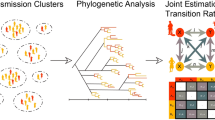
Abstract
The growth of human immunodeficiency virus (HIV) sequence databases resulting from drug resistance testing has motivated efforts using phylogenetic methods to assess how HIV spreads1,2,3,4. Such inference is potentially both powerful and useful for tracking the epidemiology of HIV and the allocation of resources to prevention campaigns. We recently used simulation and a small number of illustrative cases to show that certain phylogenetic patterns are associated with different types of epidemiological linkage5. Our original approach was later generalized for large next-generation sequencing datasets and implemented as a free computational pipeline6. Previous work has claimed that direction and directness of transmission could not be established from phylogeny because one could not be sure that there were no intervening or missing links involved7,8,9. Here, we address this issue by investigating phylogenetic patterns from 272 previously identified HIV transmission chains with 955 transmission pairs representing diverse geography, risk groups, subtypes, and genomic regions. These HIV transmissions had known linkage based on epidemiological information such as partner studies, mother-to-child transmission, pairs identified by contact tracing, and criminal cases. We show that the resulting phylogeny inferred from real HIV genetic sequences indeed reveals distinct patterns associated with direct transmission contra transmissions from a common source. Thus, our results establish how to interpret phylogenetic trees based on HIV sequences when tracking who-infected-whom, when and how genetic information can be used for improved tracking of HIV spread. We also investigate limitations that stem from limited sampling and genetic time-trends in the donor and recipient HIV populations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wertheim, J. O. et al. Social and genetic networks of HIV-1 transmission in New York City. PLoS Pathog. 13, e1006000 (2017).
Pillay, D. et al. PANGEA-HIV: phylogenetics for generalised epidemics in Africa. Lancet Infect. Dis. 15, 259–261 (2015).
Lewis, F., Hughes, G. J., Rambaut, A., Pozniak, A. & Leigh Brown, A. J. Episodic sexual transmission of HIV revealed by molecular phylodynamics. PLoS Med. 5, e50 (2008).
Poon, A. F. et al. Near real-time monitoring of HIV transmission hotspots from routine HIV genotyping: an implementation case study. Lancet HIV 3, e231–e238 (2016).
Romero-Severson, E. O., Bulla, I. & Leitner, T. Phylogenetically resolving epidemiologic linkage. Proc. Natl Acad. Sci. USA 113, 2690–2695 (2016).
Wymant, C. et al. PHYLOSCANNER: analysing within- and between-host pathogen genetic diversity to identify transmission, multipleInfection, recombination and contamination. Preprint at bioRxiv https://doi.org/10.1101/157768 (2017).
Leitner, T. & Albert, J. Reconstruction of HIV-1 transmission chains for forensic purposes. AIDS Rev. 2, 241–251 (2000).
Abecasis, A. B. et al. Science in court: the myth of HIV fingerprinting. Lancet Infect. Dis. 11, 78–79 (2011).
Bernard, E. J., Azad, Y., Vandamme, A. M., Weait, M. & Geretti, A. M. HIV forensics: pitfalls and acceptable standards in the use of phylogenetic analysis as evidence in criminal investigations of HIV transmission. HIV Med. 8, 382–387 (2007).
Romero-Severson, E., Skar, H., Bulla, I., Albert, J. & Leitner, T. Timing and order of transmission events is not directly reflected in a pathogen phylogeny. Mol. Biol. Evol. 31, 2472–2482 (2014).
Volz, E. M., Romero-Severson, E. & Leitner, T. Phylodynamic inference across epidemic scales. Mol. Biol. Evol. 34, 1276–1288 (2017).
Gottlieb, G. S. et al. Dual HIV-1 infection associated with rapid disease progression. Lancet 363, 619–622 (2004).
Grobler, J. et al. Incidence of HIV-1 dual infection and its association with increased viral load set point in a cohort of HIV-1 subtype C-infected female sex workers. J. Infect. Dis. 190, 1355–1359 (2004).
Yang, O. O. et al. Human immunodeficiency virus type 1 clade B superinfection: evidence for differential immune containment of distinct clade B strains. J. Virol. 79, 860–868 (2005).
Smith, D. M. et al. Lack of neutralizing antibody response to HIV-1 predisposes to superinfection. Virology 355, 1–5 (2006).
Smith, D. M. et al. Incidence of HIV superinfection following primary infection. JAMA 292, 1177–1178 (2004).
Korber, B., Hraber, P., Wagh, K. & Hahn, B. H. Polyvalent vaccine approaches to combat HIV-1 diversity. Immunol. Rev. 275, 230–244 (2017).
Ypma, R. J., van Ballegooijen, W. M. & Wallinga, J. Relating phylogenetic trees to transmission trees of infectious disease outbreaks. Genetics 195, 1055–1062 (2013).
McNearney, T. et al. Relationship of human immunodeficiency virus type 1 sequence heterogeneity to stage of disease. Proc. Natl Acad. Sci. USA 89, 10247–10251 (1992).
Wolfs, T. F. W., Zwart, G., Bakker, M. & Goudsmit, J. HIV-1 genomic RNA diversification following sexual parenteral virus transmission. Virology 189, 103–110 (1992).
Zhang, L. Q. et al. Selection for specific sequences in the external envelope protein of human immunodeficiency virus type 1 upon primary infection. J. Virol. 67, 3345–3356 (1993).
Salazar-Gonzalez, J. F. et al. Genetic identity, biological phenotype, and evolutionary pathways of transmitted/founder viruses in acute and early HIV-1 infection. J. Exp. Med. 206, 1273–1289 (2009).
Fischer, W. et al. Transmission of single HIV-1 genomes and dynamics of early immune escape revealed by ultra-deep sequencing. PLoS ONE 5, e12303 (2010).
Shankarappa, R. et al. Consistent viral evolutionary changes associated with the progression of human immunodeficiency virus type 1 infection. J. Virol. 73, 10489–10502 (1999).
Keele, B. F. et al. Identification and characterization of transmitted and early founder virus envelopes in primary HIV-1 infection. Proc. Natl Acad. Sci. USA 105, 7552–7557 (2008).
Rieder, P. et al. Characterization of human immunodeficiency virus type 1 (HIV-1) diversity and tropism in 145 patients with primary HIV-1 infection. Clin. Infect. Dis. 53, 1271–1279 (2011).
Leitner, T., Escanilla, D., Franzén, C., Uhlén, M. & Albert, J. Accurate reconstruction of a known HIV-1 transmission history by phylogenetic tree analysis. Proc. Natl Acad. Sci. USA 93, 10864–10869 (1996).
Lemey, P. et al. Molecular footprint of drug-selective pressure in a human immunodeficiency virus transmission chain. J. Virol. 79, 11981–11989 (2005).
Didelot, X., Fraser, C., Gardy, J. & Colijn, C. Genomic infectious disease epidemiology in partially sampled and ongoing outbreaks. Mol. Biol. Evol. 34, 997–1007 (2017).
Jombart, T. et al. Bayesian reconstruction of disease outbreaks by combining epidemiologic and genomic data. PLoS Comput. Biol. 10, e1003457 (2014).
Kenah, E., Britton, T., Halloran, M. E. & Longini, I. M. Jr. Molecular infectious disease epidemiology: survival analysis and algorithms linking phylogenies to transmission trees. PLoS Comput. Biol. 12, e1004869 (2016).
Kumar, A. et al. Infant transmitted/founder HIV-1 viruses from peripartum transmission are neutralization resistantto paired maternal plasma. PLoS Pathog. 14, e1006944 (2018).
Patel, P. et al. Estimating per-act HIV transmission risk: a systematic review. AIDS 28, 1509–1519 (2014).
Li, H. et al. High multiplicity infection by HIV-1 in men who have sex with men. PLoS Pathog. 6, e1000890 (2010).
Romero-Severson, E. O. et al. Donor–recipient identification in para- and poly-phyletic trees under alternative HIV-1 transmission hypotheses using approximate Bayesian computation. Genetics 207, 1089–1101 (2017).
Aldovini, A. & Young, R. A. Mutations of RNA and protein sequences involved in human immunodeficiency virus type 1 packaging result in production of noninfectious virus. J. Virol. 64, 1920–1926 (1990).
Rusert, P. et al. Quantification of infectious HIV-1 plasma viral load using a boosted in vitro infection protocol. Virology 326, 113–129 (2004).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Foley, B. et al. HIV Sequence Compendium 2015 (Los Alamos National Laboratory, 2015).
Leitner, T., Kumar, S., & Albert, J. Tempo and mode of nucleotide substitutions in gag and env gene fragments in human immunodeficiency virus type 1 populations with a known transmission history. J. Virol. 71, 4761–4770 (1997); erratum 72, 2565 (1998).
Leitner, T., Korber, B. T., Daniels, M., Calef, C. & Foley, B. in HIV Sequence Compendium 2005 (eds Leitner, T. et al.) 41–48 (Theoretical Biology and Biophysics, Los Alamos National Laboratory, 2005).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Acknowledgements
We thank N. Hengartner for advice on statistical analyses, C. Fraser for suggesting the use of Bayes’ rule to illustrate the broader inference problem, and J. Macke and W. Abfalterer for help with database annotation and searches. This study was supported by a NIH/NIAID grant (R01AI087520) and a NIH–DOE interagency agreement (AI2013183).
Author information
Authors and Affiliations
Contributions
T.L. designed the study and compiled the data, T.L. and E.R.-S. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–3 and Supplementary Table 1 footnote.
Supplementary Table 1
Excel file summarizing data features per epidemiological pair.
Rights and permissions
About this article
Cite this article
Leitner, T., Romero-Severson, E. Phylogenetic patterns recover known HIV epidemiological relationships and reveal common transmission of multiple variants. Nat Microbiol 3, 983–988 (2018). https://doi.org/10.1038/s41564-018-0204-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0204-9
This article is cited by
-
HIV-1 drug resistance and genetic transmission network among newly diagnosed people living with HIV/AIDS in Ningbo, China between 2018 and 2021
Virology Journal (2023)
-
Broadly neutralizing plasma antibodies effective against autologous circulating viruses in infants with multivariant HIV-1 infection
Nature Communications (2020)
-
Inferring HIV-1 transmission networks and sources of epidemic spread in Africa with deep-sequence phylogenetic analysis
Nature Communications (2019)