mouse embryonic stem cells: the lacZ reported is activated by splicing, reflects endogenous gene expression, and is mutagenic in mice. Genes Dev. 6, 903–918 (1992).
Clyne, P.J., Brotman, J.S., Sweeney, S.T. & Davis, G. Green fluorescent protein tagging Drosophila proteins at their native genomic loci with small P elements. Genetics 165, 1433–1441 (2003).
Morin, X., Daneman, R., Zavortink, M. & Chia, W. A protein trap strategy to detect GFP-tagged proteins expressed from their endogenous loci in Drosophila. Proc. Natl. Acad. Sci. USA 98, 15050–15055 (2001).
Zambrowicz, B.P. et al. Disruption and sequence identification of 2,000 genes in mouse embryonic stem cells. Nature 392, 608–611 (1998).
Nomura, M. Ribosomal RNA Genes, RNA Polymerases, Nucleolar Structures, and Synthesis of rRNA in the Yeast Saccharomyces cerevisiae (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Muratani, M. et al. Metabolic-energy-dependent movement of PML bodies within the mammalian cell nucleus. Nat. Cell Biol. 4, 106–110 (2002).
Shedden, K. & Cooper, S. Analysis of cell-cycle-specific gene expression in human cells as determined by microarrays and double-thymidine block synchronization. Proc. Natl. Acad. Sci. USA 99, 4379–4384 (2002).
Liron, Y., Paran, Y., Zatorsky, N.G., Geiger, B. & Kam, Z. Laser autofocusing system for high-resolution cell biological imaging. J. Microsc. 221, 145–151 (2006).
Ghosh, R.N. et al. Quantitative cell-based high-content screening for vasopressin receptor agonists using transfluor technology. J. Biomol. Screen. 10, 476–484 (2005).
Acknowledgements
We thank Quantomix for plasmid construction; Z. Kam for microscopy guidance; B. Zimmerman for cytoskeleton staining; E. Ariel and A. Sharp from the Weizmann flow cytometry unit for assistance, and M. Springer, P. Bordalo, O. Zuk and members of the Alon lab for discussions of the manuscript. We thank the Kahn Family Foundation and the Israel Science Foundation for support. R.M. thanks the Horowitz Complexity Science Foundation for support. A.E.C. thanks the Novartis Life Science Foundation for support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Integration positions of the YFP CD-tag. (PDF 107 kb)
Supplementary Fig. 2
Distribution of cell cycle lengths for 20 nuclear clones. (PDF 61 kb)
Supplementary Fig. 3
Determination of the relative durations of cell cycle phases. (PDF 91 kb)
Supplementary Fig. 4
Automatic segmentation based on fluorescence levels in the nuclei. (PDF 135 kb)
Supplementary Fig. 5
Raw accumulation measurements without any correction. (PDF 84 kb)
Supplementary Fig. 6
Synchrogram before cell centering and rotation, and before elimination of neighboring cells. (PDF 441 kb)
Supplementary Table 1
Localization of CD-tagged clones compared to previous studies. (PDF 108 kb)
Supplementary Video 1
Panel of 9 proteins, shown for one complete division, approximately 20 hours. Protein identities (localizations), first row, left to right: MSN (cytoplasm), BSG (plasma membrane), ATP1A1 (plasma membrane). Second row: FBL (nucleoli), RPL4 (cytoplasm and nucleoli), RTN4 (endoplasmic reticulum). Third row: H2AFV (DNA bound), USP7 (nucleus, nuclear bodies), LMNB1 (nuclear membrane). (MPG 1091 kb)
Supplementary Video 2
Histone H2AFV transmitted light. Duration shown: 42 hours. (MPG 4861 kb)
Supplementary Video 3
Histone H2AFV fluorescence. Duration shown: 42 hours. (MPG 2360 kb)
Supplementary Video 4
USP7 translocation to nuclear bodies. Duration shown: 42 hours. (MPG 5660 kb)
Rights and permissions
About this article
Cite this article
Sigal, A., Milo, R., Cohen, A. et al. Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins. Nat Methods 3, 525–531 (2006). https://doi.org/10.1038/nmeth892
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth892
This article is cited by
-
Spatial proteomics: a powerful discovery tool for cell biology
Nature Reviews Molecular Cell Biology (2019)
-
Dynamic proteomics reveals bimodal protein dynamics of cancer cells in response to HSP90 inhibitor
BMC Systems Biology (2017)
-
Improving drug discovery with high-content phenotypic screens by systematic selection of reporter cell lines
Nature Biotechnology (2016)
-
Survey statistics of automated segmentations applied to optical imaging of mammalian cells
BMC Bioinformatics (2015)
-
CellProfiler Tracer: exploring and validating high-throughput, time-lapse microscopy image data
BMC Bioinformatics (2015)