Abstract
Background
There is currently a lack of experimental evidence for horizontal gene transfer (HGT) mechanisms in the human gut microbiota. The aim of this study was therefore to experimentally determine the HGT potential in the microbiota of a healthy preterm infant twin pair and to evaluate the global occurrence of the mobilized elements.
Methods
Stool samples were collected. Both shotgun metagenome sequencing and bacterial culturing were done for the same samples. A range of experimental conditions were used to test DNA transfer for the cultured isolates. Searches for global distribution of transferable elements were done for the ~120,000 metagenomic samples in the Sequence Read Archive (SRA) database.
Results
DNA transfer experiments demonstrated frequent transmission of an ESBL encoding IncI1 plasmid, a high copy number ColEI plasmid, and bacteriophage P1. Both IncI1 and ColE1 were abundant in the stool samples. In vitro competition experiments showed that transconjugants containing IncI1 plasmids outcompeted the recipient strain in the absence of antibiotic selection. The SRA searches indicated a global distribution of the mobilizable elements, with chicken identified as a possible reservoir for the IncI1 ESBL encoding plasmid.
Conclusion
Our results experimentally support a major horizontal transmission and persistence potential of the preterm infant gut microbiota mobilome involving genes encoding ESBL.
Similar content being viewed by others
Introduction
It has been estimated that by 2050 more humans will die due to antibiotic-resistant bacteria than cancer if no action is taken.1 Resistant Enterobacteriaceae represents the largest antimicrobial resistance health threat in the world, especially within the pediatric population.1 Although pathogenic Enterobacteriaceae have been extensively studied, there is a major knowledge gap related to the non-pathogenic strains serving as a reservoir for mobile genetic elements (MGEs), and multidrug resistance.2 Recent culture independent work has indicated high levels of horizontal gene transfer (HGT) in the human gut.3 This enables the potential for transmission of antibiotic resistance (ARG) and virulence genes within the preterm infant gut microbiota.4
Preterm infants have an immature immune system and are particularly vulnerable to a range of infections, for which antibiotics are the only treatment option.5,6 Furthermore, the preterm gut microbiota has low complexity, being in many instances dominated by a single bacterium,7 with Enterobacteriaceae being predictive for disease development.7,8
The Enterobacteriaceae contains a rich repertoire of conjugative plasmids with the potential of transmission.9 Due to the presence of their advanced replication and transfer systems, conjugative plasmids can replicate and transfer autonomously.10 Plasmid addiction systems, such as plasmid partitioning, toxin/antitoxin, and stability genes, ensure the stability of these plasmids within the microbial populations.10 They are often regarded as parasites of the bacterial cell. In addition, these plasmids can also harbor accessory elements, such as transposons and integrons, which can carry several ARGs as gene cassettes.11 In some cases, the transfer of specific plasmids is dependent on the conjugative machinery of other plasmids for their mobilization;10 in addition, bacteriophages may also utilize these conjugal pili as targets for cell entry.12
Therefore, the main aim of our study was to experimentally determine the HGT potential of MGEs within the gut microbiota of a preterm twin pair. We focused on conjugative plasmids since these elements are known for their carriage of multidrug resistance genes and virulence factors,13 in addition to serving as vectors for transmission of non-conjugative plasmids and bacteriophages.14
Methods
The experimental approach is outlined in Fig. 1. Details about quantitative PCR, DNA sequencing, and proteome analyses are presented in Supplementary Methods.
Outline of experimental design. One stool sample was collected from each of a preterm twin pair. Shotgun metagenome sequencing bacterial culturing was done in parallel. A range of conditions were used for DNA transfer. The DNA transfer was determined by subtracting the genetic content of the recipient strain, and the quantity determined by mapping against the shotgun data. The mobilizable elements were subsequently identified for all sequences present in the Sequence Read Archive (SRA) database, with inference about the distribution being deduced from the assocaited metaddata. Finally stability experiments were conducted testing the stability of the transmitted elements without antibiotic selective pressure
Sample description
Fecal samples were collected 20 days after birth from a preterm twin pair (Twin I and II) that was part of a prospective, single-center, observational study at the University and Polytechnic Hospital La Fe in Valencia, Spain. The study protocol was approved by the Comité de Ética e Investigación Médica (CEIM) and parents approved and signed informed consent in all cases. They were born preterm (gestational age 30) by emergency cesarean section. Weight at birth was 1410 g (Twin I) and 1630 g (Twin II), respectively, and both were breastfed. No antibiotics were given to the babies prior to sampling. The collected fecal samples were frozen within 1 h and kept at −80 °C for later analysis.
Bacterial strain isolation from fecal samples
Mueller Hinton (MH) agar (Sigma-Aldrich, Madrid, Spain) containing sulfonamide (300 µg/ml) was used to plate 0.2 g of fecal sample diluted to up to 10−4 from the corresponding twins in order to select integrons, which are commonly associated with ARG and MGEs. The plates were incubated at 37 °C overnight. Colonies (22 from Twin I and 52 from Twin II) were picked to cover all the colony morphologies from the 10−3 and 10−4 dilutions and streaked onto fresh MH agar plates to verify pure culture. All the isolates were species identified by Sanger sequencing of the 16S ribosomal RNA gene V3 and V4 regions by GATC (Konstanz, Germany). The isolated strains were then stored with 18% glycerol at −80 °C until further analysis.
Antibiotic susceptibility test for the isolated strains
Antibiotic susceptibility was determined for the 74 isolates using disk diffusion method according to EUCAST (http://www.eucast.org/).15 The susceptibilities of the isolates were tested for six different antibiotics classes belonging to penicillins (amoxicillin-clavulanic acid; 30 µg/disc); cephalosporin (cefpodoxime; 10 µg/disc); fluoroquinolones (ciprofloxacin; 5 µg/disk); aminoglycosides (gentamicin; 5 µg/disk); trimethoprim; 5 µg/disk and sulfamethoxazole; 25 µg/disc. The antibiotic susceptibility cartridges were obtained from Oxoid, Thermo Fisher Scientific (Waltham, MA, USA). For testing, bacterial suspensions were adjusted to a turbidity of 0.5 McFarland standard and inoculated onto MH agar plates (Sigma-Aldrich, Norway). The antimicrobial discs were placed on the surface of the agar plate and was incubated at 37 °C overnight. The diameter of the zones of inhibition surrounding the antimicrobial discs were interpreted according to the EUCAST guidelines.16
DNA extraction
For the DNA isolation, 200 µl of the overnight-incubated broth (or 100 mg stool sample) were mixed with 200 µl of S.T.A.R buffer (Roche, Oslo, Norway). In addition to this, 0.25 g of acid-washed glass beads <106 µm (Sigma-Aldrich) was added and the cells were lysed in FastPrep96 (MP Biomedicals, France) at 1800rpm for 40 s for three rounds. The lysed cells were centrifuged at 13,000 rpm for 5 min and 50 µl of the supernatant was used for the DNA isolation. An automated protocol based on paramagnetic particles (LGC Genomics, UK) was used for the DNA isolation in a KingFisher Flex instrument (Thermo Fisher Scientific). The concentration of the eluted DNA (1.5–30.6 ng/µl) was determined by fluorescence using a Qubit System (Invitrogen). The DNA was then stored at −40 °C until further use.
Plasmid extraction
Twenty seven strains were grown in LB broth overnight using a plasmid extraction protocol modified from Zaman et al. 17 The cells were pelleted by centrifugation and resuspended in 100 µl resuspension buffer (50 mM glucose, 10 mM EDTA, and 10 mM Tris-Cl, pH 8.0). Then, 200 µl lysis solution (0.2 M NaOH and 1% sodium dodecyl sulfate) was added. For all the post-lysis steps gentle inversion of the tube was done to mix. One-hundred fifty microliters of 7.5 M ammonium acetate and 150 µl chloroform was added and the samples were left on ice for 10 min. Then, the tubes were centrifuged for 10 min at 5000 rpm. The supernatant was transferred to a new tube containing 200 µl precipitation solution (30% polyethylene glycol 8000 and 1.5 M NaCl). The tube was then left on ice for 15 min before centrifuging 5000 rpm for 5 min to pellet the DNA. Finally, the DNA was resuspended in 50–100 µl deionized sterile water.
Transmission experiment
The recipient strain used in this study was an Escherichia coli DH5α-RifR, which was resistant to 320 µg/ml rifampicin. We used both solid agar mating and liquid broth mating. For the liquid mating, 500 µl overnight culture of the recipient and 10 µl of the donor were mixed in 4 ml of LB broth and incubated at 37 °C for 4, 8, and 24 h. For the solid agar mating, 1 µl loop of donor and recipient colonies were mixed together on LB agar and incubated at 37 °C for 4, 8, and 24 h. The mixed colonies were diluted up to 10−2 using phosphate-buffered saline. To select for the transconjugants, two different methods were used. In the first experiment, the mixtures from the liquid and solid agar mating were grown on MH agar plates containing 320 µg/ml rifampicin. Discs with antimicrobial agents corresponding to the resistance profiles of the donor strains were placed onto the surface of the agar plates. Presumptive transconjugants were picked from the colonies growing within the inhibition zones on the rifampicin-containing plates. For the rest of the conjugation experiments, a selective antibiotic was added in the MH agar together with the rifampicin. In this way, the number of transconjugants could be counted and the transmission efficiency calculated (colony-forming unit (CFU)/ml of transconjugants divided by CFU/ml of donor). Selection antibiotics used were 300 µg/ml sulfamethoxazole or 1 µg/ml cefotaxime.
To confirm the colonies growing inside the inhibition zones or on the selective agar were transconjugants and not mutant donors, the colony sizes were examined, as transconjugant colonies are much smaller than the donor colonies. In addition, the colonies were streaked out on lactose agar (Norwegian Veterinary Institute, Oslo, Norway) to test the ability for lactose fermentation. The donor strains, unlike the recipient, are able to ferment lactose and will hence give a yellow change of color in the blue lactose agar. In total, six transmission experiments were done.
The susceptibility to the DNA-damaging agent mitomycin c was tested on donor strain L-II and the recipient DH5α-RifR. The donors were grown in concentrations up to 2 µg/ml, while the recipient was grown in concentrations up to 0.05 µg/ml.
The transmission experiments are summarized in Table S4.
Competition experiment
Transconjugants were grown to stationary phase at 37 °C in 5 ml MH broth in the background of the recipient strain without transconjugation elements. Then it was diluted to 1.5 × 108 CFU/ml based on OD measurements, and regrown to stationary phase. The number of generations was calculated based on change in OD. The process was repeated until 100 generations was reached. MGEs were quantified by quantitative PCR (qPCR) from the first inoculation, and then after 15, 30, 45, 60, 75, 90, and 100 generations.
Statistical methods
Statistical data analyses were performed using the MATLAB® R2016a software (The MathWorks Inc., MA, USA). Curve fitting for competition experiments were done using third-order polynomials (Excel 2013, Microsoft Inc., WA, USA). Specific statistical tests are noted in the figure legends or tables, as applicable.
Searches in the SRA database
A search was done for both IncI1 plasmid and P1 bacteriophage using the recently developed SRA searching tool18 to identify similar sequences among the available raw sequence data from metagenomic samples in the NCBI SRA. The search tool uses PARTIE19 to separate metagenomic and amplicon sequences, bowtie2 for read alignment,20 and the compute resources of XSEDE.21
The reads were filtered in order to ensure hits for complete sequence elements were high accuracy read hits (alignment score >40), with at least 50 read hits covering the whole sequence elements (with hits within 5000 bp of both ends of the query sequence). The final filterings were done in the MATLAB programming environment.
Results
Shotgun metagenome sequencing
For the collective metagenome assembly of the stool microbiota for Twin I and II, we obtained 1.8 billion nucleotides from 8.3 million reads with an average read length of 222 bp. We obtained 185 contigs (>1000 bp) after assembly, with a collective size of 5.3 million bp. The N50 was 64,272 bp, indicating assembly of the majority of the metagenome.
The shotgun analyses revealed highly similar microbiota composition for the twins, with Spearman’s correlation of 0.94 (p < 0.0005) for sequence assembly coverage.
RAST annotation of the contigs indicate that about 1.6% of all the annotated genes belonged to MGEs, including plasmids, transposable elements, and bacteriophages. Virulence, disease, and defense genes accounted for about 3% of all the annotated genes, these included adhesion, invasion, toxin, bacteriocin, and antibiotic resistance genes.
Bacterial isolates and antimicrobial susceptibility tests
In total, 74 strains were isolated from the fecal samples, with 22 strains originating from preterm Twin I and 52 strains from preterm Twin II (Table S2). From the qPCR screening, 45 (Twin I, n = 9; Twin II, n = 36) of the 74 strains were identified as E. coli, while the rest of the strains belonged to Enterococcus spp. (n = 29) or Staphylococcus epidermidis (n = 1). The antibiotic susceptibility testing showed 71 strains resistant to at least one antimicrobial agent, representing 14 patterns (patterns I–XIV). Resistance to cephalosporin was most prevalent, as 89% (n = 66) of the isolates were resistant to cefpodoxime. We found that 23% (n = 17) of the isolates were resistant to gentamicin. All the isolates resistant to amoxicillin-clavulanic acid (n = 19) were also resistant to cefpodoxime (Table S2).
Genome sequencing of strain isolates
Seventeen strains (Twin I, n = 6; Twin II, n = 11) representing all the resistance patterns I to XIV were selected for genome sequencing (Table S3). These comprised 12 Escherichia coli and 5 Enterococcus faecalis. On average, 827,634 reads were generated per genome with read length from 35 301 bp. We obtained an average of 73 contigs over 1000 bp in length. The average N50 length was 210,522 bp, with at least 10 contigs ≥N50 length/sample.
The contigs from all the strains were submitted to PlasmidFinder for detection of plasmid incompatibility groups in the genome sequences. Only the E. coli strains contained identifiable conjugative plasmids (Table S3). The strain J-II harbored IncFII and IncFIB conjugative plasmids. The rest of the strains contained IncFIB and IncI1 conjugative plasmids (Table S3).
In vitro DNA transfer experiments
Six of the E. coli strains representing AR patterns V, IX, XII, and XIII were chosen for DNA transfer experiments. The transfer experiments involved conjugations under different conditions, including solid- and liquid-phase mating, in addition to DNA damage induction (see Table S4 for experimental details).
All the selected transconjugants, except transconjugant 16 (L-II), showed transfer of the IncI plasmid. In transmission experiment numbers 1 to 4, the P1 bacteriophage transferred to 24 of 49 transconjugants when using sulfamethoxazole for selection, while when using cefotaxime as the selective antibiotic (experiments 5 and 6) no P1 transfer was detected. The ColEI plasmid transferred to 114 out of 138 transconjugants—irrespective of antibiotic selection. However, neither the P1 bacteriophage nor the ColEI plasmid transferred when using solid media for the transfer experiments (Table S5).
In transfer experiments 2 to 6, transconjugants were also tested for ESBL production. Transconjugants were scored as ESBL positive when resistance to cefotaxime, ceftazidime, and cefpodoxime was observed, and had an increase of >5 mm in zone diameter for cefpodoxime/clavulanic acid combination disk compared to the zone diameter of cefpodoxime alone. Of all of the 123 transconjugants tested, 98 (78%) showed the ESBL resistance phenotype (Table S5).
Two conjugation experiments (numbers 4 and 5) with mitomycin c were performed to determine if DNA damage stress has an effect on transmission efficiency. There was an increase in the overall DNA transfer efficiency with increasing mitomycin c concentration (Table S5).
Genome sequencing of transconjugants
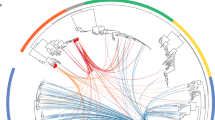
To address the heterogeneity of ARG and MGE, 27 of the transconjugants were shotgun sequenced. The pooled transconjugant plasmid extracts were first sequenced. From the sequences of the pooled plasmid extracts, 18 DNA contigs were detected with a collective size of 223,997 bp. Then, each individual transconjugant was mapped towards the 18 contigs derived from the plasmid extracts. Contigs belonging to the same genetic element were subsequently identified by hierarchal clustering based on their coverage in the different transconjugants. These analyses revealed four main clusters corresponding to bacteriophage P1, transposon Tn21, conjugative plasmid IncI1, and the ColEI-like plasmid (Fig. 2), as confirmed by BLAST searches (Table S6).
Four distinct clusters of genetic elements transmitted in the conjugation experiments. Clustering was based on the coverage of the contigs in the different transconjugants. The dendrograms were derived based on average clustering of Z-scores using Heatmapper40
Abundance of mobilizable elements in infant stool
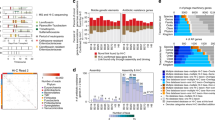
To determine the abundance of the mobilizable elements in the original stool samples the elements were mapped towards the metagenome contigs derived from the stool samples. The metagenome contigs mapping to Tn21, IncI1, and ColEI elements were all overrepresented with respect to coverage (abundance), with the ColEI contig mapping having the overall highest coverage in the original samples. In contrast, bacteriophage P1 seemed to have low coverage in the stool samples, and was not detected in any of the contigs (Fig. 3).
Gene annotations of the mobilized elements
The mobilizable elements were annotated based on shotgun sequence assemblies from both the bacterial strains isolated and the transconjugants.
IncI1 conjugative plasmid
The contigs related to the IncI conjugative plasmid harbored all the transfer genes (traA-traY) and the pilus genes (pil genes) indicating the plasmid, and the ESBL encoding gene (blaSHV-12) it carries, can likely transfer to other cells. This de novo assembled IncI plasmid consisted of a complex transfer system extending to over 50 kb with two types of conjugative pilus regions (Fig. S7a). In addition to the presence of the transfer system, the de novo assembled IncI plasmid also harbored a plasmid SOS system (psiA-psiB family) and replication initiation genes. The same type of assembled IncI plasmid was observed across all of the 11 strains that had the IncI plasmid (Fig. S1a).
P1 bacteriophage
The contigs related to the bacteriophage were identified as a P1 bacteriophage with phage capsid, tail and baseplate proteins, together with other genes necessary for phage particle formation. The lytic repressor protein C1 was present, in addition to a prophage addiction system-like toxin/antitoxin Doc/Phd system (Fig. S1b). The assembled P1 bacteriophage from strain L-II was compared to the assembled P1 bacteriophages from F-I, H-II, C-II, G-II, and L-II. The presence of bacteriophages was observed in five of the six strains, while the sixth strain (I-I) was identical over the same regions as the other five, but contained additional genes (Fig. S1b). These genes include a phage lysin protein (EC 3.2.1.17 lysozyme), a phage portal protein, phage tail fiber proteins, and additional mobile element proteins (Fig. S1b).
ColE1-like plasmid
A ColEI-like plasmid was detected in the strains after shotgun sequencing of plasmid extracts from the transconjugants. The genes harbored by the plasmid included a putative replication regulatory protein, an IncI1 plasmid conjugative transfer pilus-tip adhesion protein, a mobile element protein, and a few hypothetical proteins (Fig. S1d). The plasmid was present in most of the strains isolated from the twins (Table S3). However, no antibiotic resistance or bacteriocin genes were located on the ColEI-like plasmid, nor were any additional transfer genes identified.
Tn21 transposon
A Tn21 transposon was present in all the strains that showed multidrug resistance patterns (resistance towards more than two antibiotics). The AR genes associated with the transposon were blaTEM-1B conferring β-lactamase resistance, sul3 conferring sulfonamide resistance, dfrA14 conferring trimethoprim resistance, mph(A) conferring macrolide resistance, cmlA1 conferring chloramphenicol resistance, and aadA1/aadA2 conferring streptomycin and spectinomycin resistance. In addition, the transposon harbored an integron integrase gene (Fig. S1e). In the strains used as donors in the transmission experiments, the transposon was present in contigs associated to the IncI1 conjugative plasmid. However, after transmission experiments the transposon was present in contigs related to both the IncI1 and the P1 bacteriophage, or on the P1 bacteriophage alone.
Proteome analyses
Proteome analyses were conducted in order to address the overall host influence of the mobilized elements. These analyses support a low burden of the transconjugants, with the protein composition showing a correlation coefficient ≥0.98 between the recipient strain and the transconjugants (Fig. S2). The IncI1-encoded pilin (PilS) was the only mobility associated protein detected, although at a low level (<0.1%) in the transconjugants. For antibiotic resistance, no proteins were identified for streptomycin resistance in transconjugant #16 and 17, and β-lactamases in transconjugant #17 and 55.
In vitro competition experiments
Finally, in order to investigate the competitiveness of transconjugants in the absence of antibiotic selection competition experiments were performed. The transconjugants were selected to cover the mobilizable elements detected in this work. For all competition experiments, the abundance of IncI1 plasmid increased after about 60 generations, even in the cases with only 1% IncI1 plasmid as the start concentration (Fig. 4a–c). Although not as pronounced, ColEI plasmids also showed the same pattern (Fig. 4h, i), while for P1 bacteriophage there seemed to be a concentration-dependent decline. Furthermore, it seemed that the P1 bacteriophage was more stable when the IncI1 plasmid was not present (Fig. 4d–f).
The IncI1 plasmid is persistent in the absence of antibiotic selection. Four transconjugants with different mobile genetic element profiles and antibiotic resistance patterns were chosen for a plasmid/phage stability experiments (transconjugants 16, 22, 30, and 55). Three parallel experiments were conducted for each set of strains. Parallel 1: contained the transconjugant only (a, d, and g), Parallel 2: contained 1% transconjugant + 99% DH5α-RifR (b, e, and h), Parallel 3: contained 99% transconjugant + 1% DH5α-RifR (c, f, and i). a–c show the relative amount of IncI1, d–f the relative amount of P1, and g–i the relative amount of ColEI
Global distribution of IncI1 and P1
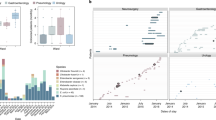
The search using the SRA search toolkit resulted in 60,000 and 30,000 metagenomic sample hits for IncI1 plasmid and P1 bacteriophage, respectively. Further filtering by alignment quality and number of hits led to 351 samples (SRRs) satisfying the matching criteria for IncI1 and 401 samples for P1. These samples belonged to 70 and 78 study projects (SRPs), respectively. Both IncI1 and P1 were identified globally in human, environment, and animal sample sources (Fig. 5, Table S7 and S8). For IncI1 we identified a relative overabundance of samples associated with the infant gut (Fig. 5a), while bacteriophage P1 seemed more associated with adult gut samples originating from a wider range of study projects (Fig. 5b). Quantitatively, one study project (SPR114955) of preterm infants show a mean level of IncI1 mapping fragments >1% (Fig. S3a). Interestingly, P1 showed particularly high levels in several study projects, representing both adult and infant gut microbiota datasets (Fig. S3b).
Prevalence of InciI1 (a) and P1 (b) in the Sequence Read Archive (SRA) database. The pie charts illustrate the distribution samples with IncI1 (a) and P1 (b) across study projects. The study projects representing >1% of the samples are color coded, while those >2% are marked with names. The SRA mining was done in April 2019
A Blast search for the annotated IncI1 sequence (accession number MH422553) in NCBI gave closest hit (97% query cover, 99.99% identity, and total score of 1.870e + 05) to a plasmid isolated from an E. coli strain found in cecal contents of broiler chicken (accession number MK070495.1).22 Furthermore, the chicken metagenome study project (SRP081435) identified by the SRA search indicated that seven out of ten samples contained the IncI1 plasmid.
Discussion
Although there are several lines of bioinformatic evidence of extensive HGT in the human commensal gut microbiota, this has not yet been experimentally proven.3,4,23,24,25 In the current work, we have experimentally shown frequent transmission of MGE between gut-associated bacteria and laboratory strains. Also, mapping these MGE elements back to the original samples of isolation have demonstrated their importance in the commensal gut microbiota in the infants from which they were originally isolated. Furthermore, searches in the SRA database identified the mobilizable elements as prevalent in human gut-associated datasets, supporting a widespread distribution of the elements identified in the current work.
SRA searches identified the IncI1 plasmid as prevalent in both infant and preterm infant study projects, with nearly half of the hits coming from these sources. Quantitatively the level of IncI1 was also higher in the infant study projects. In addition to the human infant datasets, we identified IncI1 as highly prevalent in a chicken gut metagenome study project. Particularly intriguing was the identification of a highly mobilizable IncI1 plasmid in chickens,22 nearly identical to the plasmid identified in our work. The IncI1-containing chickens were from Belgium and the Netherlands, representing nearly one-quarter of the world’s export of fresh chicken (www.worldstopexports.com/chicken-exports-by-country). This supports chickens from these countries as a possible reservoir for transmission of ESBL encoding genes to the microbiome of humans, potentially through the food chain.26
Although antibiotic usage in food production has become restricted, antibiotic resistance is surprisingly persistent.27 For the IncI1 plasmid identified here, persistence can potentially be explained by the plasmid-containing cells being able to outcompete cells without plasmids even in the absence of antibiotic selection pressure. IncI1 plasmids have been notoriously associated with the spread of multidrug resistance in both human and animal populations.28,29,30,31,32 The mechanisms for their ability to outcompete non-plasmid carrying strains remains unknown. One possibility, however, could be high frequency of conjugal transfer.33 The transmission pattern of Tn21 also provides evidence for frequent transposition during the transmission; Tn21 is identified as a major part of the floating genome.34
Upon sequencing of plasmids from the transconjugants, we also discovered the transmission of a ColEI-like plasmid. This plasmid is known to co-transfer with conjugative plasmids.10,35 The plasmid was also highly prevalent in the stool samples, supporting the high transmission potential. Although the plasmid does not contain antibiotic resistance genes, it has been suggested to be involved in cell division, and regulation of indole production.36
The relatively high frequency of bacteriophage P1 transmission during conjugation also illustrates mobility of this bacteriophage. Bacteriophage P1 was the first transducing bacteriophage discovered, serving as a model organism for HGT for decades.37 Despite extensive investigations, the transmission mechanisms during natural conditions remains unknown. Several attempts on mobilizing wild-type bacteriophages have failed.38 However, in this study we detected a high frequency of concurrent IncI1 plasmid and P1 phage transmission during liquid conjugation, suggesting the co-transfer mechanisms. This is also supported for the SRA searches where bacteriophage P1 showed the highest prevalence for study projects prevalent in IncI1. P1, however, was also detected in study projects where IncI1 plasmids were not detected, suggesting that P1 is not strictly dependent on IncI1 plasmid. Interestingly, P1 showed high levels in a study project involving fecal microbiota transfer (FMT), with the highest levels representing more than 1% of the reads. This may indicate conditions of P1 induction during FMT, and that P1 may affect the efficiency of FMT.39 Similar mechanisms could also potentially explain why a very low amount of P1 was detected in the original stool samples, while a relatively high frequency of P1 was detected for the isolates. For the isolates case, the conditions used for culturing from stool samples could potentially induce P1 transmission during culturing.
The extensive horizontal transmission potential of the preterm infant commensal gut microbiota detected here illustrates the challenges in preventing transmission of antibiotic resistance genes. Likely much of the transmission occurs without antibiotic selection. Thus, given resistance genes that are associated with elements such as the Tn21 transposon and integrons can co-transfer with selfish MGEs (elements that can transmit themselves); this may help explain the persistence of antibiotic resistance, even in the absence of selection pressure by the use of antibacterial agents.
In conclusion, this study has demonstrated frequent transmission in commensal microbiota as a persistence mechanism, supporting previous in vitro experiments.33 This study therefore provides further support that future strategies to prevent antibiotic resistance transmission should not only target the vectors of transmission, but also the resistance genes and mechanisms of resistance themselves. In particular, Tn21 transposons and IncI1 plasmids should be considered as elements of special importance.
Disclaimer
The views presented in this manuscript do not necessarily reflect those of the U.S. Food and Drug Administration.
Data availability
The following annotated sequences were deposited in the NCBI database. IncFIB from L-II was deposited with accession number MH422552, IncI1 from L-II with accession number MH422553, P1 bacteriophage from L-II with accession number MH445381, and P1 from transconjugant 2 (L-II) with accession number MH445380. Sequencing treads for the shotgun sequence data for the isolated bacterial strains and the transconjugants are deposited in Sequence Read Archive (SRA) with accession number SRP148649.
References
Tacconelli, E. & Magrini, N. Global Priority List of Antibiotic-resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics (World Health Organization, Geneva, 2017).
Logan, L. K. & Weinstein, R. A. The epidemiology of carbapenem-resistant enterobacteriaceae: the impact and evolution of a global menace. J. Infect. Dis. 215, S28–S36 (2017).
Brito, I. L. et al. Mobile genes in the human microbiome are structured from global to individual scales. Nature 535, 435−439 (2016).
Ravi, A. et al. Association of the gut microbiota mobilome with hospital location and birth weight in preterm infants. Pediatr. Res. 82, 829–838 (2017).
Morrow, A. L. et al. Early microbial and metabolomic signatures predict later onset of necrotizing enterocolitis in preterm infants. Microbiome 1, 13 (2013).
Westerbeek, E. A. et al. The intestinal bacterial colonisation in preterm infants: a review of the literature. Clin. Nutr. 25, 361–368 (2006).
Ward, D. V. et al. Metagenomic sequencing with strain-level resolution implicates uropathogenic E. coli in necrotizing enterocolitis and mortality in preterm infants. Cell Rep. 14, 2912–2924 (2016).
Wang, Y. et al. 16S rRNA gene-based analysis of fecal microbiota from preterm infants with and without necrotizing enterocolitis. ISME J. 3, 944–954 (2009).
Fernandez-Lopez, R., Redondo, S., Garcillan-Barcia, M. P. & de la Cruz, F. Towards a taxonomy of conjugative plasmids. Curr. Opin. Microbiol 38, 106–113 (2017).
Smillie, C., Garcillan-Barcia, M. P., Francia, M. V., Rocha, E. P. & de la Cruz, F. Mobility of plasmids. Microbiol. Mol. Biol. Rev. 74, 434–452 (2010).
Kovalevskaya, N. P. Mobile gene cassettes and integrons. Mol. Biol. 36, 196–201 (2002).
Rumnieks, J. & Tars, K. Diversity of pili-specific bacteriophages: genome sequence of IncM plasmid-dependent RNA phage M. BMC Microbiol. 12, 277 (2012).
Carattoli, A. Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 2227–2238 (2009).
Thomas, C. M. Paradigms of plasmid organization. Mol. Microbiol. 37, 485–491 (2000).
Bauer, A. W., Kirby, W. M., Sherris, J. C. & Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 45, 493–496 (1966).
Matuschek, E., Brown, D. F. J. & Kahlmeter, G. Development of the EUCAST disk diffusion antimicrobial susceptibility testing method and its implementation in routine microbiology laboratories. Clin. Microbiol. Infect. 20, O255–O266 (2014).
Zaman, M. A., Pasha, M. H. & Akhter, M. Plasmid curing of Escherichia coli cells with ethidium bromide, sodium dodecyl sulfate and acridine orange. Banglad. J. Microbiol. 27, 28–31 (2010).
Levi, K., Rynge, M., Abeysinghe, E. A., & Edwards, R. Searching the sequence read archive using Jetstream and Wrangler. in PEARC '18: Proceedings of the Practice and Experience on Advanced Research Computing, Pittsburgh, PA, USA, July 22–26 (ACM, New York, 2018).
Torres, P. J., Edwards, R. A. & McNair, K. A. PARTIE: a partition engine to separate metagenomic and amplicon projects in the Sequence Read Archive. Bioinformatics 33, 2389–2391 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Towns, J., Cockerill, T., Dahan, M. et al. XSEDE: accelerating scientific discovery. Comput. Sci. Eng. 16, 62–74 (2014).
Lambrecht, E., Van Meervenne, E. & Boon, N. et al. Characterization of cefotaxime- and ciprofloxacin-resistant commensal Escherichia coli originating from belgian farm animals indicates high antibiotic resistance transfer rates. Microb. Drug Resist. 24, 707–717 (2018).
Ravi, A., Avershina, E. & Foley, S. L. et al. The commensal infant gut meta-mobilome as a potential reservoir for persistent multidrug resistance integrons. Sci. Rep. 5, 15317 (2015).
Ravi, A., Valdes-Varela, L., Gueimonde, M. & Rudi, K. Transmission and persistence of IncF conjugative plasmids in the gut microbiota of full-term infants. FEMS Microbiol. Ecol. 94, fix158 (2018).
Smillie, C. S. et al. Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480, 241–244 (2011).
Leverstein-van Hall, M. A., Dierikx, C. M., Cohen Stuart, J. et al. Dutch patients, retail chicken meat and poultry share the same ESBL genes, plasmids and strains. Clin. Microbiol. Infect. 17, 873–880 (2011).
Hughes, D. & Andersson, D. I. Persistence of antibiotic resistance in bacterial populations. FEMS Microbiol. Rev. 35, 901–911 (2011).
Sidjabat, H. E. et al. Expansive spread of IncI1 plasmids carrying blaCMY-2 amongst Escherichia coli. Int. J. Antimicrob. Agents 44, 203–208 (2014).
Riccobono, E. et al. Characterization of IncI1 sequence type 71 epidemic plasmid lineage responsible for the recent dissemination of CTX-M-65 extended-spectrum beta-lactamase in the Bolivian Chaco region. Antimicrob. Agents Chemother. 59, 5340–5347 (2015).
Madec, J. Y., Haenni, M., Metayer, V., Saras, E. & Nicolas-Chanoine, M. H. High prevalence of the animal-associated bla CTX-M-1 IncI1/ST3 plasmid in human Escherichia coli isolates. Antimicrob. Agents Chemother. 59, 5860–5861 (2015).
Madec, J. Y. Sequence type 48 Escherichia coli carrying the blaCTX-M-1 IncI1/ST3 plasmid in drinking water in France. Antimicrob. Agents Chemother. 60, 6430–6432 (2016).
Norizuki, C. et al. Specific blaCTX-M-8/IncI1 plasmid transfer among genetically diverse Escherichia coli isolates between humans and chickens. Antimicrob. Agents Chemother. 61, e00663-17 (2017).
Lopatkin, A. J. et al. Persistence and reversal of plasmid-mediated antibiotic resistance. Nat. Commun. 8, 1689 (2017).
Liebert, C. A., Hall, R. M. & Summers, A. O. Transposon Tn21, flagship of the floating genome. Microbiol. Mol. Biol. Rev. 63, 507–522 (1999).
Shintani, M., Sanchez, Z. K. & Kimbara, K. Genomics of microbial plasmids: classification and identification based on replication and transfer systems and host taxonomy. Front. Microbiol. 6, 242 (2015).
Gaimster, H. & Summers, D. Plasmids in the driving seat: the regulatory RNA Rcd gives plasmid ColE1 control over division and growth of its E. coli host. Plasmid 78, 59–64 (2015).
Yarmolinsky, M. B. Bacteriophage P1 in retrospect and in prospect. J. Bacteriol. 186, 7025–7028 (2004).
Yang, L. et al. Characterization of a P1-like bacteriophage carrying CTX-M-27 in Salmonella spp. resistant to third generation cephalosporins isolated from pork in China. Sci. Rep. 7, 40710 (2017).
Park, H. et al. The success of fecal microbial transplantation in Clostridium difficile infection correlates with bacteriophage relative abundance in the donor: a retrospective cohort study. Gut Microbes. (2019). https://doi.org/10.1080/19490976.2019.1586037.
Babicki, S., Arndt, D. & Marcu, A. et al. Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res. 44, W147–W153 (2016).
Acknowledgements
We thank Norwegian University of Life Sciences for the financial support, and the Spanish Ministry of Science and Universities for the grant AGL2015-70487-P. Travels and stays in Spain and Norway for this study were supported by EEA Coordinated Mobility of Researchers NILS Science and Sustainability Project 017-CM-01-2013.
Authors contributions
M.V., M.C.C., A.R., and G.P.-M. collected the clinical material and isolated the bacteria. M.H. did the main transmission experiments. I.L.A., A.R., and J.L. contributed with the sequencing and sequence interpretation. M.S., S.L.F., and D.B.D. contributed with the phenotypic characterization of the strains. K.R. wrote the paper with input from all the coauthors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hagbø, M., Ravi, A., Angell, I. et al. Experimental support for multidrug resistance transfer potential in the preterm infant gut microbiota. Pediatr Res 88, 57–65 (2020). https://doi.org/10.1038/s41390-019-0491-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-019-0491-8
This article is cited by
-
Outbreak of OXA-48-producing Enterobacteriaceae in a neonatal intensive care unit in Western Sweden
European Journal of Clinical Microbiology & Infectious Diseases (2023)
-
Capturing the antibiotic resistome of preterm infants reveals new benefits of probiotic supplementation
Microbiome (2022)
-
Nasogastric enteral feeding tubes modulate preterm colonization in early life
Pediatric Research (2022)